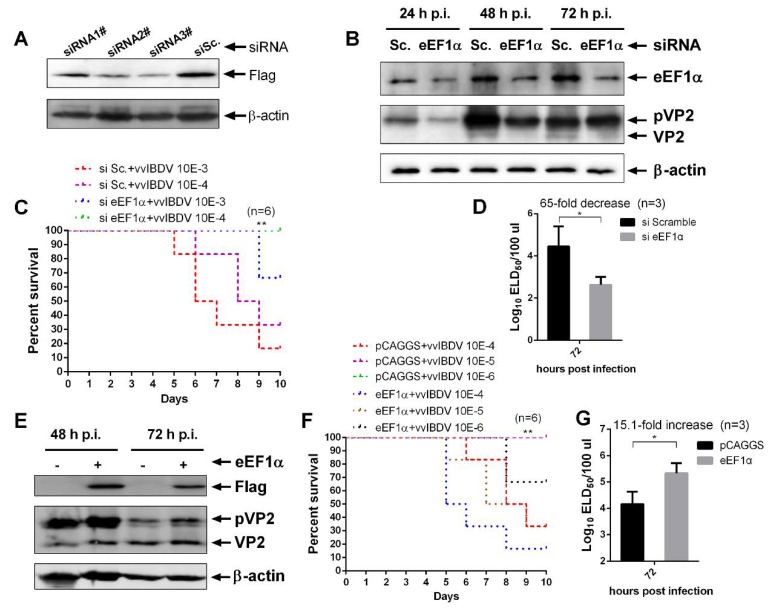
Figure 2.
Chicken eukaryotic translation elongation factor 1α (cheEF1α) promoted the replication of vvIBDV. (A) Effects of siRNAs on the cheEF1α expression. 293T cells were transfected with Flag-tagged cheEF1α accompanied with siRNAs (RNAi#1, RNAi#2, or RNAi#3) or scrambled siRNA. Cell lysates were harvested 48 h after transfection and tested by Western blotting with anti-Flag antibody. Endogenous β-actin expression was used as an internal control. (B) Yields of viral proteins in cheEF1α knockdown cells. Viral proteins loads were analyzed by Western blotting at 24 h, 48 h, and 72 h post-infection (p.i.) with vvIBDV Gx at multiplicity of infection (MOI) = 1 compared the potent cheEF1α siRNA treatment to scrambled siRNA treatment. The amount of cheEF1α indicated the efficiency of siRNA silencing and β-actin was used as the loading control. (C) Typical survival curve of chicken embryos infected with extracellular progeny virus in cheEF1α knockdown cells. Progeny viruses in extracellular of cells with or without cheEF1α knockdown were harvested at 72 h p.i., and chicken embryos were infected with 10-fold diluted progeny viruses, with the number of deaths for 9 days post-infection being recorded. One of three independent tests was shown as survival curve, and these three independent tests of 50% embryo lethal dose (ELD50) were summarized in (D). (E) Yields of viral proteins in cheEF1α over-expression cells. Viral protein loads were analyzed by Western blotting at 48 h and 72 h p.i. with vvIBDV Gx at MOI = 1 with or without cheEF1α over-expression. (F) Typical survival curve of chicken embryos infected with extracellular progeny virus in cheEF1α over-expression cells. The progeny virus titers based on 50% embryo lethal dose (ELD50) are summarized in (G). Error bars and mean ± SD were calculated on the basis of three independent experiments. The statistical analysis of survival curve was analyzed with GraphPad Prism 6 software, and the other statistical analysis was performed with t-tests considered significant with one asterisk as p < 0.05 and two asterisks as p < 0.01.