Abstract
Accurate target recognition in transcript degradation is crucial for regulation of gene expression. In the fission yeast Schizosaccharomyces pombe, a number of meiotic transcripts are recognized by a YTH-family RNA-binding protein, Mmi1, and selectively degraded by the nuclear exosome during mitotic growth. Mmi1 forms nuclear foci in mitotically growing cells, and the nuclear exosome colocalizes to such foci. However, it remains elusive how Mmi1 and the nuclear exosome are connected. Here, we show that a complex called MTREC (Mtl1-Red1 core) or NURS (nuclear RNA silencing) that consists of a zinc-finger protein, Red1, and an RNA helicase, Mtl1, is required for the recruitment of the nuclear exosome to Mmi1 foci. Physical interaction between Mmi1 and the nuclear exosome depends on Red1. Furthermore, a chimeric protein involving Mmi1 and Rrp6, which is a nuclear-specific component of the exosome, suppresses the ectopic expression phenotype of meiotic transcripts in red1Δ cells and mtl1 mutant cells. These data indicate that the primary function of MTREC/NURS in meiotic transcript elimination is to link Mmi1 to the nuclear exosome physically.
Author summary
Since abnormal gene expression is detrimental to proliferation, cells possess a number of mechanisms to regulate gene expression at transcriptional and post-transcriptional levels. In particular, expression of meiotic genes is rigorously repressed in somatic cells because their aberrant expression causes severe cellular defects including genome instability and tumorigenesis. In the fission yeast Schizosaccharomyces pombe, selective degradation of meiotic transcripts is employed to prevent their deleterious expression during mitotic growth. Meiotic transcripts are recognized by a YTH-family RNA-binding protein, Mmi1. Mmi1 then induces their selective degradation by the nuclear exosome, which is a highly conserved exonuclease complex. However, little is known how Mmi1 cooperates with the nuclear exosome. Here, we demonstrate that the interaction of Mmi1 with the nuclear exosome is mediated by a complex termed MTREC/NURS that is composed of a conserved zinc-finger protein, Red1, and an RNA helicase, Mtl1. Our findings shed light on the target recognition mechanisms of post-transcriptional regulation.
Introduction
Stability control of transcripts is crucial to establish appropriate gene expression profiles in a wide range of biological processes [1, 2]. In the fission yeast Schizosaccharomyces pombe, expression of a number of meiotic genes is strictly repressed during the mitotic cell cycle by selective RNA elimination, in addition to transcriptional suppression, since their aberrant expression is highly deleterious to cell growth [3, 4]. Selectivity in meiotic RNA degradation is guaranteed by a particular sequence element termed DSR (determinant of selective removal) on meiotic transcripts, which is composed of the repetitive hexanucleotide motif UNAAAC [3, 5–7]. Meiotic transcripts are recognized by a YTH family RNA-binding protein, Mmi1, through direct binding of the YTH domain with UNAAAC DSR motifs [3, 6]. The mode of Mmi1 interaction with RNAs is different from that of other YTH proteins, and Mmi1 does not recognize N6-methyladenosine-containing RNAs [8–11].
Mmi1 induces selective degradation of DSR-containing meiotic transcripts by the nuclear exosome [3, 12]. The exosome is a highly conserved 3′-5′ exonuclease complex that localizes to both the nucleus and cytoplasm [13–15]. The cytoplasmic exosome consists of a nine-subunit inert complex and a catalytically active subunit, Dis3. In the nucleus, an additional exonuclease, Rrp6, binds to the nine-subunit complex in addition to Dis3. Recent genome-wide analyses have revealed that the exosome targets various classes of coding and non-coding transcripts [16–18]. The exosome cooperates with a variety of adapter complexes for selective degradation of divergent target transcripts. Mmi1-mediated selective elimination of meiotic transcripts requires polyadenylation-related factors, including a canonical poly(A) polymerase, Pla1, and a poly(A)-binding protein, Pab2 [12, 19, 20]. Another pivotal factor engaged in Mmi1-mediated RNA degradation is a zinc-finger protein, Red1 [21, 22]. Red1, together with Mtr4-like RNA helicase protein, Mtl1, constitutes a complex termed MTREC (Mtl1-Red1 core) or NURS (nuclear RNA silencing), and mediates the selective degradation of various transcripts, including not only meiotic RNAs, but also cryptic unstable transcripts (CUTs), the latter of which are independent of Mmi1 [23–25]. The machinery of Mmi1-mediated RNA degradation also triggers facultative heterochromatin formation at a subset of its target genes [5, 26, 27]. Furthermore, it regulates transcription termination of its targets [28–30], suggesting that Mmi1-mediated multilayered regulation rigorously shapes appropriate gene expression profiles in mitotically growing cells.
Mmi1 forms nuclear foci in vegetatively growing cells [3]. Most, if not all, factors cooperating with Mmi1, including the nuclear exosome, colocalize with Mmi1 at nuclear foci [12, 21, 22, 24, 31, 32]. We have recently shown that DSR-containing meiotic transcripts are tethered to the Mmi1 foci [33]. Tethering of meiotic transcripts leads to prevention of their nuclear export and deleterious expression of meiotic proteins, even when RNA elimination is compromised [33]. Foci formation of Mmi1 requires its self-interaction with the assistance of a homolog of Enhancer of Rudimentary, Erh1, which induces Mmi1 dimerization [33, 34]. Mmi1 mutant cells lacking the self-interaction domain (SID) and erh1 deletion mutant cells do not form Mmi1 foci. In these mutant cells, DSR-containing transcripts are liberated from the nuclear foci and escape degradation. These findings highlight the importance of spatial control in Mmi1-mediated regulation. However, it remains largely unknown how factors involved in Mmi1-mediated regulation are incorporated into the nuclear foci. The Red1-containing complex MTREC/NURS binds to both Mmi1 and the nuclear exosome [23–25], suggesting that Red1 acts as a bridge between the nuclear exosome and Mmi1. However, Red1 function in Mmi1-mediated transcript elimination remains to be elucidated.
Here, we show that recruitment of the nuclear exosome to nuclear foci is Red1-dependent. We further demonstrated that Red1 mediates physical interactions between Mmi1 and the nuclear exosome. Our current study sheds new light on our understanding of accurate target recognition in nuclear RNA degradation.
Results
Red1 is crucial for nuclear exosome foci formation
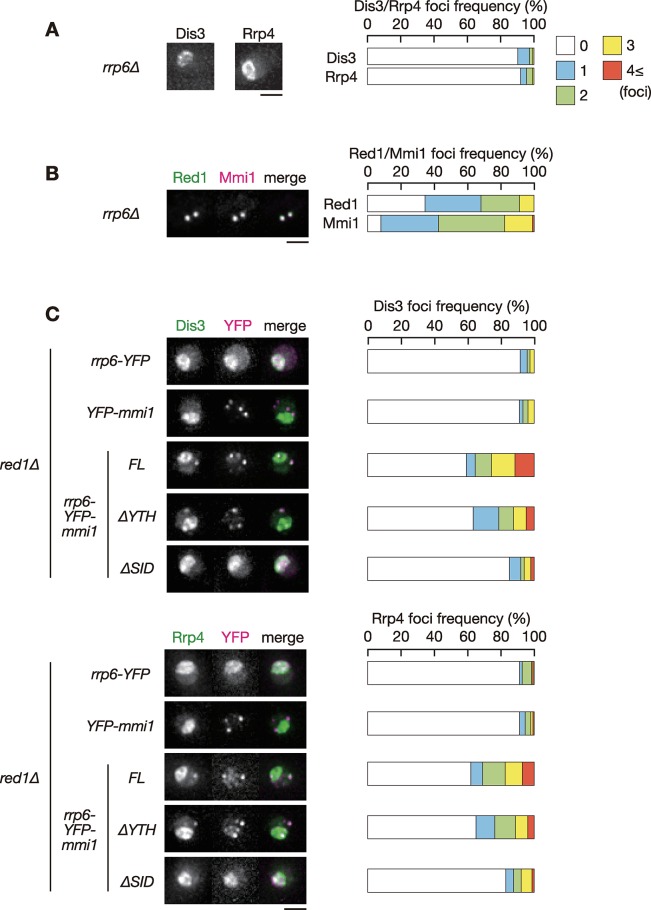
To elucidate the mechanisms underlying exosome foci formation, we searched for factors required for the proper localization of Rrp6. We first examined the effect of deletion of the red1, mmi1, and pab2 genes, whose gene products colocalize with nuclear foci of Rrp6 [12, 21]. Deletion of mmi1 causes severe growth defects due to the ectopic expression of meiotic transcripts. Growth defects of mmi1 deletion can be alleviated by deletion of mei4, a crucial target of Mmi1 [3]. Thus, we used a mmi1Δ mei4Δ double mutant for examining the impact of mmi1 deletion. Wild-type cells showed nucleolar localization of Rrp6. Rrp6 also formed foci in both nucleoplasm and nucleolus (Fig 1A and S1A Fig). Strikingly, nuclear foci formation of Rrp6 was severely impaired in red1Δ cells, whereas nucleolar accumulation was maintained. Cells lacking mmi1 exhibited only a modest reduction in the frequency of Rrp6 foci formation and the number of foci per cell, indicating weak contribution of Mmi1 in Rrp6 foci formation. Foci formation of Mmi1 and Red1 was not interdependent (Fig 1A and S1A Fig) [22, 33], and they might play divergent roles in Rrp6 foci formation. The localization of Rrp6 in pab2Δ cells was comparable to that observed in wild-type cells. We also found that deletion of red1 strongly impeded foci formation of both Dis3 and Rrp4, the core components of the exosome (Fig 1B and S1B Fig). Since the fluorescence signal of Dis3 and Rrp4 in the nucleolus in wild-type cells was more intense than that of Rrp6, the frequency of foci formation in or on the border of the nucleolus, and in nucleoplasm, might be underestimated. Deletion of red1 had no severe impact on nuclear foci formation of Pla1 and Pab2, although percentages of cells containing 1 focus were increased (Fig 1C and 1D, S1C Fig and S1D Fig). From these observations, we concluded that Red1 is a key player in nuclear foci formation of the exosome and regulates the localization of a subset of factors involved in Mmi1-mediated RNA degradation.
