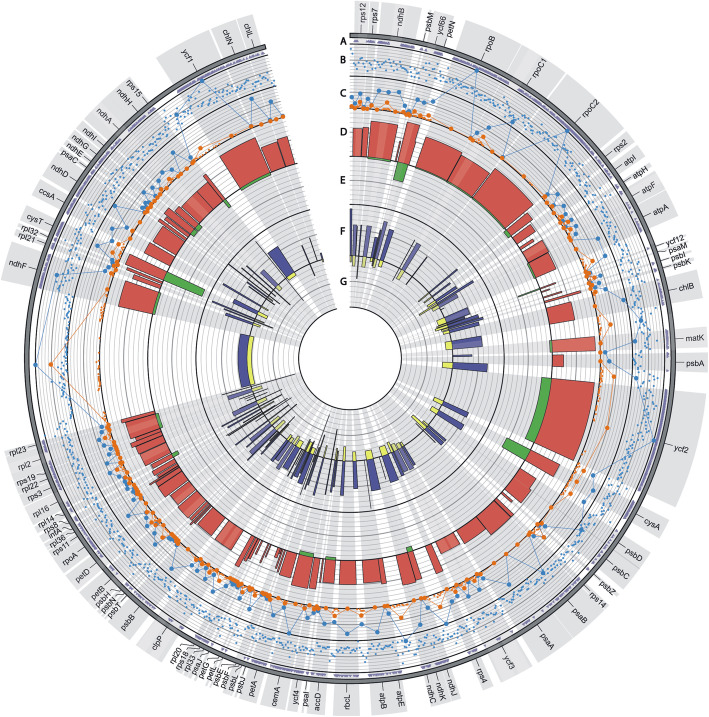
Fig. 2.
SNP and indel variation among plastomes of Calypogeia. Track A shows nonsynonymous SNP occurrence within genes. Track B and C represent identified SNPs (small blue dots) and indels (small orange dots) per 100 bp window size (maximum value = 40). Line plot, comprising B and C track, represents SNPs (blue line) and indels (orange line) within each exon, intron or intergenic spacer (snp max. Value = 400, indel max. Value = 100). Track D represents percent of SNPs per CDS length (maximum value = 22) while track E represents percent of indels per CDS length (maximum value = 2). Track F represents percent of SNPs per noncoding region length (max. Value = 30) while track G represents percent of indels per noncoding region length (max value = 20)