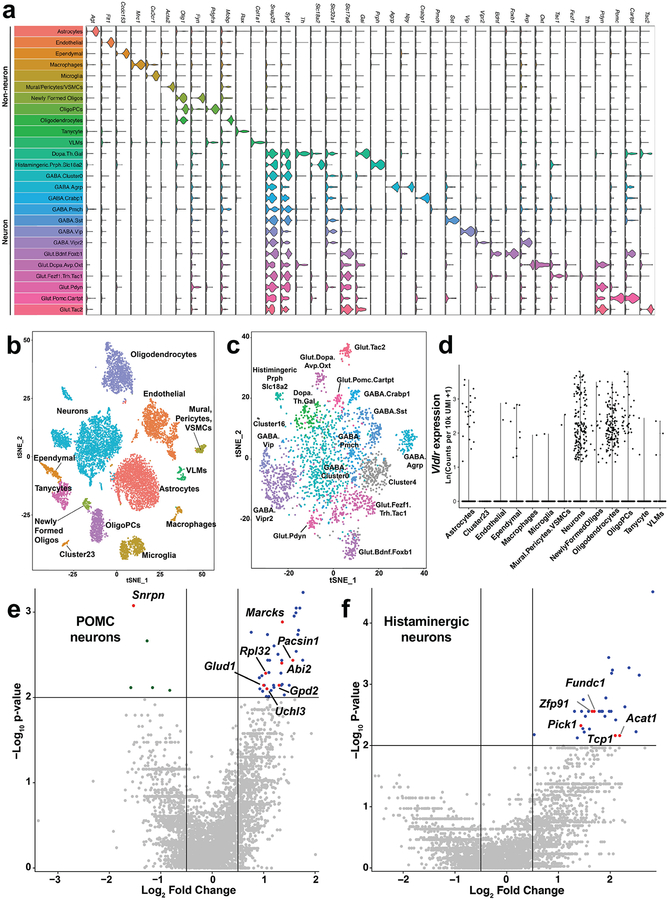
Extended Data Figure 6: Single cell RNA sequencing examination of the transcriptional landscape of the hypothalamus with Drop-seq.
Clustering analysis combined with expression profiling of a panel of marker genes allowed us to discriminate 26 unique clusters of cells in the hypothalamus. (a) Violin plots demonstrate the expression patterns of the 38 marker genes used to identify the cell clusters. Individual data points indicating the magnitude of gene expression in a single cell are superimposed on a probability density plot for the distribution of the data; the expression analysis is based on the data collected from n=11,453 single cells. (b) Global gene expression relationships in the 11,453 single cells isolated from the hypothalamic tissues of six mice projected onto two dimensions using t-distributed Stochastic Neighbor Embedding (tSNE). The clusters were defined using shared nearest neighbor graph-based clustering. (c) tSNE plot of the neuronal cells identified in the Drop-seq experiment (n=3369 single cells). (d) Violin plot demonstrating that Vldlr is only appreciably expressed in neuron and oligodendrocyte cell populations. Individual data points indicating the magnitude of gene expression in a single cell; the expression analysis is based on the unique molecular identities (UMI) data collected from n=11,453 single cells. (e-f) Volcano plots of the differentially expressed genes analyzed by two-sided Wilcoxon rank sum tests in (e) POMC+ (n=24 WT and n=26 Idol−/− cells) and (f) Histaminergic neurons (n=23 WT and n=11 Idol−/− neurons. Labeled genes are linked to whole body metabolic homeostasis – see Supplemental Data Table 2 for details.