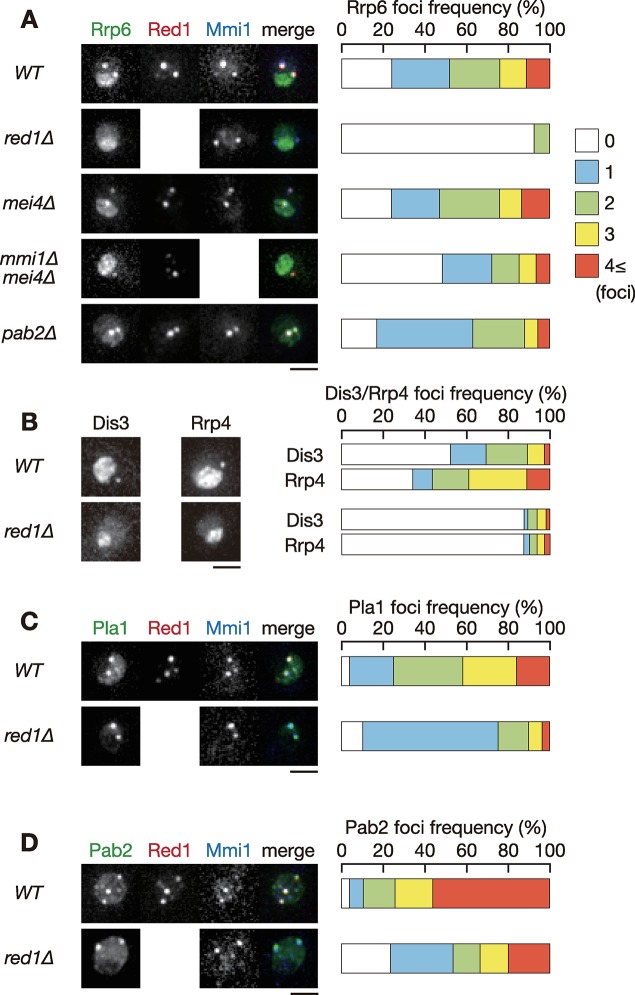
Fig 1. Red1 is required for nuclear foci formation of exosome.
(A) Localization of Rrp6, Red1 and Mmi1 in wild-type, red1Δ, mei4Δ, mmi1Δ mei4Δ, and pab2Δ cells. Cells expressing Rrp6-YFP (green), Red1-mCherry (red) or CFP-Mmi1 (blue) from the respective endogenous loci were observed during exponential growth in YE liquid medium. Images of the nuclear region are shown. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Rrp6 foci are indicated on the right (n > 100). (B) Localization of Dis3 and Rrp4 in wild-type and red1Δ cells. red1Δ cells expressing Dis3-GFP or Rrp4-GFP from the respective endogenous loci were observed. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Dis3 or Rrp4 foci are indicated (n > 100). (C) Localization of Pla1, Red1 and Mmi1 in wild-type and red1Δ cells. Cells expressing Pla1-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Pla1 foci are indicated (n > 100). (D) Localization of Pab2, Red1, and Mmi1 in wild-type and red1Δ cells. Cells expressing Pab2-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Pab2 foci are indicated (n > 100). Scale bars: 2 μm.
Rrp6 foci formation is dependent on an N-terminal domain of Red1
We next determined which region of Red1 was essential for nuclear foci formation. Since it has been reported that the C-terminal zinc-finger motif is required for the elimination of meiotic transcripts but not for localization [21], we constructed N-terminal deletion mutants according to secondary structure predictions by JPred (Fig 2A) [35]. Comparable expression of the Red1 deletion series was confirmed by western blot analysis (S2A Fig). The deletion of residues 2 to 195 did not alter the localization of Red1 (Fig 2A and 2B). When residues 2 to 347 was deleted, Red1 failed to form nuclear foci and was, rather, uniformly localized in the nucleus. The deletion of residues 196 to 347 also resulted in uniform nuclear localization. We then divided the domain involving residues 196 to 347 into three regions, and found that residues 196 to 245 were responsible for nuclear foci formation of Red1.
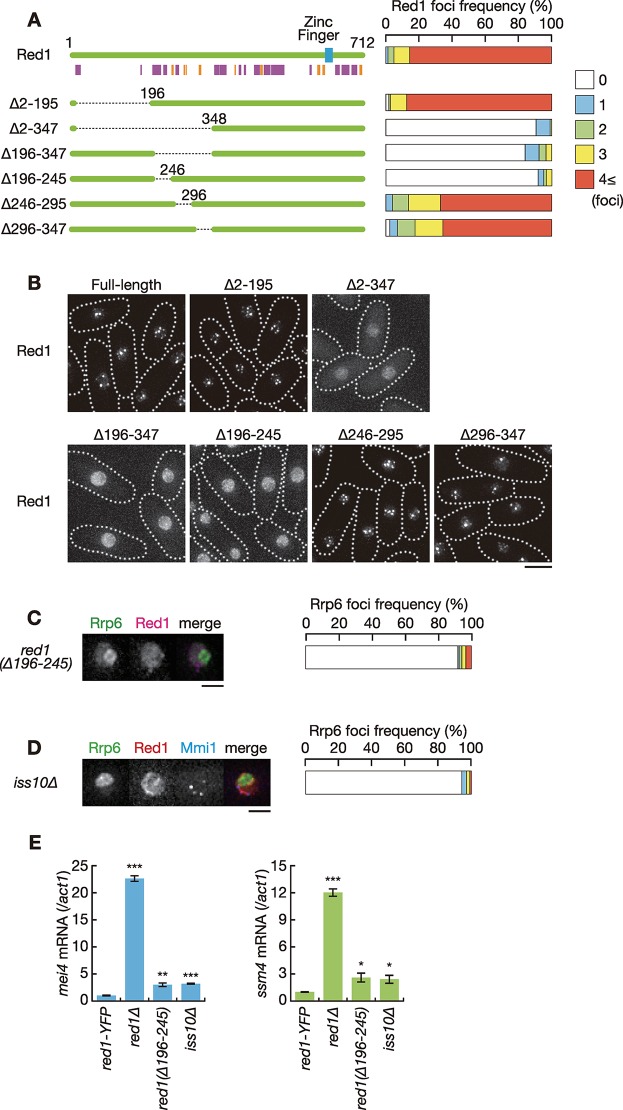
Fig 2. Red1 forms nuclear foci through the N-terminal domain.
