Abstract
Cancer-associated fibroblasts (CAFs) are a prominent stromal cell type in solid tumors and molecules secreted by CAFs play an important role in tumor progression and metastasis. CAFs coexist as heterogeneous populations with potentially different biological functions. Although CAFs are a major component of the breast cancer stroma, molecular and phenotypic heterogeneity of CAFs in breast cancer is poorly understood. In this study, we investigated CAF heterogeneity in triple-negative breast cancer (TNBC) using a syngeneic mouse model, BALB/c-derived 4T1 mammary tumors. Using single-cell RNA sequencing (scRNA-seq), we identified six CAF subpopulations in 4T1 tumors including: 1) myofibroblastic CAFs, enriched for α-smooth muscle actin and several other contractile proteins; 2) ‘inflammatory’ CAFs with elevated expression of inflammatory cytokines; and 3) a CAF subpopulation expressing major histocompatibility complex (MHC) class II proteins that are generally expressed in antigen-presenting cells. Comparison of 4T1-derived CAFs to CAFs from pancreatic cancer revealed that these three CAF subpopulations exist in both tumor types. Interestingly, cells with inflammatory and MHC class II-expressing CAF profiles were also detected in normal breast/pancreas tissue, suggesting that these phenotypes are not tumor microenvironment-induced. This work enhances our understanding of CAF heterogeneity, and specifically targeting these CAF subpopulations could be an effective therapeutic approach for treating highly aggressive TNBCs.
Keywords: cancer-associated fibroblasts, breast cancer, mammary fat pad, gene expression profiling, scRNA-seq, CAF heterogeneity, normal fibroblasts, myofibroblasts, inflammatory fibroblasts, pancreatic cancer
1. Introduction
Tumors are heterogeneous cellular entities in which progression depends on the dynamic crosstalk between cancer cells and other cells present in the stromal microenvironment [1]. The stroma is composed of supporting cells including fibroblasts, vascular endothelial cells, pericytes, mesenchymal stem cells and various types of immune cells. These cells are surrounded by fibrous structural proteins that comprise a dense extracellular matrix [2]. Cancer-associated fibroblasts (CAFs) are a predominant stromal cellular component in most solid tumors including breast, prostate and pancreatic cancers [3,4]. Previously it has been shown that resident tissue fibroblasts, bone marrow-derived mesenchymal stem cells, hematopoietic stem cells, adipocytes and endothelial cells can all give rise to CAFs, each by different mechanisms [5,6]. In addition, CAFs may arise directly from the cancer cells themselves via epithelial-mesenchymal transition (EMT) [7]. Factors released from CAFs into the tumor microenvironment play crucial roles in tumor growth, angiogenesis, metastasis, and resistance to therapy [5]. Thus, targeting CAFs directly to turn off their downstream effects or inhibiting CAF-secreted factors that stimulate tumor development and progression could represent a potential strategy for treating solid tumors.
Breast cancer is the most frequent cancer in women where an estimated 1.7 million women are diagnosed with breast cancer worldwide, every year [8]. Triple-negative breast cancer (TNBC) is a highly aggressive form of breast cancer, characterized by the absence of three receptors: estrogen (ER), progesterone (PR) and epidermal growth factor receptor 2 (HER2) [9]. TNBCs are resistant to most available targeted therapies because these treatments require the presence of at least one of these receptors to be effective in killing the cancer cells [9]. In addition, ~10–20% of all invasive breast cancers are TNBC [10]. TNBC metastasizes earlier and more frequently than other types of breast tumors; the 5-year survival rate for patients with TNBC is ~77% compared to 93% for all other breast cancer types [11]. Therefore, the development of novel therapeutic strategies for treating TNBC is urgently needed. Understanding the crosstalk between CAF subtypes and other cells in the TNBC microenvironment could open potential new avenues for cancer treatment.
Cancer-associated fibroblasts (CAFs) also coexist as heterogeneous populations, and several CAF subtypes with distinct molecular profiles have been identified in various cancers [6,12,13,14,15,16]. Although CAFs are the most prominent stromal components in solid tumors, identifying all possible subtypes and their specialized functions is far from complete. It has been previously shown that CAFs express high levels of alpha-smooth muscle actin (α-SMA/Acta2), CD90 (Thy1), platelet-derived growth factor receptors α or β (Pdgfra/b), integrin β1/CD29, podoplanin (Pdpn), osteonectin (Sparc), fibroblast activation protein (Fap), fibroblast-specific protein 1 (S100a4), caveolin 1 (Cav1) and vimentin (Vim) [3,14]. Several recent studies have used these markers to identify and characterize CAFs in various cancers [14,17,18,19]. However, these markers are far from being all-encompassing or completely specific to these cell subtypes, preventing us from identifying subtle differences among CAF subtypes using conventional methods. Single-cell RNA sequencing (scRNA-seq) allows us to profile gene expression in individual cells in a tissue with complex architecture and provides a high-resolution window into transcriptional differences. In turn, these molecular differences may lead to a better understanding of the function of each specific cell [20]. Furthermore, scRNA-seq enables us to discover rare cell types that until now may have been overlooked by traditional methods [21]. Several studies have utilized scRNA-seq to investigate CAF heterogeneity in solid tumors including pancreatic, breast and colorectal cancer, advancing our understanding of CAF heterogeneity [3,15,16], but no study to date has compared CAF subpopulations in various tumor types and also to fibroblast subpopulations present in healthy, normal tissues.
In this study, we characterized the fibroblast heterogeneity in a mouse allograft model of TNBC. Syngeneic mammary fat pad tumors were generated by injecting 4T1 breast cancer cells into BALB/c mice. Palpable tumors were dissected, and gene expression was profiled at single-cell level. The scRNA-seq analysis identified six CAF subpopulations in 4T1 mammary fat pad tumors including: 1) a CAF subpopulation with elevated expression of α-smooth muscle actin (α-SMA) and other contractile proteins including Tnc, Tagln and Myl9; 2) a subpopulation enriched in Ly6c1 and inflammatory cytokines Cxcl12, Il6 and Ccl2; and 3) a CAF subpopulation expressing Cd74 and other MHC class II proteins. Furthermore, we compared the CAF signatures of 4T1 tumors to those of pancreatic tumors from a genetically engineered mouse model (GEMM), the KPC mouse [22], and from subcutaneous allografts with a cell line (mT3) derived from the KPC mice [23], and of normal tissue resident fibroblasts to determine their similarities and differences. α-SMA-high CAFs, inflammatory CAFs and MHC class II-expressing CAFs were found in both breast and pancreatic tumors and shared highly similar transcriptional profiles. Interestingly, cells with inflammatory CAF profile and MHC class II-expressing CAF profile were also found endogenous to healthy breast/pancreas tissues, suggesting that these types of fibroblasts are not induced by the tumor microenvironment and may play important roles in tissue homeostasis.
