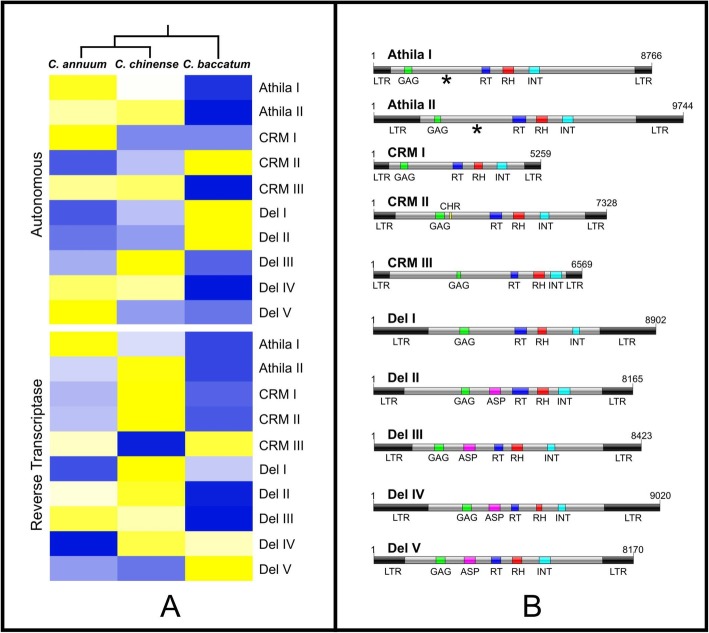
Fig. 3.
Comparative distribution of the putative autonomous LTR-RTs and the reverse transcriptase sequences along the C. annuum, C. chinense, and C. baccatum datasets. a The Annuum clade, composed by C. annuum and C. chinense and the Baccatum clade (C. baccatum) can be distinguished by the dendrogram on top of the heatmap. In the heatmap, lower and higher accumulation (blue, intermediate and yellow, respectively) represent the amount of conserved sequences found in each dataset. Image shows that Athila/Tat elements accumulated more in C. annuum and C. chinense (marked in yellow and light-yellow), the CRM groups were differentially accumulated, highlighting the absence of CRM III and the predominance of Del I and II in C. baccatum, and bigger accumulation of Del III, IV, and V in C. annuum and C. chinense. The reverse transcriptase of these elements exhibits a similar pattern of distribution than the one observed for the complete elements, Athila/Tat I and II were more accumulate in C. annuum and C. chinense, respectively. CRM I and II were more accumulated in C. chinense, while the group CRM III was more accumulated in C. baccatum. Capsicum annuum and C. chinense had a bigger accumulation of Del I, II and II than C. baccatum, while the groups Del III and IV exhibited more accumulation in C. baccatum. b Graphical representation of LTR-RTs groups. LTR – long terminal repeat, GAG – nucleocapsid, RT – reverse transcriptase, RH – RNAse H, INT – integrase, ASP – aspartase. Asterisks present in Athila illustrations refer to the hallmark ORF for this lineage. Note a difference in extension (bp length) among elements, including GAG and POL positioning, and LTR sizes. Note also that only CRM II exhibits a chromodomain sequence and that all the Del elements present an additional aspartase locus