(A) Schematic of the structure of Red1 and its truncation series. The blue rectangle represents a putative zinc-finger motif. Predicted alpha helices and beta sheets are shown in magenta and orange, respectively. Frequencies of cells carrying 0, 1, 2, 3, or 4 and more Red1 foci are indicated on the right (n > 100). (B) Localization of full-length and truncated Red1. YFP-tagged full-length or truncated Red1 were expressed from the endogenous locus. Dotted lines indicate the shape of cells. Scale bar: 5 μm. (C) Localization of Rrp6 and Red1 in red1(Δ196–245) cells. red1(Δ196–245) cells expressing Rrp6-YFP (green) and Red1-mCherry (magenta) from the respective endogenous loci were observed. Images of the nuclear region are shown. Frequency of cells containing 0, 1, 2, 3, or 4 and more Rrp6 foci is indicated on the right (n > 100). Scale bar: 2 μm. (D) Localization of Rrp6, Red1 and Mmi1 in iss10Δ cells. iss10Δ cells expressing Rrp6-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Frequency of cells containing 0, 1, 2, 3, or 4 and more Rrp6 foci is indicated (n > 100). Scale bar: 2 μm. (E) Expression of mei4 mRNA and ssm4 mRNA in wild-type (red1-YFP), red1Δ, red1(Δ196–245), and iss10Δ cells. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01; ***P < 0.001 compared with the wild-type red1-YFP strain (Student’s t-test).
We next examined Rrp6 localization in cells expressing Red1 lacking residues 196 to 245 (red1(Δ196–245)), and found that this domain is vital for Rrp6 nuclear foci formation (Fig 2C and S2B Fig). We also tested the effect of deletion of iss10, which is essential for Red1 nuclear foci formation [22, 24]. iss10Δ cells exhibited a strong reduction in Rrp6 foci formation frequency (Fig 2D and S2C Fig), suggesting that the proper localization of Red1 is crucial for nuclear foci formation of Rrp6.
The growth profile of red1(Δ196–245) cells was similar to that of wild-type cells, while red1Δ cells grew more slowly than wild-type cells and exhibited severe cold sensitivity (S3A Fig) [21, 22]. iss10Δ cells showed mild cold sensitivity, as previously demonstrated [22]. The cold sensitivity of red1Δ is not suppressed by deletion of mei4, indicating that the defect arises independently of ectopic accumulation of meiotic transcripts [32, 33]. Meiotic transcripts, such as mei4, ssm4 and rec8 mRNAs, were accumulated in red1(Δ196–245) and iss10Δ cells during vegetative growth, albeit to a lesser extent than red1Δ cells (Fig 2E and S3B Fig). The effect of the deletion of residues 196 to 245 was less prominent on expression of spo5 mRNAs, which are also targeted by Mmi1. These results suggest that Red1 has some activity without this region or in the absence of Iss10 and that residues 196 to 245 is required to fully induce meiotic transcript degradation.
Red1 has been shown to mediate degradation of CUTs in an Mmi1-independent manner [25]. We then examined the expression of the promoter upstream transcripts (PROMPTs) of the cti6 gene and the rpl402 gene, which are known to be CUTs transcribed from the promoter region of cti6 or rpl402 in anti-sense direction. In contrast to meiotic transcripts such as ssm4, rec8 and spo5, ectopic expression of the cti6 and rpl402 PROMPT was detected in red1Δ and rrp6Δ cells, but not in mmi1Δ cells (S3C Fig), as shown previously [25]. red1(Δ196–245) and iss10Δ cells showed a modest, if any, defect in elimination of the cti6 and rpl402 PROMPT, as was the case with meiotic transcripts (S3D Fig). Altogether, the 196–245 domain of Red1 was required for nuclear foci formation of itself and Rrp6, and played a significant role in regulation of meiotic transcript and CUT expression, although it was dispensable for growth at low temperatures.
Red1 connects Rrp6 with Mmi1 for recruiting target RNAs to the exosome
Red1 physically interacts with both Rrp6 and Mmi1 [21]. To test whether Red1 mediates the interaction between Mmi1 and Rrp6, we carried out co-immunoprecipitation assays. Mmi1 was specifically co-purified with Rrp6 (S4A Fig). RNase treatment did not affect the interaction of Mmi1 with Rrp6 (S4B Fig), indicating that this interaction is independent of RNA molecules. The interaction between these proteins was abolished in the absence of red1 (Fig 3A), suggesting that Red1 physically links Mmi1 with the exosome. The deletion of residues 196 to 245 in Red1 mildly impaired the interaction between Mmi1 and Rrp6 owing to reduction of the interaction of Red1 with Mmi1 (Fig 3B).
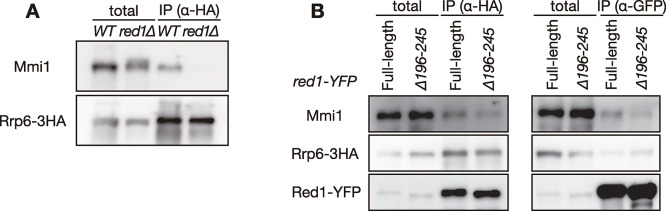
Fig 3. Red1 is required for interaction between Rrp6 and Mmi1.
(A) Co-immunoprecipitation of Rrp6 and Mmi1 in wild-type and red1Δ cells. Native cell extracts were prepared from exponentially growing cells expressing Rrp6-3HA in liquid MM medium and subjected to immunoprecipitation with anti-HA antibody. Precipitates and 10% total cell extracts were then immunoblotted with anti-Mmi1 and anti-HA antibodies. (B) Co-immunoprecipitation of Rrp6 and Mmi1 (left), or Red1 and Mmi1 (right) in wild-type and red1(Δ196–295) cells. Native cell extracts were subjected to immunoprecipitation with anti-HA antibody or anti-GFP antibody. Precipitates and 10% total cell extracts were then immunoblotted with anti-Mmi1, anti-HA and anti-GFP antibodies.
The above results implied that gene expression defects in red1Δ cells could be ascribed to impaired interaction between Mmi1 and Rrp6. We investigated this possibility by expressing a chimeric Mmi1-Rrp6 protein in red1Δ cells. We constructed a plasmid carrying a chimeric gene in which GFP was inserted between full-length rrp6 and mmi1 (rrp6-GFP-mmi1) ORFs. The chimeric protein Rrp6-GFP-Mmi1 was able to suppress the growth defect, and ectopic accumulation of meiotic transcripts and CUTs in rrp6Δ cells (S5A and S5B Fig). Rrp6-GFP-Mmi1 also suppressed the growth defect and ectopic accumulation of meiotic transcripts in temperature-sensitive mmi1 mutant cells (mmi1-ts3) (S5C and S5D Fig). In these experiments, cells were cultured in minimal medium to induce expression of the chimeric protein. Since mmi1Δ cells show growth retardation in minimal medium even when combined with the mei4 deletion, we used the temperature-sensitive mmi1 mutant instead of mmi1Δ. These data indicated that Rrp6-GFP-Mmi1 maintains the functions of both Rrp6 and Mmi1. Intriguingly, aberrant accumulation of mei4, ssm4, rec8 and spo5 mRNAs in red1Δ cells was abrogated when Rrp6-GFP-Mmi1 was expressed, whereas not by either Rrp6 or Mmi1 alone (Fig 4A and S6A Fig). It should be noted that the use of minimal medium in these experiments resulted in the lower accumulation of meiotic transcripts than that in rich medium (S3 Fig). Meanwhile, cti6 and rpl402 PROMPTs still accumulated in red1Δ cells even when Rrp6-GFP-Mmi1 was expressed (Fig 4A and S6A Fig). Rrp6-GFP-Mmi1 did not suppress the cold sensitivity of red1Δ cells (S6B Fig), consistently with that the growth defect caused by red1 deletion is irrelevant to ectopic expression of meiotic transcripts [32, 33]. We found that the slight growth retardation of red1Δ cells at 25 to 36˚C, which has been shown previously [21, 22], was suppressed by expression of Rrp6, Mmi1 or Rrp6-GFP-Mmi1 as well as Red1, although the underlying mechanism remains unclear.