2. Results
2.1. scRNA-seq Reveals Transcriptional Profiles of CAFs in Murine Mammary Tumors
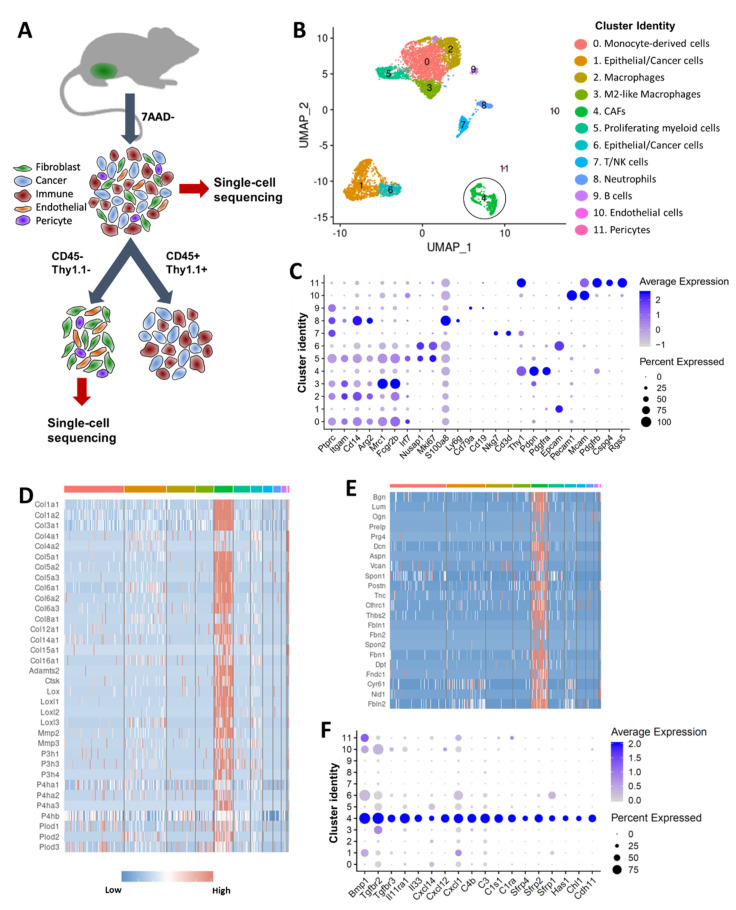
scRNA-seq was conducted on viable cells isolated from BALB/c-derived 4T1 orthotopic tumors using the 10x Genomics Chromium platform (Figure 1A). Of cells sequenced, 6420 cells met our quality control metrics and were further analyzed to identify various cell types in the tumor. A graph-based clustering using Seurat [24] identified 12 cell clusters (Figure 1B). By cross-referencing genes differentially expressed in each cluster to previously published cell-type specific markers, we assigned each cluster to its putative cell-type identity (Figure 1B). Cells in clusters 0, 2, 3, 5, 7, 8 and 9 expressed CD45 (Ptprc) and several other markers of myeloid and lymphoid lineages, and were classified as immune cells (Figure 1B,C, Table S1). Immune cells accounted for 66.4% of all sequenced cells. Clusters expressing high levels of Epcam (clusters 1 and 6) were identified as epithelial/cancer cells and accounted for ~24.5% of all cells (Figure 1B,C, Table S1). Cells in cluster 4 had high levels of Thy1, Pdpn and Pdgfra [25] and were identified as CAFs (Figure 1B,C, Table S1). This cluster included 535 cells and accounted for ~8% of all cells analyzed. Cells in cluster 10 expressed high levels of Pecam1 and Mcam and were identified as endothelial cells (Figure 1B,C, Table S1). We also identified a small population of pericytes (cluster 11) (Figure 1B). Interestingly, pericytes shared many markers with CAFs including Thy1 and Pdgfrb but also had unique markers such as NG2 (Cspg4), Mcam and Rgs5 [26,27] (Figure 1C, Table S1).
Figure 1.
Single cell analysis of 4T1 mouse mammary tumors. (A) Graphical representation of the experimental workflow. 4T1 syngeneic tumors were dissociated into single cells, and two cell fractions were generated: (1) a viable cell fraction (7AAD−) and (2) immune-depleted stromal cell fraction (obtained by depleting CD45+ immune cells and Thy1.1+ cancer cells). Cells from both fractions were subjected to single cell sequencing using the 10x Genomics Chromium platform. (B) Cell clusters from 10x Genomics scRNA-seq analysis visualized by Uniform Manifold Approximation and Projection (UMAP). Colors indicate clusters of various cell types (CAFs in black circle). (C) Dot plot showing the expression of selected markers of various cell types. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression level in that cluster. (D) Heatmap showing high levels of collagens and collagen-processing enzymes in CAFs. (E) Heatmap showing high levels of key proteoglycans and glycoproteins in CAFs. (F) Dot plot showing the expression of a subset of genes enriched in CAFs. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster.
Next, we compared the gene expression profile of CAFs (cluster 4) to other cell types in the tumor. Although we were only able to identify fewer CAFs in 4T1 mammary tumors than generally expected (~8% CAFs vs ~66% immune cells and ~24% cancer cells), potentially due to the limitations of the single cell isolation method, our analysis identified a large number of genes significantly enriched in CAFs compared to other cell types (Table S1). CAFs have been shown to play a major role in extracellular matrix (ECM) synthesis and organization [28]. Consistent with these functions, 4T1-derived CAFs showed enrichment for collagens (Col1a1, Col1a2, Col3a1, Col5a1-3, Col6a1-3 etc.), proteoglycans (Dcn, Lum, Bgn, Prg4 etc.) and glycoproteins (Postn, Dpt, Tnc, Fbln, Fbn1 etc.) (Figure 1D,E). Although CAFs expressed basement membrane collagens Col4a1 and Col4a2, the highest expression for these two genes was detected in pericytes and endothelial cells. CAFs also expressed high levels of several enzymes involved in processing and assembly of collagen into fibrils including Adamts-2, prolyl-4-hydroxylases (P4ha1-3), lysyl hydroxylases (Plod1-3) and lysyl oxidases (Lox, Loxl1-3) [29] (Figure 1D). Several ECM catabolic enzymes including matrix metalloproteinase-2 (Mmp2), matrix metalloproteinase-3 (Mmp3) and cathepsin K (Ctsk) were also enriched in CAFs (Figure 1D). CAFs also showed enrichment for bone morphogenic protein 1 (Bmp1), Tgf-β receptors (Tgfbr2, Tgfrb3), Wnt signaling pathway inhibitors (Sfrp1, Sfrp2, Sfrp4), complement pathway genes (C1ra, C1s1, C3, C4b), cytokines and cytokine receptors including Cxcl1, Cxcl12, Cxcl14, Il1r1 and Il11ra1(Figure 1F, Table S1). Cell adhesion proteins Cdh11 and Chl1 were also enriched in CAFs (Figure 1F).
