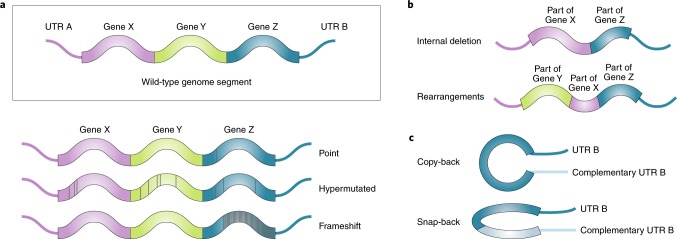
Fig. 1. Classes of DVGs.
a, Mutations in the viral genomes can lead to the generation of DVGs. These mutations can be single point mutations, hypermutations or frameshift mutations that alter virus replication, viral protein expression and/or viral protein function. b, Deletion-type DVGs occur when the polymerase skips part of the genome during replication and generates a truncated version of the genome. Deletion DVGs could also involve genomic rearrangements and gene duplication. Deletion generally results in lost or altered expression of one or more genes. c, Snap-back and copy-back DVGs are generated in nsRNA viruses when a sequence is duplicated in reverse complement to create theoretical panhandle structures for copy-back DVGs or hairpin structures for snap-back DVGs. Complementary sequence duplication occurs when the polymerase is released from the template strand and reattaches back to the nascent strand, copying back the end of the nascent genome. They can also occur from the polymerase copying back another complementary genome bound in trans. Most described snap-back and copy-back DVGs are generated from the 5’ end of the genome with the complementary end containing a duplication of the unstranslated region (UTR). Various degrees of complementation within Gene Z in the illustrated model can be found in copy-back DVGs. These types of genomes are not transcribed but can be replicated by the viral polymerase.