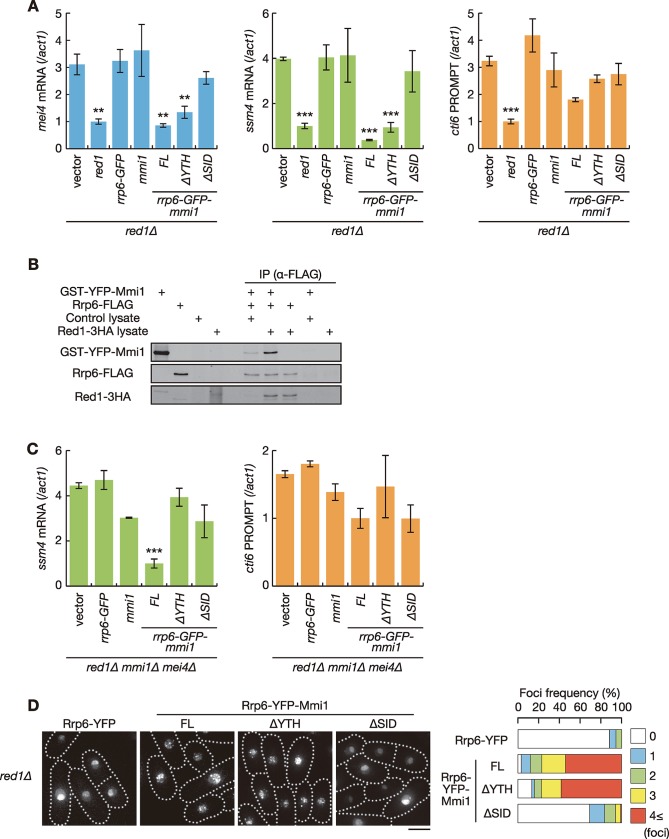
Fig 4. Interaction between Rrp6 and Mmi1 via Red1 is essential for meiotic transcript elimination.
(A) Expression of mei4 mRNA, ssm4 mRNA and cti6 PROMPT in red1Δ cells expressing Red1, Rrp6-GFP, Mmi1 or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium. Transcripts of each gene were analyzed by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. **P < 0.01; ***P < 0.001 compared with cells carrying empty vector (Student’s t-test). (B) In vitro interaction between Mmi1 and Rrp6. Purified Rrp6-FLAG was incubated with purified GST-YFP-Mmi1 and reticulocyte lysate expressing Red1-3HA or control lysate, and was precipitated with anti-FLAG antibody. Precipitates were then immunoblotted with anti-GFP, anti-HA and anti-FLAG antibodies. (C) Expression of ssm4 mRNA and cti6 PROMPT in red1Δ mmi1Δ mei4Δ cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium. Transcripts were analyzed by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. ***P < 0.001 compared with cells carrying empty vector (Student’s t-test). (D) Localization of Rrp6-Mmi1 fusion proteins. red1Δ cells expressing Rrp6-YFP or chimeric proteins composed of Rrp6, YFP, and full-length or truncated Mmi1 from plasmids were observed during exponential growth. Frequencies of cells containing 0, 1, 2, 3, or 4 and more nuclear foci are indicated on the right (n > 100). Dotted lines indicate the shape of cells. Scale bar: 5 μm.
We next confirmed the Red1-mediated interaction between Mmi1 and Rrp6 by in vitro binding assay (Fig 4B). The weak direct interaction of Mmi1 with Rrp6 was observed in our experimental conditions. Addition of Red1 greatly increased the interaction, indicating the role of Red1 in connecting Mmi1 to the exosome. The direct binding of Red1 with Rrp6 was also observed (Fig 4B).
From these results, we concluded that the physical interaction between Rrp6 and Mmi1 is crucial for the elimination of meiotic transcripts, but not for other function(s) involving Red1.
Self-interaction of Mmi1 is vital for meiotic transcript elimination by Rrp6
We previously demonstrated that Mmi1 interacts with itself via the self-interaction domain, SID, which is essential for the proper function and localization of Mmi1 [33]. Given that Rrp6-GFP-Mmi1 functions by interacting with endogenously expressed Mmi1, the YTH-RNA-binding domain in the chimeric protein could be unnecessary for meiotic transcript elimination because target transcripts might be recognized by endogenous Mmi1 and delivered to Rrp6 in the chimeric protein through Mmi1-Mmi1 interaction. To test this hypothesis, we constructed a plasmid carrying a fusion gene of rrp6, GFP or YFP, and mmi1-ΔYTH or mmi1-ΔSID, lacking the region encoding the YTH-RNA-binding domain or the SID, respectively. We confirmed that chimeric proteins were expressed in a comparable amount, although removal of the SID resulted in an increase of the expression level (S6C Fig). Both chimeric proteins maintained the function of Rrp6 (S5A and S5B Fig), but lost the ability to act as Mmi1 (S5C and S5D Fig). The chimeric protein containing Mmi1-ΔYTH showed a negative effect in mmi1-ts3 cells at permissive temperatures (S5C and S5D Fig). This is because Mmi1 lacking the YTH domain exerts a dominant-negative activity, as shown previously [33]. Rrp6-GFP-Mmi1-ΔYTH amended the degradation defect of mei4, ssm4, rec8 and spo5 mRNAs in red1Δ cells, but Rrp6-GFP-Mmi1-ΔSID did not (Fig 4A and S6A Fig). Rrp6-GFP-Mmi1-ΔYTH did not suppress the degradation defect in red1Δ mmi1Δ cells (Fig 4C and S6D Fig), indicating that Rrp6-GFP-Mmi1-ΔYTH requires endogenous Mmi1 to exert the function in the absence of Red1. The interaction between Rrp6-GFP-Mmi1 and endogenous Mmi1 was demonstrated by co-immunoprecipitation (S6E Fig). This interaction was dependent on the SID but not on the YTH domain, as predicted.
We next examined the localization of the chimeric proteins. YFP instead of GFP was used for the observation to accommodate the microscope filter set. Rrp6-YFP-Mmi1 formed nuclear foci in red1Δ cells (Fig 4D). The absence of the YTH domain did not affect the localization of the chimeric protein. However, SID deletion significantly reduced the frequency of foci formation. These observations suggested that Mmi1 self-interaction is crucial for foci formation of the chimeric proteins, and for meiotic transcript degradation by Rrp6, although we cannot exclude the possibility that the SID is required for the interaction of Mmi1 with other factors required for meiotic RNA elimination.
Rrp6 is required for foci formation of other exosome components
Rrp6 mediates the interaction between core components of the exosome and various exosome-related factors [36]. To examine whether Rrp6 acts as a mediator between Red1 and other exosome components, we observed the localization of core components of the exosome in rrp6Δ cells. Dis3 and Rrp4 failed to form foci in the absence of rrp6 (Fig 5A and S7A Fig), while Red1 and Mmi1 showed similar localization to wild-type cells (Fig 5B and S7B Fig). In red1Δ cells, in which foci formation of Rrp6, Dis3 and Rrp4 was severely impaired (Fig 1A and 1B), Rrp6-YFP-Mmi1 partially recovered the foci formation frequencies of Dis3 and Rrp4 (Fig 5C and S7C Fig). Foci formation frequency was similar even when the YTH domain was absent. The recovery of Dis3 and Rrp4 foci formation depended on the SID of Mmi1 in the chimeric protein. These results suggested that Rrp6 plays a key role in the assembly of the exosome at nuclear foci containing Mmi1 and Red1.
Fig 5. Rrp6 is vital for nuclear foci formation of other exosome components.
(A) Localization of Dis3 and Rrp4 in rrp6Δ cells. rrp6Δ cells expressing Dis3-GFP or Rrp4-GFP from the respective endogenous loci were observed during exponential growth in liquid YE medium. Images of the nuclear region are shown. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Dis3 or Rrp4 foci are indicated on the right (n > 100). (B) Localization of Red1 and Mmi1 in rrp6Δ cells. rrp6Δ cells expressing Red1-YFP (green) and CFP-Mmi1 (magenta) were examined. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Red1 or Mmi1 foci are indicated on the right (n > 100). (C) Localization of Dis3 and Rrp4 in red1Δ cells expressing Rrp6-YFP, YFP-Mmi1, or chimeric proteins composed of Rrp6, YFP, and full-length or truncated Mmi1. Dis3-mCherry and Rrp4-mCherry were expressed from the respective endogenous loci (green) and YFP-containing chimeric proteins were expressed from plasmids (magenta) in liquid SD medium. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Dis3 or Rrp4 are indicated on the right (n > 100). Scale bars: 2 μm.