2.2. Six Distinct Subtypes of CAFs Were Detected in Murine Mammary Tumors
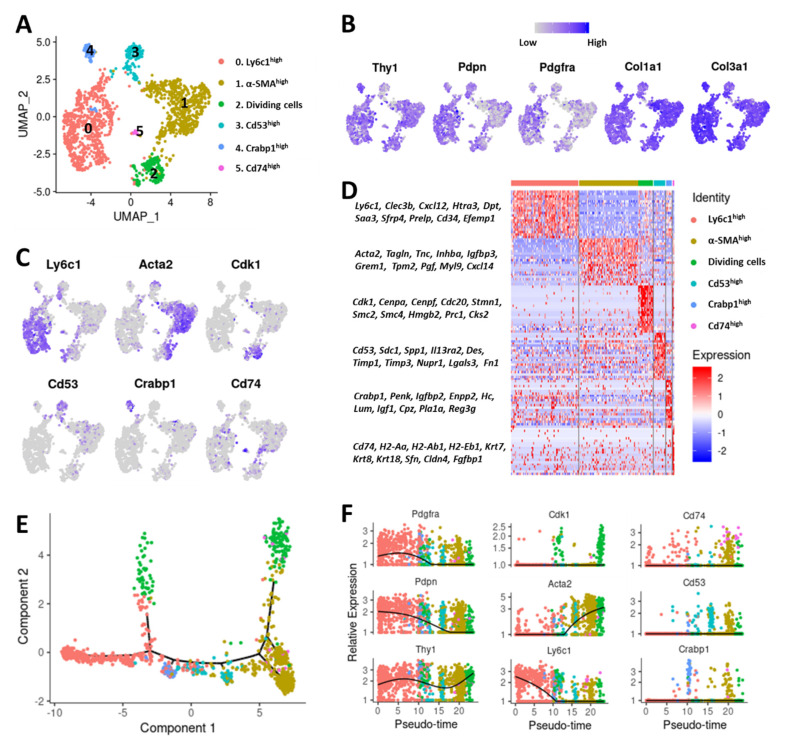
To better understand CAF heterogeneity in syngeneic 4T1 mammary tumors, we performed scRNA-seq on an immune (CD45+) and cancer cell (Thy1.1+) depleted fraction of 4T1-Thy1.1 tumor single cell suspension (Figure 1A). Among the ~4000 cells sequenced, we detected epithelial (Epcam+), endothelial (Pecam1+), pericytes (Mcam+, Rgs5+) and low levels of immune cells (CD45+); these cells were excluded from subsequent analysis (Figure S1A,B). The remaining ~1600 cells were classified as CAFs based on the expression of commonly used CAF markers (Figure S1A,C). Unsupervised clustering of these CAFs identified six distinct clusters with unique gene expression signatures (Figure 2A and Figure S1D–E). CAF markers Pdpn, Thy1 and Pdgfra [25] and CAF-secreted collagens Col1a1 and Col3a1 were expressed in all these clusters (Figure 2B). Cells in cluster 0 expressed high levels of lymphocyte antigen 6 complex, locus C1 (Ly6c1) while cells in cluster 1 were highly enriched for alpha smooth muscle actin (α-SMA/Acta2) (Figure 2C); as such, these genes were chosen to represent these clusters. Cells in cluster 2 expressed high levels of cyclin-dependent kinase 1 (Cdk1) and other cell cycle genes including Cenpa and Cenpf and were identified as ‘dividing cells’ (Figure 2C). Leukocyte surface antigen Cd53 was highly enriched in cluster 3 and was used as a marker for this cluster (Figure 2C). Cells in cluster 4 showed significant enrichment for cellular retinoic acid-binding protein 1 (Crabp1), while cells in cluster 5 showed enrichment for Cd74 (Figure 2C). Crabp1 and Cd74 were chosen as representative markers of clusters 4 and 5, respectively. We also found that these clusters had distinct gene expression profiles with a significant number of genes differentially expressed between these six clusters (Figure 2D, Table S2).
Figure 2.
Characterization of CAF subtypes in 4T1 breast cancer. (A) Cell clusters from 10x Genomics scRNA-seq analysis visualized by UMAP. Colors indicate various CAF subtypes. (B) Feature plots showing the expression of commonly used CAF markers in various CAF subtypes. Legend shows a color gradient of normalized expression. (C) Markers of various CAF subtypes that were used to denote each subtype. (D) Heatmap showing a subset of genes differentially expressed between the 6 CAF subtypes. (E) Monocle pseudospace trajectory colored based on CAF clusters in (A). (F) Expression of CAF markers on a pseudotime scale (colored based on CAF clusters in (A)).
To explore the relationship among these CAF subtypes, we constructed a transcriptional trajectory of these cells on a pseudotime scale using Monocle [30]. Cells in cluster 0 (Ly6c1high) were distributed at one end of the pseudotemporal trajectory whereas cells in cluster 1 (α-SMA/Acta2high) resided at the other end, suggesting that these clusters are most divergent from each other (Figure 2E,F). Pseudotime analysis also suggested that dividing/cycling cells from cluster 2 (Cdk1high) included cells diverging from both Ly6c1high and Acta2high clusters (Figure 2E,F). Cluster 3 (Cd53high) cells existed along the trajectory. Cluster 4 (Crabp1high) resided next to Ly6c1high cluster while cluster 5 (Cd74high) cells were mainly distributed at the other end of the trajectory, closer to the α-SMAhigh cluster (Figure 2E,F).
