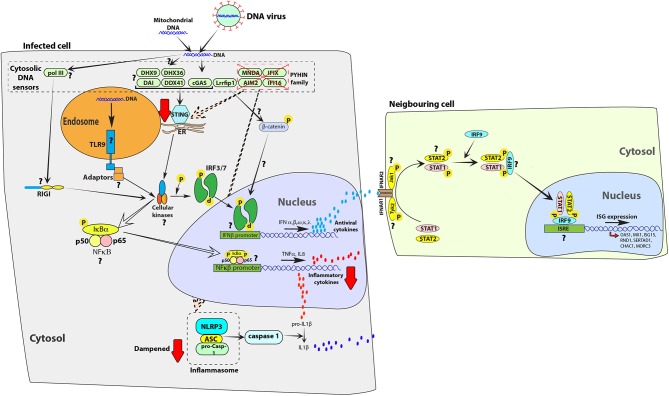
Figure 2.
Exogenous and self-DNA sensing pathways are dampened in bat cells. In human cells, endosomal TLR9 (35) and cytosolic receptors from the PYHIN family (86) [absent in melanoma 2 (AIM2), IFN-inducible gene 16 (IFI16), myeloid cell nuclear differentiation antigen (MNDA) and IFN-inducible protein X (IFIX)], cyclic GMP-AMP synthase (cGAS) (87), DNA-dependent activator of IFN-regulatory factor (DAI) (88), RNA polymerase III (PolIII) (89), LRR binding FLII interacting protein 1 (Lrrfip1) (90), DDX41 (91), DExH-Box helicase 9 (DHX9) and DEAH-Box helicase 36 (DHX36) (92) can detect exogenous and self-DNA (86, 93). On binding to DNA, these receptors signal through adaptor proteins to activate cellular kinases, such as TANK-binding kinase 1 protein (TBK1), which in-turn activates transcription factors, such as interferon regulatory factor 3 (IRF3) or IRF7 and nuclear factor kappa-light-chain-enhancer of activated B cells (NFκB) to induce expression of antiviral interferons (IFNs) and pro-inflammatory cytokines, respectively. PYHIN family of cytosolic receptors signal through STING and the NLRP3 inflammasome. This family of receptors has been negatively selected for and lost in the genomic sequences of bats (94). A downstream signal mediator of cGAS, stimulator of IFN genes (STING) is less functional in bat cells, relative to human cells (95). The attenuated function of STING would likely extend to other DNA sensors that signal through STING, such as DDX41, DHX9, DHX36, and DAI. The presence and function of additional homologs of DNA sensors, such as TLR9, PolIII, and Lrrfip1 have not been characterized in bats. In the figure, red arrows indicate a dampened response in the pathway, relative to human cells. Question marks (?) highlight pathways and molecular homologs that have not yet been characterized or identified in bats. The data have been compiled from studies in different species and one finding may not represent a universal bat response. ER, endoplasmic reticulum.