Red1 links Mmi1 to the exosome in cooperation with Mtl1
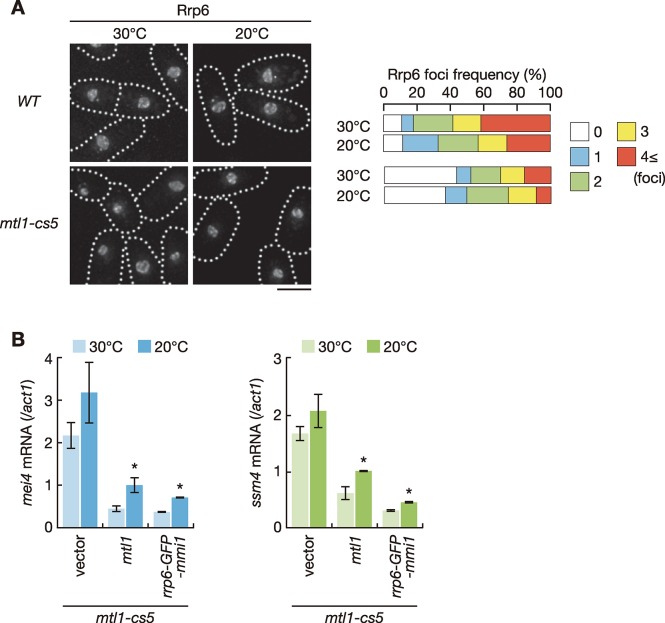
We next investigated whether the function of Red1 to connect Mmi1 with the exosome required Mtl1, a component of the Red1-containing MTREC/NURS complex [23, 24]. Nuclear foci formation of Rrp6 was compromised in mtl1 mutant cells (Fig 6A). The relatively mild effect compared to that in red1Δ cells (Fig 1A) might arise from the use of a conditional cold-sensitive mutant, since Mtl1 is essential for cell growth. We further examined the expression of meiotic transcripts in the mtl1-cs5 cells expressing Rrp6-GFP-Mmi1. Expression of the chimeric protein suppressed aberrant accumulation of meiotic mRNAs as efficiently as that of wild-type Mtl1 (Fig 6B and S8 Fig). These observations suggest that Mtl1 contributes to connect Mmi1 and the nuclear exosome by forming the MTREC/NURS complex with Red1.
Fig 6. Mtl1 mediates interaction between Rrp6 and Mmi1 to eliminate meiotic transcripts.
(A) Localization of Rrp6 in wild-type and mtl1-cs5 cells. Cells expressing Rrp6-mCherry from the endogenous locus were observed during exponential growth at 30˚C in liquid YE medium and shifted to 20°C for 2 hours. Dotted lines indicate the shape of cells. Frequencies of cells containing 0, 1, 2, 3, or 4 and more Rrp6 foci are indicated on the right (n > 100). Scale bar: 5 μm. (B) Expression of mei4 mRNA and ssm4 mRNA in mtl1-cs5 cells expressing Mtl1 and Rrp6-GFP-Mmi1 from plasmids. Cells were grown in liquid MM medium at 30°C and shifted to 20°C for 2 hours. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05 compared with cells carrying empty vector at 20˚C (Student’s t-test).
Discussion
In this study, we demonstrated that the primary function of Red1 in meiotic transcript elimination is to connect Mmi1 and the nuclear exosome physically. We have also shown that Rrp6 is crucial for nuclear foci formation of the core exosome. This observation is consistent with the result of previous co-purification assays, which showed that the interaction between exosome subunits and Red1 depends on Rrp6 [25]. Red1 forms the complex, MTREC or NURS, which includes an RNA helicase, Mtl1 [23, 24]. Our observations also imply that the MTREC/NURS complex, rather than Red1 alone, may link Mmi1 to the exosome.
We found that foci formation of Rrp6 was severely impaired by the deletion of residues 196 to 245 in Red1. Meanwhile, ectopic expression levels of meiotic transcripts were moderate in red1(Δ196–245) cells compared to those in red1Δ cells. These results suggest that nuclear foci, to which Rrp6 and Mmi1 localize, might not be the exclusive sites for Mmi1-mediated RNA degradation. Indeed, the localization of factors involved in Mmi1-mediated degradation is not limited to nuclear foci; Mmi1, Red1 and Rrp6 have been shown to localize to the loci encoding target transcripts including mei4 by chromatin immunoprecipitation experiments [23, 24, 26, 27, 32, 37], although their colocalization with the mei4 locus was not observed by microscopic observations [24, 37]. Thus, it is plausible that Mmi1-mediated RNA degradation also takes place at the genetic loci where its targets are transcribed. Further clarification of the site(s) of RNA degradation would be important for understanding of Mmi1-mediated regulation.
Red1 exerts various functions beyond meiotic transcript elimination. For instance, Red1 plays an essential role in degradation of CUTs by the nuclear exosome, independently of Mmi1 [25]. Red1 is also required for growth at low temperatures [21, 22], although it remains elusive what Red1 does under cold conditions. It is interesting whether Red1 physically links a target-recognition factor to the degradation machinery, such as the nuclear exosome, in other situations as in meiotic transcript elimination. Another intriguing question is whether Red1 function is conserved in other organisms. A human zinc-finger protein, ZFC3H1, which is suggested to be a counterpart of Red1, forms a complex termed PAXT (poly(A) tail exosome targeting) together with an Mtl1-ortholog, hMtr4, and plays an important role in the selective elimination of polyadenylated nuclear RNAs [38, 39]. Intriguingly, ZFC3H1 is required not only for RNA degradation but also retention of target transcripts to nuclear foci [39, 40]. We have demonstrated that Red1 is dispensable for tethering of meiotic transcripts to nuclear foci, while Mmi1 prevents nuclear export of its targets by sequestering them to foci even when RNA degradation is dampened [33]. Future studies may clarify the conservation of Red1 function in higher eukaryotes.
Materials and Methods
Yeast strains and general genetic methods
The S. pombe strains used in this study are listed in S1 Table. The general genetic manipulation procedures and growth media have been previously described [41, 42]. Deletion mutants and epitope-tagged strains were constructed using PCR-based gene-targeting protocols [43, 44].
To construct truncated red1 mutant strains, we cloned the red1 open reading frame (ORF) with 1 kb upstream and downstream flanking regions and removed the indicated regions using the PrimeStar Mutagenesis kit (Takara, Shiga, Japan). PCR-amplified DNA fragments encompassing each truncated red1 ORF with flanking regions were introduced into the red1::ura4+ strain. Transformants were counter-selected on medium containing 5-fluoroorotic acid.
Plasmids expressing chimeric rrp6-GFP-mmi1 gene and its variants were constructed by cloning the PCR-amplified rrp6 and full-length or truncated mmi1 ORFs into pREP81 carrying the green fluorescent protein (GFP) ORF, in which the fusion genes were expressed under the control of a modified thiamine-repressive nmt1 promoter [45]. To observe the localization and expression levels of chimeric proteins (Figs 4D and 5C, S6C Fig and S7C Fig), GFP was replaced with yellow fluorescent protein (YFP) and the fusion genes were expressed from the modified constitutively active adh1 promoter.
Fluorescence microscopy
Cells were grown in logarithmic phase at 30°C and mounted onto lectin-coated glass bottom culture dishes (MatTek, Ashland, MA) filled with liquid growth medium. Images were acquired using the DeltaVision-SoftWoRx system (GE Healthcare, Chicago, IL) by collecting 12 optical sections along the z-axis at 0.5 μm intervals. All images were deconvolved and merged into single projections using SoftWoRx software.
RNA preparation and reverse-transcription quantitative PCR (RT-qPCR) analysis
Total RNA extraction and RT-qPCR analysis were performed as previously described [33] by using ReverTra Ace qPCR Master Mix (TOYOBO, Osaka, Japan) and a LightCycler96 instrument (Roche, Basel, Switzerland) with SYBR Premix Ex Taq II (Takara). Normalization was performed using act1, which encodes actin. Primers used in this study are listed in S2 Table.