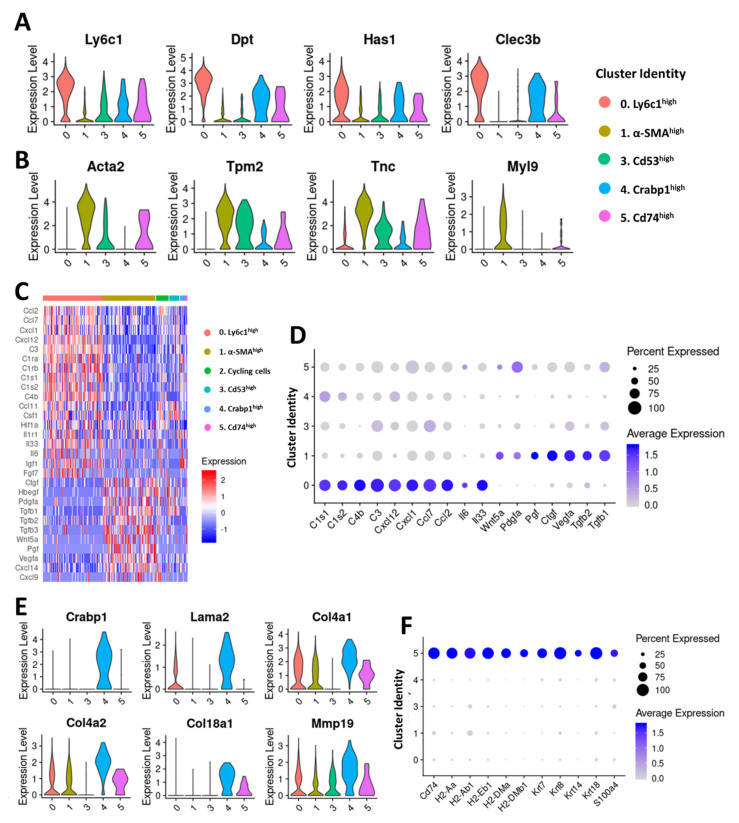
The Ly6c1high (cluster 0) and α-SMAhigh (cluster 1) clusters together constituted ~79% of fibroblasts in the 4T1 tumor (Figure 2A) and were further analyzed to gain insights into their potential roles in tumor development and progression (Figure 3A,B). Ly6c1 is an antigen present in neutrophils, monocytes, dendritic cells, and T cells [31] and its function in fibroblasts is not yet known. The Ly6c1high cluster also expressed other marker genes including ECM protein dermatopontin (Dpt), plasminogen-binding C-type lectin tetranectin (Clec3b), and hyaluronan synthase 1 (Has1), an enzyme responsible for cellular hyaluronan synthesis [32] at higher levels (Figure 3A). In addition, Ly6c1high cluster showed enrichment for stem cell antigen-1 (Ly6a/Sca-1), serum amyloid A3 (Saa3), collagen 14a1 (Col14a1) which plays a regulatory role in collagen fibrillogenesis [33], a small leucine-rich proteoglycan osteoglycin (Ogn), proteoglycan 4 (Prg4), prolargin (Prelp), EGF-containing fibulin-like extracellular matrix protein 1 (Efemp1), and HtrA serine peptidase 3 (Htra3) (Figure S2A). Hyaluronan, a major non-protein glycosaminoglycan component of the ECM, has been shown to promote cancer cell proliferation, migration, invasion, adhesion, EMT and cancer stem cell activation [34]. Increased Has1 expression may contribute to increased hyaluronan synthesis in Ly6c1high CAFs. Ogn plays a restrictive role in cancer progression [35,36]. Htra3 may also have a tumor-suppressive function [37]. Efemp1 and Prg4 can have either tumor-promoting or tumor-suppressive function [38,39,40,41]. We also observed elevated expression of transcripts encoding immune modulatory cytokines including interleukin-6 (Il6), interleukin 33 (Il33), chemokine (C-X-C motif) ligand 1 (Cxcl1), C-X-C motif chemokine 12 (Cxcl12), monocyte chemoattractant protein-1 (MCP-1/Ccl2) and monocyte-chemotactic protein 3 (MCP3/Ccl7) and several members of compliment pathway including C3, C4b, C1s1 and C1s2 in Ly6c1high cluster (Figure 3C,D), suggesting that Ly6c1high CAFs play a role in regulating the immune responses in the tumor microenvironment.
Figure 3.
Characterization of CAF clusters in 4T1 mammary tumors. Violin plots showing the expression of Ly6c1high (A) and α-SMAhigh (B) cluster markers in all clusters except the dividing/cycling cells. (C) Heatmap depicting the expression profiles of growth factors and immune/inflammatory signaling mediators differentially expressed between CAF subpopulations. (D) Dot plots showing the expression of selected immune/inflammatory signaling molecules enriched in Ly6c1high CAFs and growth factors enriched in α-SMAhigh CAFs. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster. (E) Violin plots showing the expression of selected markers enriched in Crabp1high CAFs. (F) Dot plots showing the expression of selected markers of Cd74high CAFs. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster.
The α-SMAhigh cluster (cluster 1), had a molecular signature similar to the myofibroblasts, which have been shown to play crucial roles in wound healing and pathological tissue remodeling [42,43]. Genes highly expressed in the α-SMAhigh cluster but absent or significantly reduced in other clusters included: contractile proteins tropomyosins 1 and 2 (Tpm1, Tpm2) and myosin light chain 9 (Myl9), transgelin (Tagln), calponins (Cnn2 and Cnn3), insulin-like growth factor-binding protein 3 (Igfbp3), tenascin C (Tnc) and transmembrane Protein 119 (Tmem119) (Figure 3B and Figure S2A). Interestingly, the α-SMAhigh cluster showed enrichment for several growth factor genes including transforming growth factor β (Tgfb1 and Tgfb2), connective tissue growth factor (CCN2/Ctgf), placental growth factor (Pgf), vascular endothelial growth factor A (Vegfa) and Wnt5a (Figure 3D). These growth factors have been implicated in various aspects of cancer development and progression including cell proliferation, migration, invasion, EMT and angiogenesis, suggesting that α-SMAhigh CAFs may promote tumor growth and progression [44,45,46,47,48]. We also observed that Ly6c1high CAFs expressed higher levels of platelet derived growth factor receptor alpha (Pdgfra), a commonly used CAF marker, whereas platelet derived growth factor receptor beta (Pdgfrb) expression was higher in cells from the α-SMAhigh cluster (Figure S2A).
Although small in size, Cd53high (cluster 3), Crabp1high (cluster 4) and Cd74high (cluster 5) clusters also displayed unique gene expression profiles, suggesting that these CAFs may also have distinct functions in the tumor microenvironment (Figure 2A,D). Cells in Cd53high cluster showed significant transcriptional enrichment for desmin (Des), a cytoplasmic intermediate filament protein which plays a crucial role in structural integrity and function of muscle [49] (Figure S2B). Other genes enriched in this cluster included matrix glycoprotein fibronectin 1 (Fn1) which has a well-established role in tumor development and progression [50,51,52], integrin alpha 1 (Itga1), syndecan 1 (Sdc1), matrix metalloproteinase inhibitors Timp1, Timp2 and Timp3 and galectin 3 (Lgals3), a potential regulator of cell migration, proliferation, angiogenesis, EMT and apoptosis [53] (Figure S2B). Basement membrane collagens Col4a1, Col4a2, Col18a1, laminin A2 (Lama2) and Mmp19, a protease which has been reported to degrade several basement membrane proteins [54] were among the genes highly enriched in the Crabp1high cluster (Figure 3E). This cluster also showed enrichment for perlecan (Hspg2), a major component of basement membranes, ECM proteins lumican (Lum), decorin (Dcn) and spondin 1 (Spon1), and insulin-like growth factor 1 (Igf1) (Figure S2B). Cells in the Cd74high cluster uniquely expressed high levels of MHC class II genes (H2-Aa, H2-Ab1, H2-Eb, Cd74 etc.) which are normally expressed by antigen-presenting cells (Figure 3F). This cluster also expressed commonly used CAF markers such as Pdpn, Pdgfra, Thy1, Col1a1, Col3a1 and Dcn at comparable levels to other CAF clusters confirming that these Cd74high cells are CAFs and not immune cells (Figure S2C). Expression of the pan-CAF markers such as Pdpn, Pdgfra and Dcn were extremely low in immune cells and epithelial cells whereas MHC class II genes were expressed in antigen-presenting immune cells at high levels (Figure S2D). We also detected enrichment for transcripts encoding fibroblast-specific protein 1 (FSP1/S100a4), keratins Krt7, Krt8, Krt14 and Krt18 and claudins Cldn3, Cldn4 and Cldn7 in this cluster which were also enriched in epithelial/cancer cells (Figure 3F and Figure S2B,D).
