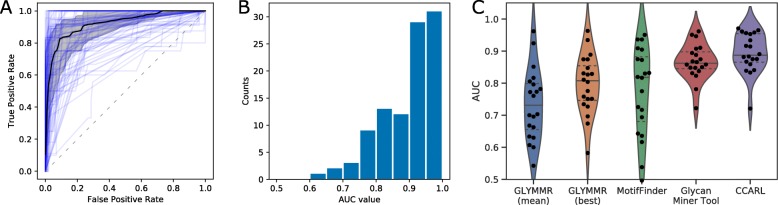
Fig. 7.
Classification performance across a range of different lectins. a Receiver-operator characteristic (ROC) curves across a number of different glycan microarray experiments. Individual ROC curves are shown in light blue. The median ROC curve is shown in black, with shading representing 25th-75th percentiles. The dashed line indicates an uninformative (random) classifier. b Area Under the Curve (AUC) values for all glycan microarray experiments examined. See Table 1 and Additional file 5 for a full list of lectins examined. c Classification performance of CCARL compared to existing glycan motif tools. Area Under the Curve (AUC) values were calculated across a number of different glycan microarray experiments using stratified 5-fold cross-validation (with the exception of MotifFinder, which was evaluated using a single fold). Motifs were extracted using GLYMMR, MotifFinder, the Glycan Miner Tool and CCARL, and assessed using a logistic regression model (with the exception of MotifFinder, which outputs predicted RFU values). Motifs from GLYMMR were extracted at several minimum support thresholds, and both the mean AUC value and best AUC value reported for each microarray experiment. Median and interquartile range are indicated by solid and dashed grey lines respectively