Immunoprecipitation and western blot analysis
Immunoprecipitation was performed as previously described [46]. Hemagglutinin (HA)-tagged Rrp6 was precipitated using mouse anti-HA-tag magnetic beads M180-11 (MBL, Nagoya, Japan). Red1-YFP and GFP-containing chimeric proteins were precipitated and detected using anti-GFP-tag magnetic beads D153-11 (MBL) and anti-GFP antibody [47], respectively. Rabbit anti-Mmi1 TB0514 (our laboratory preparation) and mouse anti-HA 12CA5 (Sigma-Aldrich, St. Louis, MO) were used to detect Mmi1 and Rrp6-3HA, respectively. For immunoprecipitation with RNase treatment, cell lysates were incubated for 30 min at room temperature with 0.5 U of RNase A and 20 U of RNase T1 (RNase Cocktail Enzyme Mix, Thermo Fisher Scientific, Waltham, MA). In S2A Fig and S6C Fig, harvested cells were disrupted with glass beads in 20% trichloroacetic acid. Anti-GFP antibody [47] was used to detect Red1-YFP, Rrp6-YFP and Rrp6-YFP-Mmi1. Anti-γ-tubulin GTU-88 (Sigma-Aldrich) was used for a loading control.
in vitro binding assay
PCR-amplified YFP-mmi1 and rrp6-FLAG were cloned into pCold-GST (Takara). Glutathione S-transferase (GST)-tagged YFP-Mmi1 and Rrp6-FLAG proteins purified from E. coli were separated by NGC Chromatography System (Bio-rad, Hercules, CA) with HiTrap Heparin HP and HiTrap Q HP column (GE Healthcare) for GST-YFP-Mmi1 and GST-Rrp6-FLAG, respectively. It should be noted that most of GST-Rrp6-FLAG was cleaved between GST and Rrp6-FLAG during the preparation. HA-tagged Red1 was expressed in Rabbit Reticulocyte Lysate System, Nuclease Treated (Promega, Madison, WI) according to the manufacturer’s instruction. mRNA containing the His-tagged red1-3HA gene was transcribed using T7-Scribe Standard RNA IVT Kit (Cellscript, Madison, WI). Capping and poly(A)-tailing were performed using ScriptCap m7G Capping System (Cellscript), ScriptCap 2′-O-Methyltransferase Kit (Cellscript), and A-Plus Poly(A) Polymerase Tailing Kit (Cellscript). For negative controls, we parallelly performed in vitro translation without mRNA. Proteins were incubated at the room temperature for 30 min in the binding buffer (20 mM HEPES pH 7.5, 150 mM NaCl, 5 mM MgCl2, 1 mM dithiothreitol (DTT), 0.5% Triton-X100, 5% glycerol), and then mixed with Anti-DDDDK-tag mAb-Magnetic Beads M185-11 (MBL) at 4°C for 30 min. Beads were washed by wash buffer (20 mM HEPES pH 7.5, 300 mM NaCl, 5 mM MgCl2, 1 mM DTT, 1% Triton-X100) for 5 times, and subjected to western blotting. Rabbit anti-GFP ab6556 (abcam, Cambridge, UK), mouse anti-FLAG M2 (Sigma-Aldrich), and rat anti-HA 3F10 (Sigma-Aldrich) were used to detect GST-YFP-Mmi1, Rrp6-FLAG, and Red1-3HA, respectively. Images were acquired by Odyssey CLx Infrared Imaging System (LI-COR, Lincoln, NE).
Supporting information
(A) Localization of Rrp6, Red1 and Mmi1 in wild-type, red1Δ, mei4Δ, mmi1Δ mei4Δ, and pab2Δ cells. Cells expressing Rrp6-YFP (green), Red1-mCherry (red) or CFP-Mmi1 (blue) from the respective endogenous loci were observed during exponential growth in YE liquid medium. Dotted lines indicate the shape of cells. Boxed regions are magnified in Fig 1A. (B) Localization of Dis3 and Rrp4 in wild-type and red1Δ cells. red1Δ cells expressing Dis3-GFP or Rrp4-GFP from the respective endogenous loci were observed. Boxed regions are magnified in Fig 1B. (C) Localization of Pla1, Red1 and Mmi1 in wild-type and red1Δ cells. Cells expressing Pla1-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Boxed regions are magnified in Fig 1C. (D) Localization of Pab2, Red1, and Mmi1 in wild-type and red1Δ cells. Cells expressing Pab2-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Boxed regions are magnified in Fig 1D. Scale bars: 5 μm.
(PDF)
(A) Expression levels of truncated Red1 proteins. Cell extracts were prepared from exponentially growing cells expressing wild-type or truncated Red1-YFP in liquid YE medium and immunoblotted with anti-GFP antibody. γ-tubulin was used as a loading control. The asterisks indicate non-specific bands. (B) Localization of Rrp6 and Red1 in red1(Δ196–245) cells. red1(Δ196–245) cells expressing Rrp6-YFP (green) and Red1-mCherry (magenta) from the respective endogenous loci were observed. Dotted lines indicate the shape of cells. Boxed region is magnified in Fig 2C. Scale bar: 5 μm. (C) Localization of Rrp6, Red1 and Mmi1 in iss10Δ cells. iss10Δ cells expressing Rrp6-YFP (green), Red1-mCherry (red) and CFP-Mmi1 (blue) were examined. Boxed region is magnified in Fig 2D. Scale bar: 5 μm.
(PDF)
(A) Growth profiles of wild-type (red1-YFP), red1Δ, iss10Δ and red1(Δ196–245) cells. Ten-fold serial dilutions of cells were spotted on YE medium and incubated at the indicated temperatures. (B) Expression of rec8 mRNA and spo5 mRNA in wild-type (red1-YFP), red1Δ, iss10Δ and red1(Δ196–245) cells. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; ***P < 0.001 compared with the wild-type red1-YFP strain (Student’s t-test). (C) Expression of ssm4 mRNA, rec8 mRNA, spo5 mRNA, cti6 PROMPT and rpl402 PROMPT in wild-type, mmi1Δ mei4Δ, red1Δ and rrp6Δ cells. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01; ***P < 0.001 compared with the wild-type strain (Student’s t-test). (D) Expression of cti6 PROMPT and rpl402 PROMPT in wild-type (red1-YFP), red1Δ, red1(Δ196–245), and iss10Δ cells. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01; ***P < 0.001 compared with the wild-type red1-YFP strain (Student’s t-test).
(PDF)
(A) Co-immunoprecipitation of Rrp6 and Mmi1 in wild-type cells. Native cell extracts were prepared from exponentially growing cells expressing Rrp6-3HA from a plasmid in MM medium and subjected to immunoprecipitation with anti-HA antibody. Cells carrying an empty vector were used as a negative control. Precipitates and 10% total cell extracts were then immunoblotted with anti-Mmi1 and anti-HA antibodies. (B) The effect of RNase treatment on interaction between Rrp6 and Mmi1. Native cell extracts were incubated with or without RNase and subjected to immunoprecipitation with anti-HA antibody.
(PDF)
(A) Growth profiles of rrp6Δ cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids. Ten-fold serial dilutions of cells were spotted on MM medium and incubated at the indicated temperatures. (B) Expression of mei4 mRNA, ssm4 mRNA, and cti6 PROMPT in rrp6Δ cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. **P < 0.01; ***P < 0.001 compared with cells carrying empty vector (Student’s t-test). (C) Growth profiles of mmi1-ts3 cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids. Ten-fold serial dilutions of cells were spotted on MM medium and incubated at the indicated temperatures. (D) Expression of mei4 mRNA and ssm4 mRNA in mmi1-ts3 cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids. Cells were grown in liquid MM medium at 25°C and shifted to 37°C for 4 hours. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. **P < 0.01; ***P < 0.001 compared with cells carrying empty vector at 37˚C (Student’s t-test).