2.3. TNBC-Derived CAF Subtypes Share Molecular Features with Pancreatic Ductal Adenocarcinoma (PDAC)-Derived CAF Subtypes
Elyada et al. have previously demonstrated that three distinct CAF subtypes exist in PDAC: 1) “myofibroblastic CAFs” (myCAFs), that express high levels of α-SMA and other contractile genes such as Tagln, Myl9 and tropomyosins; 2) “inflammatory CAFs” (iCAFs) which express low levels of α-SMA but high levels of cytokines and other markers such as Ly6c1, Clec3b, Dpt and Has1; 3) antigen-presenting CAFs (apCAFs) which express MHC class II-related genes and induce T cell receptor (TCR) ligation in CD4+ T cells in an antigen-dependent manner [15]. Here, we compared the profiles of breast and pancreatic tumor-derived CAFs to gain more insights into CAF heterogeneity across different solid tumor types.
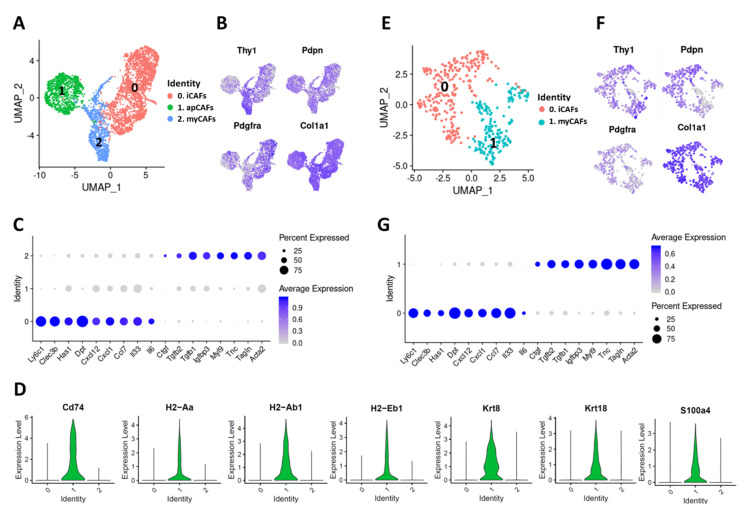
We reproduced Elyada et al.’s findings using publicly available scRNA-seq data derived from a genetic mouse model of PDAC, the KPC mouse (Kras+/LSL-G12D; Trp53+/LSL-R172H; Pdx-Cre) (GSE129455) [15], and identified the three CAF subtypes (Figure 4A). All of these CAF subtypes expressed pan-CAF markers including Thy1, Pdpn, Pdgfra and Col1a1 (Figure 4B). The CAF subtypes identified in PDAC included a Ly6c1high cluster (cluster 0) which was identified as iCAFs based on the expression of markers including Ly6c1, Clec3b, Has1, Dpt and Col14a1 and a α-SMAhigh cluster (cluster 2) with high expression of Acta2, Tagln, Myl9, Igfbp3 and Tnc which was identified as myCAFs (Figure 4A–C). As in the case for TNBC, PDAC-derived Ly6c1high CAFs (iCAFs) also expressed genes coding for chemokines and other inflammatory mediators such as Il6, Il33, Cxcl1, Cxcl12 and Ccl7 while α-SMAhigh CAFs (myCAFs) showed enrichment for growth factors Tgfb1, Tgfb2 and Ctgf (Figure 3C,D and Figure 4C). We also identified a Cd74high cluster (cluster 1) which was enriched for MHC class II-related genes Cd74, H2-Aa, H2-Ab1 and H2-Eb1 and was identified as apCAFs (Figure 4A,D). Interestingly, in both cancer types Cd74high CAFs (apCAFs) showed enrichment for keratins including Krt8 and Krt18 and Fsp1 (S100a4), a fibroblast marker (Figure 3F and Figure 4D). A comparative analysis of 4T1- and KPC-derived CAFs further confirmed that iCAFs, myCAFs and apCAFs are highly similar in both breast and pancreatic cancers (Figure S3).
Figure 4.
Characterization of CAF subtypes in murine PDAC. (A) UMAP plot showing various CAF subtypes identified in PDAC tumors from KPC mice. (B) Expression of commonly used CAF markers in KPC-derived PDAC CAFs. (C) Dot plot showing the expression of selected markers of Ly6c1high CAFs (iCAFs) and α-SMAhigh CAFs (myCAFs) in KPC-derived PDAC CAFs. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster. (D) Violin plots showing the expression of Cd74high CAF (apCAFs) markers in KPC-derived CAFs (cluster identities on X-axis). (E) CAF subtypes identified in subcutaneous mT3 tumors. (F) Expression of commonly used CAF markers in mT3 tumor-derived CAFs. (G) Dot plot showing the expression of selected iCAF and myCAF markers in mT3-derived CAFs.
Additionally, we generated syngeneic tumors in immunocompetent C57BL/6 mice by injecting the KPC-derived PDAC cell line mT3, subcutaneously (SQ) [23]. Of the ~2500 cells sequenced from an immune-depleted tumor, 434 cells were classified as CAFs, forming two distinct clusters (Figure 4E). Both CAF clusters expressed pan-CAF markers Pdpn, Thy1, Pdgfra and Col1a1 (Figure 4F). Further analysis identified these 2 CAF subtypes as: 1) Ly6c1high CAFs (iCAFs) based on enrichment for Ly6c1, Clec3b, Has1 and Dpt and 2) α-SMAhigh CAFs (myCAFs) based on enrichment for Acta2, Tagln, Tnc and Myl9 (Figure 4G). Interestingly, we did not detect any Cd74high cells in the syngeneic mT3 SQ tumors.
Regardless of the tumor type or tumor site, Ly6c1high CAFs (iCAFs) showed enrichment for Il33, Il6, Ccl7, Cxcl1 and Cxcl12 whereas α-SMAhigh CAFs (myCAFs) expressed high levels of Tgfb1, Tgfb2 and Ctgf (Figure 3D and Figure 4C,G). This comparative gene expression analysis showed high concordance between Ly6c1high, α-SMAhigh and Cd74high CAFs in both breast and pancreatic cancers, but points out that SQ tumor models have some limitations.
