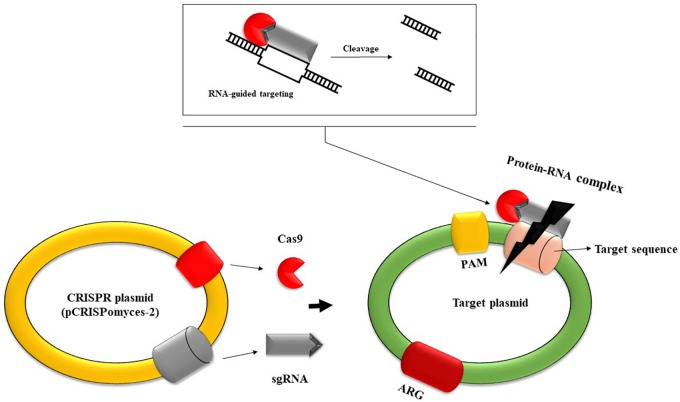
FIGURE 3.
Schematic representation of CRISPR-based plasmid system capable of removing MGE-like resistance plasmids. This system contains two sgRNA transcripts, the cas9 nuclease, and other structural elements. Firstly, sgRNA forms a complex with cas nuclease. The sgRNA transcripts guide cas9 nuclease to introduce double-stranded breaks at the ends of the target DNA, leading to cleavage. Direct target recognition is achieved through recognition of protospacer adjacent motifs (PAM), short DNA sequences that are not found in CRISPR loci, so there is no risk of self-degradation (So et al., 2017). Subsequently, the gap is filled through homologous recombination by an editing template. This system can be used to edit the genome of several antibiotic-resistant bacterial strains, leading to the removal of resistance determinants.