(PDF)
(A) Expression of rec8 mRNA, spo5 mRNA, and rpl402 PROMPT in red1Δ cells expressing Red1, Rrp6-GFP, Mmi1 or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium. Transcripts of each gene were analyzed by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01 compared with cells carrying empty vector (Student’s t-test). (B) Growth profiles of red1Δ cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids. Ten-fold serial dilutions of cells were spotted on MM medium and incubated at the indicated temperatures. (C) Expression levels of chimeric Rrp6-YFP-Mmi1 proteins. Cell extracts were prepared from exponentially growing cells expressing Rrp6-YFP or chimeric proteins composed of Rrp6, YFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium and immunoblotted with anti-GFP antibody. γ-tubulin was used as a loading control. (D) Expression of rec8 mRNA, spo5 mRNA and rpl402 PROMPT in red1Δ mmi1Δ mei4Δ cells expressing Red1, Rrp6-GFP, Mmi1, or chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium. Transcripts were analyzed by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01; ***P < 0.001 compared with cells carrying empty vector (Student’s t-test). (E) Co-immunoprecipitation of endogenous Mmi1 and chimeric Rrp6-GFP-Mmi1 proteins. Native cell extracts were prepared from exponentially growing cells expressing chimeric proteins composed of Rrp6, GFP, and full-length or truncated Mmi1 from plasmids in liquid MM medium and subjected to immunoprecipitation with anti-GFP antibody. Precipitates and 10% total cell extracts were then immunoblotted with anti-Mmi1 and anti-GFP antibodies.
(PDF)
(A) Localization of Dis3 and Rrp4 in rrp6Δ cells. rrp6Δ cells expressing Dis3-GFP or Rrp4-GFP from the respective endogenous loci were observed during exponential growth in liquid YE medium. Dotted lines indicate the shape of cells. Boxed regions are magnified in Fig 5A. (B) Localization of Red1 and Mmi1 in rrp6Δ cells. rrp6Δ cells expressing Red1-YFP (green) and CFP-Mmi1 (magenta) were examined. Boxed region is magnified in Fig 5B. (C) Localization of Dis3 and Rrp4 in red1Δ cells expressing Rrp6-YFP, YFP-Mmi1, or chimeric proteins composed of Rrp6, YFP, and full-length or truncated Mmi1. Dis3-mCherry and Rrp4-mCherry were expressed from the respective endogenous loci (green) and YFP-containing chimeric proteins were expressed from plasmids (magenta) in liquid SD medium. Boxed regions are magnified in Fig 5C. Scale bars: 5 μm.
(PDF)
Expression of rec8 mRNA and spo5 mRNA in mtl1-cs5 cells expressing Mtl1 and Rrp6-GFP-Mmi1 from plasmids. Cells were grown in liquid MM medium at 30°C and shifted to 20°C for 2 hours. Transcripts were quantified by RT-qPCR and normalized to act1. Error bars represent standard error of three independent samples. *P < 0.05; **P < 0.01 compared with cells carrying empty vector at 20˚C (Student’s t-test).
(PDF)
(PDF)
(PDF)
(XLSX)
Acknowledgments
We thank H. Kato, T. Urano and J. Nakayama for providing the anti-GFP antibody, S. Grewal and the National BioResource Project Japan for fission yeast strains, A. Nakade for technical support, and S. Iwasaki for helpful support.
Data Availability
All relevant data are within the manuscript and its Supporting Information files.
Funding Statement
This work was supported by JSPS (https://www.jsps.go.jp/english/index.html) KAKENHI Grant Number 15H04333 to AY, 19K06649 to YO and a grant from the Naito Foundation (https://www.naito-f.or.jp/en/) to YO. YS is a recipient of a JSPS Research Fellowship (PD) 19J00920. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Bresson S, Tollervey D. Surveillance-ready transcription: nuclear RNA decay as a default fate. Open biology. 2018;8(3): 170270 10.1098/rsob.170270 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Schmid M, Jensen TH. Controlling nuclear RNA levels. Nature reviews Genetics. 2018;19(8):518–29. 10.1038/s41576-018-0013-2 [DOI] [PubMed] [Google Scholar]
- 3.Harigaya Y, Tanaka H, Yamanaka S, Tanaka K, Watanabe Y, Tsutsumi C, et al. Selective elimination of messenger RNA prevents an incidence of untimely meiosis. Nature. 2006;442(7098):45–50. 10.1038/nature04881 [DOI] [PubMed] [Google Scholar]
- 4.Folco HD, Chalamcharla VR, Sugiyama T, Thillainadesan G, Zofall M, Balachandran V, et al. Untimely expression of gametogenic genes in vegetative cells causes uniparental disomy. Nature. 2017;543(7643):126–30. 10.1038/nature21372 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Hiriart E, Vavasseur A, Touat-Todeschini L, Yamashita A, Gilquin B, Lambert E, et al. Mmi1 RNA surveillance machinery directs RNAi complex RITS to specific meiotic genes in fission yeast. EMBO J. 2012;31(10):2296–308. 10.1038/emboj.2012.105 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Yamashita A, Shichino Y, Tanaka H, Hiriart E, Touat-Todeschini L, Vavasseur A, et al. Hexanucleotide motifs mediate recruitment of the RNA elimination machinery to silent meiotic genes. Open biology. 2012;2(3):120014 10.1098/rsob.120014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kilchert C, Wittmann S, Passoni M, Shah S, Granneman S, Vasiljeva L. Regulation of mRNA Levels by Decay-Promoting Introns that Recruit the Exosome Specificity Factor Mmi1. Cell reports. 2015;13(11):2504–15. 10.1016/j.celrep.2015.11.026 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Chatterjee D, Sanchez AM, Goldgur Y, Shuman S, Schwer B. Transcription of lncRNA prt, clustered prt RNA sites for Mmi1 binding, and RNA polymerase II CTD phospho-sites govern the repression of pho1 gene expression under phosphate-replete conditions in fission yeast. RNA. 2016;22(7):1011–25. 10.1261/rna.056515.116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wang C, Zhu Y, Bao H, Jiang Y, Xu C, Wu J, et al. A novel RNA-binding mode of the YTH domain reveals the mechanism for recognition of determinant of selective removal by Mmi1. Nucleic Acids Res. 2016;44(2):969–82. 10.1093/nar/gkv1382 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Wu B, Xu J, Su S, Liu H, Gan J, Ma J. Structural insights into the specific recognition of DSR by the YTH domain containing protein Mmi1. Biochemical and biophysical research communications. 2017;491(2):310–6. 10.1016/j.bbrc.2017.07.104 [DOI] [PubMed] [Google Scholar]
- 11.Hazra D, Chapat C, Graille M. m(6)A mRNA Destiny: Chained to the rhYTHm by the YTH-Containing Proteins. Genes. 2019;10(1):E49 10.3390/genes10010049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Yamanaka S, Yamashita A, Harigaya Y, Iwata R, Yamamoto M. Importance of polyadenylation in the selective elimination of meiotic mRNAs in growing S. pombe cells. EMBO J. 2010;29(13):2173–81. 10.1038/emboj.2010.108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Ogami K, Chen Y, Manley JL. RNA surveillance by the nuclear RNA exosome: mechanisms and significance. Non-coding RNA. 2018;4(1). 10.3390/ncrna4010008 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zinder JC, Lima CD. Targeting RNA for processing or destruction by the eukaryotic RNA exosome and its cofactors. Genes & development. 2017;31(2):88–100. 10.1101/gad.294769.116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kilchert C, Wittmann S, Vasiljeva L. The regulation and functions of the nuclear RNA exosome complex. Nat Rev Mol Cell Biol. 2016;17(4):227–39. 10.1038/nrm.2015.15 [DOI] [PubMed] [Google Scholar]
- 16.Chekanova JA, Gregory BD, Reverdatto SV, Chen H, Kumar R, Hooker T, et al. Genome-wide high-resolution mapping of exosome substrates reveals hidden features in the Arabidopsis transcriptome. Cell. 2007;131(7):1340–53. 10.1016/j.cell.2007.10.056 [DOI] [PubMed] [Google Scholar]