2.4. Cells with Ly6c1high and Cd74high CAF Profiles are Present in Normal, Healthy Tissues
To understand the molecular profiles of tissue resident fibroblasts, we isolated Pdgfra-expressing fibroblasts from the mammary fat pads of normal BALB/c mice [55] and performed single-cell sequencing on these cells. The ~1600 sequenced cells clustered in 3 distinct subtypes (Figure 5A). All clusters expressed CAF/fibroblast markers Pdgfra, Dcn, Postn and Col1a1; however, Pdpn and Thy1 expression was more restricted to clusters 0 and 2 (Figure 5B,C). Interestingly, clusters 0 and 2 were enriched for markers of Ly6c1high CAFs including Ly6c1, Clec3b, Dpt, Has1 and Col14a1, and cytokines including Ccl2, Ccl7, Il6, Cxcl1 and Cxcl12, although there were some differences in the expression of these genes between these two clusters (Figure 5C,D, Table S3). This suggests that fibroblasts with a Ly6c1high CAF profile exist in normal tissues as well. Clusters 0 and 2 fibroblasts were also enriched for ECM remodeling enzymes including Mmp2, Ctsk, Adamts5 and Htra3 but showed some differences in the expression of core ECM genes (Figure 5E and Figure S4A, Table S3). We did not detect any α-SMAhigh or Cd74high clusters among normal mammary fibroblasts although an extremely small fraction of cells expressed these genes (Figure 5D). However, none of these cells expressed Cd53 or Crabp1. Cluster 1 showed enrichment for several long non-coding RNAs including maternally expressed 3 (Meg3), nuclear paraspeckle assembly transcript 1 (Neat1), X-inactive specific transcript (Xist) and metastasis associated lung adenocarcinoma transcript 1 (Malat1) and the expression of Ly6c1high CAF markers were extremely low in this cluster (Figure S4A and Figure 5C).
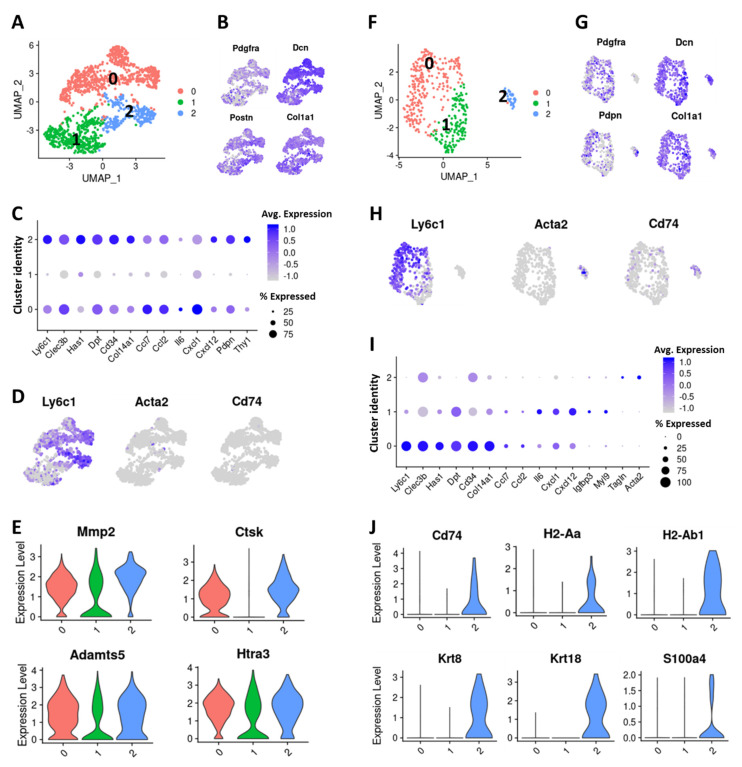
Figure 5.
Characterization of normal tissue-resident fibroblasts. (A) UMAP plot showing various fibroblast subtypes identified in normal mammary fat pad. (B) Expression of commonly used fibroblast/CAF markers in mammary fat pad-derived fibroblast subtypes (C) Dot plot showing the expression of selected iCAF markers in mammary fat pad-derived fibroblast subtypes. Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster. (D) Feature plot showing the expression of Ly6c1, Acta2 and Cd74 in mammary fat pad-derived fibroblasts. (E) Violin plot showing the expression of matrix degrading enzymes in mammary fat pad-derived fibroblast subtypes (cluster identity on X-axis). (F) UMAP plot showing various fibroblast subtypes identified in normal pancreas. (G) Expression of commonly used fibroblast/CAF markers in normal pancreas-derived fibroblasts. (H) Feature plot showing the expression of Ly6c1, Acta2 and Cd74 in pancreas-derived fibroblasts. (I) Dot plot showing the expression of selected iCAF markers in pancreas-derived fibroblast subtypes Dot size represents the fraction of cells expressing a specific marker in a particular cluster and intensity of color indicates the average expression in that cluster. (J) Violin plots showing the expression of Cd74high CAF (apCAFs) markers in pancreas-derived fibroblasts subtypes.
A comparative analysis of fibroblasts derived from normal mammary fat pad and 4T1 tumor-derived CAFs revealed that Ly6c1high CAFs from the tumor share high molecular similarity with Ly6c1high fibroblasts from normal mammary fat pad (Figure S5A–E), expressing common markers such as Ly6c1, Clec3b, Dpt, Ly6a, Htra3, Has1 and Col14a1 (Figure S5A,E). However, transcripts encoding cytokines Cxcl12 and Il33 had a significantly higher expression in the tumor-derived CAFs, suggesting a tumor microenvironment induced activation of these genes in this subpopulation (Figure S5E). Expression of markers of other CAF subtypes were mainly restricted to the tumor (Figure S5D,E).
We also analyzed a publicly available normal pancreas scRNA-seq data (GEO: GSM3577882) [56] to determine fibroblast diversity in this tissue. Fibroblasts were identified based on the expression of markers Pdgfra, Pdpn, Thy1, Dcn, Col1a1 and Col3a1 and were further analyzed to identify distinct subtypes. In normal pancreas, we identified 3 clusters expressing fibroblast markers (Figure 5F,G). Cluster 0 showed significant enrichment for Ly6c1high inflammatory CAF markers Ly6c1, Clec3b, Has1, Dpt and Col14a1 (Figure 5H,I). Cluster 1 also expressed these markers at some level, and cells in both cluster 0 and 1 transcriptionally expressed cytokines including Ccl2, Ccl7, Il6, Cxcl1 and Cxcl12 (Figure 5I). Cluster 1 also showed enrichment for long noncoding RNAs Meg3, Neat1, Xist and Malat1 that were found to be enriched in a subset of normal mammary fibroblasts (Figure S4A,B). Interestingly, cluster 2 showed enrichment for Cd74 and other genes enriched in Cd74high CAFs including H2-Ab1, H2-Aa, Krt8, Krt18 and fibroblast marker Fsp1 (S100a4), suggesting that cells with Cd74high CAF features are also present in the normal pancreas (Figure 5J).
