Figure 7.
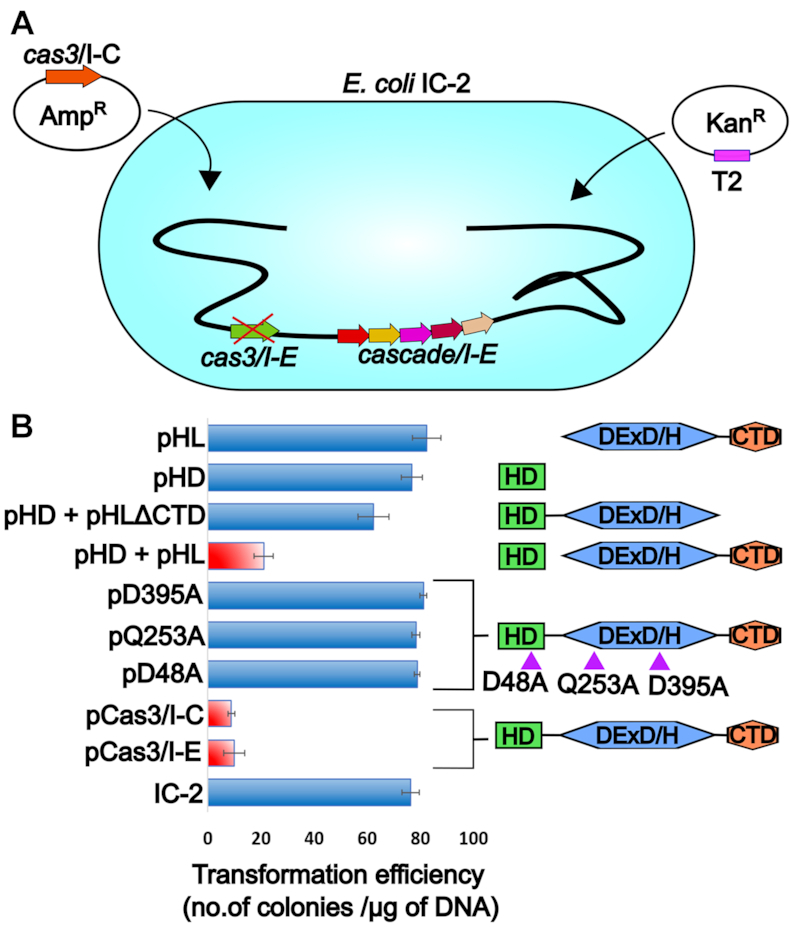
Cas3/I-C supplants Cas3/I-E in type I-E CRISPR-Cas system (A) E. coli K-12 harboring Cas3/I-E null mutant (IC-2) was used. IC-2 harbors spacer sequence targeting plasmid pRSF-11 (pT2). Cas3/I-C was heterologously expressed through IPTG inducible vector having ampicillin selection marker. Transformation efficiency was calculated after induction with 20 μM IPTG and 12 h incubation. (B) Cas3/I-C and its variants were tested for interference activity using E. coli IC-2. Heterologous expression of Cas3/I-C (pCas3/I-C) and Cas3/I-E (pCas3/I-E) showed reduced transformation efficiency (indicated as a red bar). When the HD (pHD) and helicase (pHL) domains of Cas3/I-C were co-expressed as two separate constructs, the transformation efficiency mirrored that of wild-type (indicated as a red bar). However, HD nuclease mutant (pD48A), helicase mutants (pQ253A and pD395A) and co-expressed constructs of HD and helicase without CTD (pHD + pHLΔCTD), stand-alone nuclease domain (pHD) and stand-alone helicase domain (pHL) showed high transformation efficiency. Error bar represents standard deviation measured from three independent trials. Purple triangles represent the location of the point mutations. The boundary of the domain variants are as follows: HD domain (pHD) 1-248; DExD/H domain (pHL) 249-800; pHD+pHLΔCTD 1–709.