- 17.Gudipati RK, Xu Z, Lebreton A, Seraphin B, Steinmetz LM, Jacquier A, et al. Extensive degradation of RNA precursors by the exosome in wild-type cells. Mol Cell. 2012;48(3):409–21. 10.1016/j.molcel.2012.08.018 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Schneider C, Kudla G, Wlotzka W, Tuck A, Tollervey D. Transcriptome-wide analysis of exosome targets. Mol Cell. 2012;48(3):422–33. 10.1016/j.molcel.2012.08.013 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen HM, Futcher B, Leatherwood J. The fission yeast RNA binding protein Mmi1 regulates meiotic genes by controlling intron specific splicing and polyadenylation coupled RNA turnover. PloS one. 2011;6(10):e26804 10.1371/journal.pone.0026804 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.St-Andre O, Lemieux C, Perreault A, Lackner DH, Bahler J, Bachand F. Negative regulation of meiotic gene expression by the nuclear poly(a)-binding protein in fission yeast. J Biol Chem. 2010;285(36):27859–68. 10.1074/jbc.M110.150748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Sugiyama T, Sugioka-Sugiyama R. Red1 promotes the elimination of meiosis-specific mRNAs in vegetatively growing fission yeast. EMBO J. 2011;30(6):1027–39. 10.1038/emboj.2011.32 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yamashita A, Takayama T, Iwata R, Yamamoto M. A novel factor Iss10 regulates Mmi1-mediated selective elimination of meiotic transcripts. Nucleic Acids Res. 2013;41(21):9680–7. 10.1093/nar/gkt763 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Lee NN, Chalamcharla VR, Reyes-Turcu F, Mehta S, Zofall M, Balachandran V, et al. Mtr4-like protein coordinates nuclear RNA processing for heterochromatin assembly and for telomere maintenance. Cell. 2013;155(5):1061–74. 10.1016/j.cell.2013.10.027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Egan ED, Braun CR, Gygi SP, Moazed D. Post-transcriptional regulation of meiotic genes by a nuclear RNA silencing complex. RNA. 2014;20(6):867–81. 10.1261/rna.044479.114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Zhou Y, Zhu J, Schermann G, Ohle C, Bendrin K, Sugioka-Sugiyama R, et al. The fission yeast MTREC complex targets CUTs and unspliced pre-mRNAs to the nuclear exosome. Nature communications. 2015;6:7050 10.1038/ncomms8050 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Tashiro S, Asano T, Kanoh J, Ishikawa F. Transcription-induced chromatin association of RNA surveillance factors mediates facultative heterochromatin formation in fission yeast. Genes Cells. 2013;18(4):327–39. 10.1111/gtc.12038 [DOI] [PubMed] [Google Scholar]
- 27.Zofall M, Yamanaka S, Reyes-Turcu FE, Zhang K, Rubin C, Grewal SI. RNA elimination machinery targeting meiotic mRNAs promotes facultative heterochromatin formation. Science. 2012;335(6064):96–100. 10.1126/science.1211651 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Chalamcharla VR, Folco HD, Dhakshnamoorthy J, Grewal SI. Conserved factor Dhp1/Rat1/Xrn2 triggers premature transcription termination and nucleates heterochromatin to promote gene silencing. Proc Natl Acad Sci U S A. 2015;112(51):15548–55. 10.1073/pnas.1522127112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shah S, Wittmann S, Kilchert C, Vasiljeva L. lncRNA recruits RNAi and the exosome to dynamically regulate pho1 expression in response to phosphate levels in fission yeast. Genes & development. 2014;28(3):231–44. 10.1101/gad.230177.113 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Touat-Todeschini L, Shichino Y, Dangin M, Thierry-Mieg N, Gilquin B, Hiriart E, et al. Selective termination of lncRNA transcription promotes heterochromatin silencing and cell differentiation. EMBO J. 2017;36(17):2626–41. 10.15252/embj.201796571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Sugiyama T, Wanatabe N, Kitahata E, Tani T, Sugioka-Sugiyama R. Red5 and three nuclear pore components are essential for efficient suppression of specific mRNAs during vegetative growth of fission yeast. Nucleic Acids Res. 2013;41(13):6674–86. 10.1093/nar/gkt363 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Sugiyama T, Thillainadesan G, Chalamcharla VR, Meng Z, Balachandran V, Dhakshnamoorthy J, et al. Enhancer of Rudimentary Cooperates with Conserved RNA-Processing Factors to Promote Meiotic mRNA Decay and Facultative Heterochromatin Assembly. Mol Cell. 2016;61(5):747–59. 10.1016/j.molcel.2016.01.029 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Shichino Y, Otsubo Y, Kimori Y, Yamamoto M, Yamashita A. YTH-RNA-binding protein prevents deleterious expression of meiotic proteins by tethering their mRNAs to nuclear foci. eLife. 2018;7:e32155 10.7554/eLife.32155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Xie G, Vo TV, Thillainadesan G, Holla S, Zhang B, Jiang Y, et al. A conserved dimer interface connects ERH and YTH family proteins to promote gene silencing. Nature communications. 2019;10(1):251 10.1038/s41467-018-08273-9 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Cole C, Barber JD, Barton GJ. The Jpred 3 secondary structure prediction server. Nucleic Acids Res. 2008;36(Web Server issue):W197–201. 10.1093/nar/gkn238 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Fox MJ, Mosley AL. Rrp6: Integrated roles in nuclear RNA metabolism and transcription termination. Wiley interdisciplinary reviews RNA. 2016;7(1):91–104. 10.1002/wrna.1317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Shichino Y, Yamashita A, Yamamoto M. Meiotic long non-coding meiRNA accumulates as a dot at its genetic locus facilitated by Mmi1 and plays as a decoy to lure Mmi1. Open biology. 2014;4(6):140022 10.1098/rsob.140022 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Meola N, Domanski M, Karadoulama E, Chen Y, Gentil C, Pultz D, et al. Identification of a Nuclear Exosome Decay Pathway for Processed Transcripts. Mol Cell. 2016;64(3):520–33. 10.1016/j.molcel.2016.09.025 [DOI] [PubMed] [Google Scholar]
- 39.Ogami K, Richard P, Chen Y, Hoque M, Li W, Moresco JJ, et al. An Mtr4/ZFC3H1 complex facilitates turnover of unstable nuclear RNAs to prevent their cytoplasmic transport and global translational repression. Genes & development. 2017;31(12):1257–71. 10.1101/gad.302604.117 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Silla T, Karadoulama E, Makosa D, Lubas M, Jensen TH. The RNA Exosome Adaptor ZFC3H1 Functionally Competes with Nuclear Export Activity to Retain Target Transcripts. Cell reports. 2018;23(7):2199–210. 10.1016/j.celrep.2018.04.061 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Gutz H, Heslot H, Leupold U, Loprieno N. Schizosaccharomyces pombe. In Handbook of Genetics. King RD, editor. New York: Plenum Publishing Corporation; 1974. 395–446. [Google Scholar]
- 42.Moreno S, Klar A, Nurse P. Molecular genetic analysis of fission yeast Schizosaccharomyces pombe. Methods Enzymol. 1991;194:795–823. 10.1016/0076-6879(91)94059-l [DOI] [PubMed] [Google Scholar]
- 43.Bahler J, Wu JQ, Longtine MS, Shah NG, McKenzie A 3rd, Steever AB, et al. Heterologous modules for efficient and versatile PCR-based gene targeting in Schizosaccharomyces pombe. Yeast. 1998;14(10):943–51. [DOI] [PubMed] [Google Scholar]
- 44.Sato M, Dhut S, Toda T. New drug-resistant cassettes for gene disruption and epitope tagging in Schizosaccharomyces pombe. Yeast. 2005;22(7):583–91. 10.1002/yea.1233 [DOI] [PubMed] [Google Scholar]
- 45.Basi G, Schmid E, Maundrell K. TATA box mutations in the Schizosaccharomyces pombe nmt1 promoter affect transcription efficiency but not the transcription start point or thiamine repressibility. Gene. 1993;123:131–6. 10.1016/0378-1119(93)90552-e [DOI] [PubMed] [Google Scholar]
- 46.Matsuo T, Otsubo Y, Urano J, Tamanoi F, Yamamoto M. Loss of the TOR kinase Tor2 mimics nitrogen starvation and activates the sexual development pathway in fission yeast. Mol Cell Biol. 2007;27(8):3154–64. 10.1128/MCB.01039-06 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Kato H, Okazaki K, Iida T, Nakayama J, Murakami Y, Urano T. Spt6 prevents transcription-coupled loss of posttranslationally modified histone H3. Scientific reports. 2013;3:2186 10.1038/srep02186 [DOI] [PMC free article] [PubMed] [Google Scholar]