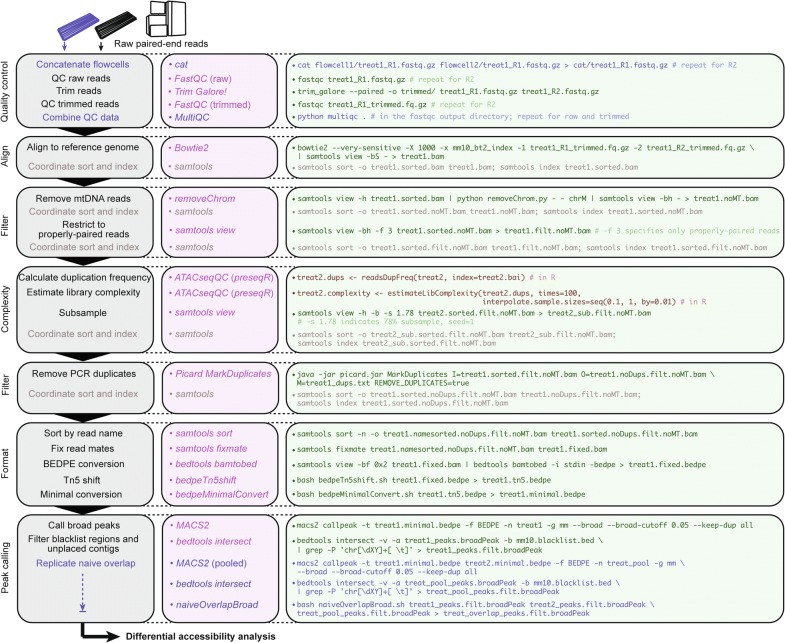
Fig. 4.
Generalized ATAC-seq data processing workflow intended for comparative analysis. Stepwise bioinformatics process and example commands for analyzing ATAC-seq data from raw reads to calling peaks for downstream differential accessibility analysis. Consider “treat1” as an example mouse ATAC-seq Illumina paired-end library. Blue text denotes optional or conditional steps dependent on experimental design and desired output. Users seeking only to discover replicate-concordant accessible regions in a singular cell state may wish to call naïve overlapping peaks, though this step is not necessary for differential accessibility analysis. Bash scripts for Tn5 coordinate shift (bedpeTn5shift.sh), minimal BEDPE format conversion (bedpeMinimalConvert.sh), and calling naïve overlap broad peaks (naiveOverlapBroad.sh) are located in the additional files section along with a machine-readable text version of this workflow