Fig. 5. CRISPR-based editing enables FOXP3 gene correction in IPEX patient cells.
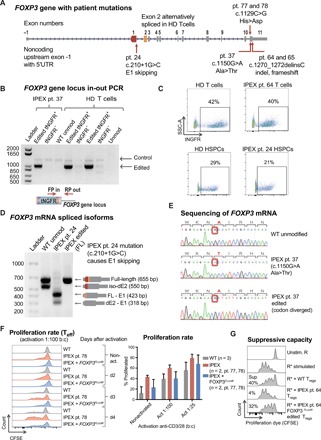
(A) Schematic of FOXP3 gene highlighting mutations of patients involved in this study. (B) Editing of IPEX and HD T cells observed at the DNA level by in-out PCR strategy. Forward primer is in the tNGFR cassette, and the reverse primer is in the FOXP3 gene locus outside the 3′ arm of homology. Positive and negative fractions after tNGFR enrichment (+/−) analyzed by PCR. (C) Flow cytometry plots of tNGFR staining. (D) Expression of FOXP3 mRNA demonstrated by RT-PCR in activated IPEX and HD T cells (schematic mRNA isoforms are shown to the right of corresponding PCR product bands). After ladder, first lane represents WT cells with two naturally occurring alternatively spliced isoforms, FL and dE2. The second lane shows aberrant skipping of E1 in IPEX pt24 mRNA that results in truncated transcripts. The third lane shows edited IPEX cells with restoration of the FL mRNA. (E) Sanger sequencing showing c.1150G>A mutation in the mRNA of pt37, resulting in an Ala>Thr change, and CRISPR-based insertion of divergent FOXP3 cDNA restoring correct amino acid sequence. (F) Proliferation of Teff cells in response to activation measured by the proliferation assay, comparing HD WT cells and IPEX pt78 cells. Quantification of average proliferation response of Teff cells from proliferation assay at day 3. (G) Functional testing of gene edited IPEX Tregs using the in vitro suppression assay. Flow cytometry plots of CFSE-stained Teff responder cells (R) cultured with or without Tregs. Calculated suppressive potential shows diminished suppressive function of IPEX pt. 64 Tregs, which was partially restored by FOXP3 gene editing.