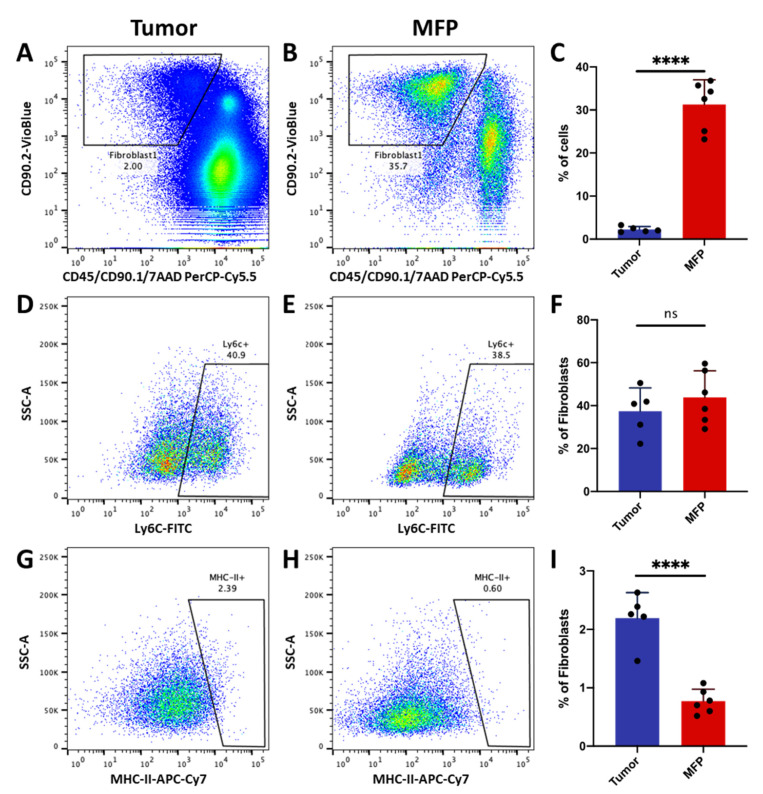
Using flow cytometry, we confirmed that Ly6c1high fibroblasts are present in both mammary tumors and naïve mammary fat pad. Fibroblasts were isolated by removing immune (anti-CD45) and cancer cells (anti-CD90.1/Thy1.1) and then selecting for cells expressing CAF marker Thy1 (CD90.2/Thy1.2) (Figure 2B). About 2.4% of cells in the tumor and ~30% of cells in the normal mammary fat pad expressed CD90.2/Thy1.2 (Figure 6A–F). About 40% of the cells in this fibroblast-enriched fraction expressed Ly6c in both the tumor and naïve mammary fat pad. Interestingly, a small proportion of these Thy1+ cells expressed MHC class II protein (Figure 6G–I), confirming that cells with Cd74high CAF profile are also present in both tumor and normal mammary tissue. Tumor tissue had a significantly higher proportion of these MHC class II-expressing cells than normal mammary fat pad (Figure 6G–I).
Figure 6.
Inflammatory fibroblasts and MHC class II-expressing fibroblasts in mammary tumors and naïve mammary fat pad. Flow cytometry plots identifying viable (7AAD−) CD90.2+/CD90.1−/CD45− fibroblasts derived from (A) tumor or (B) naïve mammary fat pad (MFP). (C) Bar graph of Thy1+ fibroblast abundance in tumor and MFP. Flow cytometry plots identifying Ly6c1high subpopulations from all Thy1+ fibroblasts in the tumor (D) and (E) MFP. (F) Bar graph of Ly6c1high subpopulations abundance in the tumor and MFP. Flow cytometry plots identifying MHC class II-expressing subpopulations from all Thy1+ fibroblasts in the tumor (G) and (H) MFP. (I) Bar graph showing the abundance of MHC class II -expressing subpopulation in the tumor and MFP. n = 5–6 performed in two independent experiments. Errors bars denote SD. **** p < 0.0001, not significant (ns) p > 0.005.
3. Discussion
In this study, we have carried out scRNA-seq of 4T1 mammary and mT3 pancreatic syngeneic tumors to investigate CAF heterogeneity across TNBC and PDAC tumor types. Our study identified six CAF subtypes in TNBC with distinct gene expression profiles: 1) Ly6c1high CAFs; 2) α-SMAhigh CAFs; 3) dividing/cycling CAFs; 4) Cd53high CAFs; 5) Crabp1high CAFs and 6) Cd74high CAFs. A comparison of this data to data from KPC mice-derived pancreatic tumors revealed that 3 of these CAF populations, Ly6c1high CAFs (iCAFs), α-SMAhigh CAFs (myCAFs) and Cd74high CAFs (apCAFs) exist in both tumor types. These 3 CAF subtypes have also been detected in human PDACs [15].
A high proportion of cells in both breast and pancreatic cancers expressed α-SMA (Acta2). While the existence of myofibroblastic (α-SMAhigh) CAFs in solid tumors are well-established [3,57], the number of α-SMA-expressing fibroblasts was extremely low in normal breast and pancreatic tissue suggesting that this subtype emerges during tumorigenesis. In 4T1 mammary fat pad tumors, α-SMAhigh CAFs showed enrichment for several growth factor transcripts including Tgfb1, Tgfb2, Ctgf, Pgf, Vegfa and Wnt5a (Figure 2C,D). TGF-β is a multifunctional cytokine with a well-established role in fibroblast to myofibroblast differentiation, EMT and immune regulation [44,58,59,60]. The protein Vegfa is a key regulator of both physiological and pathological angiogenesis [46]. Other factors secreted by α-SMAhigh CAFs such as Ctgf, Pgf and Wnt5a have been implicated in cancer cell proliferation, migration, invasion, EMT and/or angiogenesis [45,47,48], suggesting that α-SMAhigh CAFs may promote tumor development and progression. Interestingly, a previous study has shown that instead of eradicating the tumor, depletion of myofibroblasts in PDAC caused an increase in tumor invasion; an event associated with decreased survival. These findings suggest that α-SMAhigh CAFs may also function to keep the tumor in check [61].
The Ly6c1high CAFs expressed high levels of transcripts encoding inflammatory cytokines including Ccl2, Il6, Il33 and Cxcl12. Stromal fibroblast derived Ccl2 has been shown to promote tumor progression and contribute to immune evasion [62,63]. IL6 has been shown to drive EMT, metastasis and therapy resistance in cancer [64,65,66,67]. IL6 is also plays a role in the generation of tumor-associated macrophages by skewing monocyte differentiation into tumor-associated macrophage [68]. It has been shown that Il33 plays a role in tumor growth, metastasis, neo-angiogenesis, and evading programmed cell death [69]. Multiple studies have shown that Cxcl12 promotes tumor angiogenesis, tumor cell proliferation and chemoresistance [70]. Recently, Costa et al. analyzed the expression of six previously known CAF markers (FAP, integrin β1/CD29, α-SMA, S100-A4/FSP1, PDGFRβ, and CAV1) in human breast cancer and discovered four different CAF subpopulations that expressed these markers at varying levels, including a subtype which promoted immunosuppression through a Cxcl12-dependent mechanism [14]. These studies together suggest that Ly6c1high CAFs may play a role in immune suppression in the tumor microenvironment. Interestingly, we found fibroblasts with Ly6c1high CAF profile in normal breast and pancreatic tissues, suggesting that Ly6c1high CAFs might have originated from resident fibroblasts. Alternatively, the resident fibroblasts with high Ly6c1 expression got recruited into the tumor during tumor development. It has been shown that IL1 signaling through IL1R promotes the Ly6c1high CAF (iCAF) phenotype via JAK–STAT signaling and inhibition of JAK/STAT signaling shifts iCAFs to a myofibroblastic phenotype in PDAC [59]. Therefore, Ly6c1high fibroblasts present in normal tissue may differentiate into α-SMAhigh CAFs (myCAFs) during tumor progression. Further studies are required to understand the specific cues from the tumor microenvironment that drive these phenotypes.
