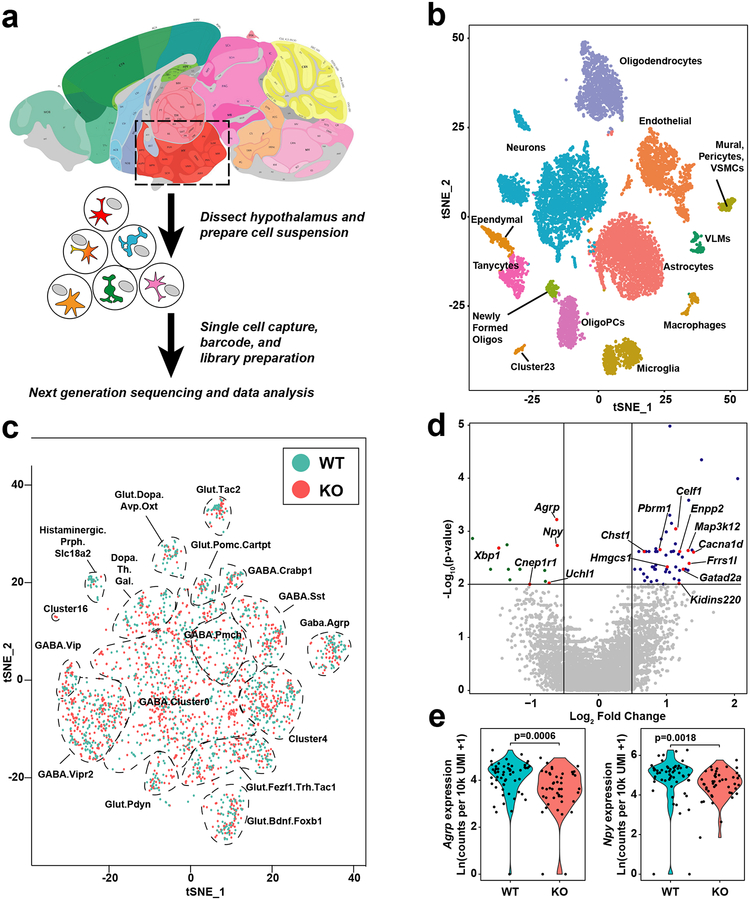
Figure 5: The single cell transcriptional landscape of the hypothalamus is affected by deletion of IDOL.
(a) Simplified schematic of the single cell RNA-sequencing experimental design. 5-week old mice were placed on a high fat, high cholesterol diet for two weeks. Hypothalamic tissues from six mice (n=3/group) were dissected and dissociated into single-cell suspension. Single cells and barcoded beads were captured into droplets followed by cDNA synthesis, amplification, and library preparation. The library was sequenced with an Illumina HiSeq 4000 next-generation instrument using the Drop-seq custom read 1B primer. (b) Global gene expression relationships in the 11,453 single cells projected onto two dimensions using t-distributed Stochastic Neighbor Embedding (tSNE). The clusters were defined using shared nearest neighbor graph-based clustering. (c) tSNE plot demonstrating the effect of IDOL knockout on the clustering of single cells in fifteen different neuron clusters; n=11,453 cells. (d) Volcano plot of the differentially expressed genes in AGRP neurons; expression data calculated as counts per 10k unique molecular identities (UMI); the p-values were calculated using a two-tailed Wilcoxon rank sum tests from n=56 WT and n=45 Idol−/− neurons identified within the AGRP cluster. Labeled genes are linked to whole-body metabolic homeostasis – see Extended Data Table 1 for details). (e) Violin plots demonstrating that IDOL deletion reduced gene expression of Agrp and Npy in AGRP neurons. Individual data points indicate the magnitude of gene expression in a single cell. These are superimposed on a probability density plot for the distribution of the data; statistics were calculated using two-tailed Wilcoxon rank sum tests on non-Ln transformed data; n=56 WT and n=45 IDOL KO neurons identified within the AGRP cluster.