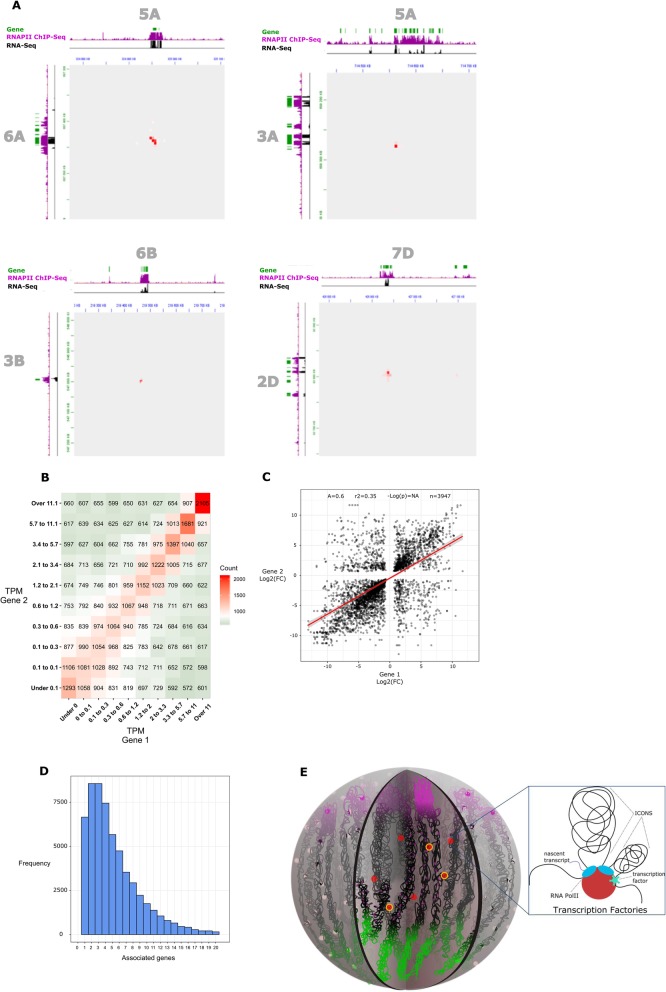
Fig. 6.
Gene interchromosomal interaction through transcription factories is associated with coregulation. a Integration of RNAPII Hi-ChIP, RNAPII ChIP-seq, and RNA-seq data. Examples of 2D heatmap with the frequency of interchromosomal interactions are presented. Green bars represent genes. b Heatmap presenting the gene expression level of gene pairs involved in RNAPII-associated interactions. Within each gene pair, each gene was classified into a decile of expression, and then the number of gene pairs with each combination of deciles was counted. c Scatterplot of log2(shoot/root fold change) for pairs of interacting genes associated through interchromosomal RNAPII-associated contacts. Interactions were identified by an RNAPII Hi-ChIP experiment (see the “Results” section). Only interchromosomal contacts conserved in shoots and roots were used in the analysis. d Histogram showing the distribution of genes interacting through RNAPII-associated contacts by numbers of partner genes. e Model of a transcription factory. Each factory could contain several RNA polymerase II molecules. These transcription factories could include many factors involved in transcription, such as coactivators, chromatin remodelers, transcription factors, histone modification enzymes, RNPs, RNA helicases, and splicing and processing factors. Multiple genes can be processed by the same factory (two are shown)