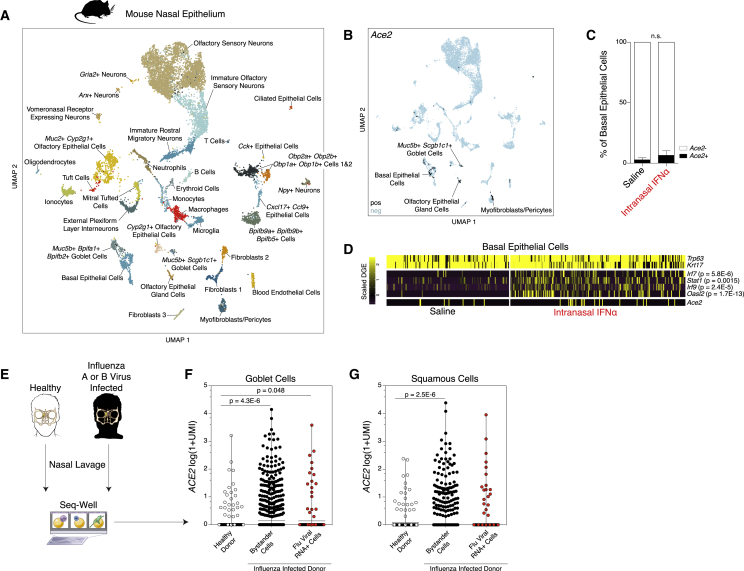
Figure 6.
In Vivo Administration of Interferons in Mice Does Not Induce Ace2, and ACE2 Is Induced in Goblet Secretory Cells during Human Influenza Infection
(A) UMAP of 11,358 single cells from mouse nasal epithelium (n = 4).
(B) UMAP projection as in (A), points colored by detection of Ace2 (SARS-CoV-2 receptor homolog). Color coding is as follows: black, RNA positive; blue, RNA negative.
(C) Percent of Ace2+ cells by treatment condition (n = 4 arrays per condition; n = 2 arrays per mouse). Black bars indicate Ace2+ cells; white bars indicate Ace2− cells. p = 0.4 by Student’s t test.
(D) Heatmap of cell-type-defining genes (Trp63 and Krt17), interferon-induced genes (Irf7, Stat1, Irf9, and Oasl2), and Ace2 among basal epithelial cells, separated by cells derived from saline-treated mice (left) and IFN-α-treated mice (right). Statistical significance by likelihood-ratio test with Bonferroni correction is shown. A full list of differentially expressed genes can be found in Table S8.
(E) Schematic for sampling cells derived from nasal washes of n = 18 human donors with and without current influenza A or B infection for Seq-Well v1 (35,840 single cells). See Cao et al., (2020).
(F and G) ACE2 expression among goblet cells (F) and squamous cells (G) by infection status. Shown are Healthy Donor cells from influenza-negative donors (white); Bystander Cells from influenza A (IAV)- or influenza B (IBV)-infected donors, no intracellular viral RNA detected (black); Flu Viral RNA+ Cells with detectable intracellular influenza A or B viral RNA (red). Statistical significance by Wilcoxon test with Bonferroni correction, n.s. for Bystander versus Flu Viral RNA+.