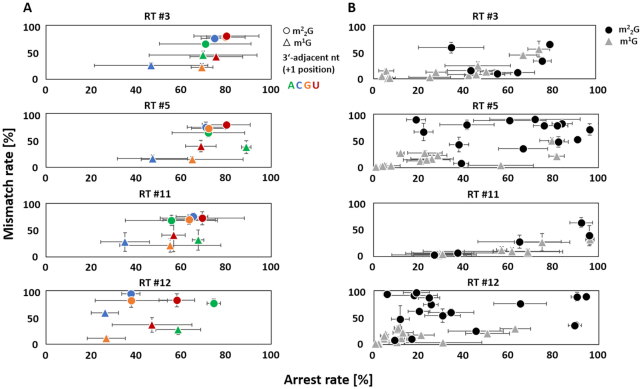
Figure 5.
m1G and m22G RT-signature comparison for the analysed reverse transcriptases RT #3, #5, #11 and #12. (A) Revolver oligo. Dot plots of mismatch and arrest RT-signatures at m22G (dots) and m1G (triangles) sites in revolver oligos for base configurations guanosine (orange), cytidine (blue), uridine (red, T in mapping profile) and adenosine (green) at position +1. Mismatch and arrest rates are given in percentage. Data was averaged from triplicates; error bars show standard deviations of arrest and mismatch rates. (B) Total tRNA from Saccharomyces cerevisiae. Dot plots of mismatch and arrest RT-signatures at m22G (black dots) and m1G (gray triangles) sites in total tRNA. Data was averaged from triplicates; error bars show standard deviations of arrest and mismatch rates. m1G and m22G sites which are present in all three total tRNA replicates and show a coverage of at least 20 reads in at least two replicates are shown (see Supplement Table S8 for more details). In general, arrest rate percentages refer to the reads covering the 3′ adjacent position of m1G/m22G (+1 position). Mismatch rate percentages refer to the reads covering the modified position.