The Cd74high CAFs (apCAFs) expressed CD74 and other MHC class II proteins which are normally expressed by antigen presenting cells in the immune system. In addition, Cd74high CAFs showed enrichment for transcripts encoding keratins including Krt7, Krt8 and Krt18 and Fsp1 (S100a4), a protein predominately expressed in fibroblasts. Elyada et al. have shown the presence of apCAFs in both mouse and human PDAC samples [15]. They also showed that apCAFs can present antigens to CD4+ T cells [15]. Another scRNA-seq study showed the presence of a cell population with ‘apCAF’ profile in normal pancreas and identified it as mesothelial cells [71]. In normal pancreas, these cells expressed Pdpn but, lacked Pdgfra expression [71]. Here we showed that cells with apCAF profile are present in both normal mammary tissue and mammary tumors. In breast/pancreatic tumors and normal breast/pancreatic tissues, Cd74high cells expressed fibroblast marker Fsp1/S100a4 (Figure 3F, Figure 4D and Figure 5J) whereas Pdgfra expression appeared to be restricted to tumor tissue only (Figure S2C, Figure 4B and Figure 5G). Future labeling and lineage tracing experiments will be able to conclusively determine the origin of these cells. It has been hypothesized that apCAFs might have an immune modulatory role in the tumor microenvironment [15], and the possibility exists that these cells also protect healthy tissue from auto-immune reactions in conditions such as lupus or pancreatitis.
Recently, using scRNA-seq, Bartoschek et al. identified four transcriptionally distinct subpopulations of CAFs known as vascular CAFs (vCAFs), matrix CAFs (mCAFs), developmental CAFs (dCAFs) and cycling CAFs (cCAFs) in the genetically engineered MMTV-PyMT mouse model of breast cancer [16]. The vCAFs showed enrichment for several genes involved in vascular development including Mcam and Rgs5 and likely have a perivascular origin [16]. Upon re-analysis of this data we detected enrichment for myofibroblast markers Acta2, Tagln, Myl9 and Tpm2 in vCAFs, suggesting that some of these cells acquired a myofibroblastic molecular profile (Figure S6A–C). It was suggested that mCAFs originate from resident fibroblasts, and the proportion of mCAFs decrease with tumor progression [16]. We found that mCAFs were enriched for Crabp1 and several other markers of Crabp1high CAFs from 4T1 tumors including Lama2, Spon1, Dcn, Lum and Mmp19. We also detected weak Ly6c1 expression in MMTV-PyMT-derived mCAFs raising the possibility that mCAFs (Crabp1high) might have originated from Ly6c1high fibroblasts (Figure S6B). In agreement with this, Crabp1high CAFs resided adjacent to Ly6c1high CAFs on the pseudotemporal trajectory (Figure 2E). Further studies are required to understand specific functions of Crabp1high CAFs in the tumor microenvironment. However, we did not detect any Cd74high CAF clusters in this dataset. Cd53high CAFs were also not identified in this dataset or in the KPC model of pancreatic cancer. However, we could identify Cd53high CAFs in our initial 4T1 scRNA-seq data (Figure S6D–F). Future studies will determine whether Cd53high CAFs are a subtype with distinct origin and function or merely represent a transitional state during the differentiation of CAFs.
To our knowledge this is the first report that comprehensively compares CAF heterogeneity in multiple tumor types in parallel to the healthy, tissue of origin, using a single cell RNA sequencing approach. We have identified six CAF subtypes in TNBC with unique transcriptomic profiles, adding several subtypes to some that have been widely described in the literature. Our findings, in line with numerous other studies, suggest that specific cues from the microenvironment are necessary to maintain these unique molecular features; many of these subtypes have been shown to acquire a myofibroblasts-like phenotype when cultured in 2D or 3D [15,72]. A major limitation of our study is that we only examined tumors at one timepoint. It is possible that these populations shift as the tumor grows in size, as it acquires resistance to therapy, or as it metastasizes [16]. Future studies should aim to compare multiple tumor types and stages to further refine our understanding of CAF heterogeneity in solid tumors as well as to begin to elucidate individual contributions to tumor function.
4. Materials and Methods
4.1. Cell Lines
4T1 cells were purchased from ATCC (Manassas, VA, USA). The 4T1-Thy1.1 cell line was graciously provided as a gift from Dr. Julian Lum [73]. 4T1-Thy1.1 cells express a non-native isoform of Thy1, Thy1.1 (CD90.1) which differs from native isoform Thy1.2 (CD90.2) in one amino acid. The mT3 cell line was developed from organoids isolated from KPC mouse PDAC lesions [23]. The cell line is syngeneic and forms tumors in immune-competent C57BL/6 mice.
4.2. Generation of Orthotopic Mammary Tumors and Tumor Digestion
10-week-old female BALB/c mice (Jackson Laboratories, Bar Harbor, ME, USA) were injected with 25,000 4T1 cells (ATCC, Manassas, VA, USA) or 4T1-Thy1.1 cells in a 1:1 suspension of Matrigel (Corning, Corning, NY, USA; Catalog no. 354234) and phosphate buffered saline (PBS) into mammary fat pad (MFP) to establish tumors. Mice were euthanized 19–26 days post injection and the tumors were dissected from these mice. Single cell suspensions were generated by passing the tumor through a syringe without a needle followed by a 1h digest with shaking at 37 °C in 100 μg/mL DNase I (Roche, Basel, Switzerland; catalog no. 11284932001), 300 U/mL collagenase/100U/mL hyaluronidase (STEMCELL Technologies, Vancouver, BC, Canada; catalog no. 07912), 0.6 U/mL Dispase II (Roche, Basel, Switzerland; catalog no. 4942078001) in DMEM/D12 with 10% fetal bovine serum (FBS) (ThermoFisher, Waltham, MA, USA). Digests were filtered through a 100 µm cell strainer prior to debris removal (Miltenyi Biotec, Bergisch Gladbach, Germany; catalog no. 130-109-398) and resuspended in BD FACS Pre-Sort Buffer (BD, Franklin Lakes, NJ, USA; catalog no. 563503) followed by red blood cell lysis (ACK lysis buffer, ThermoFisher, Waltham, MA, USA; catalog no. A1049201) prior to downstream applications. All animal experimental procedures were completed under an approved institutional animal care and use committee (IACUC) protocol at Lawrence Livermore National Laboratory and conforming to the National Institutes of Health Guide for the care and use of laboratory animals.
4.3. Single Cell Sequencing of 4T1 Tumors and Data Analysis
4T1 tumor from a female BALB/c mouse was processed and cell suspension was prepared as described above. Two subsequent washes in sterile PBS + 0.04% non-acetylated bovine serum albumin were performed to further remove debris from final suspension. Cell pellets were resuspended in PBS with 0.04% non-acetylated BSA prior to single cell sequencing preparation using Chromium Single Cell 3′ GEM, Library & Gel Bead Kit v3 (10x Genomics, Pleasanton, CA, USA; Catalog no. 1000075) on a 10 Genomics Chromium Controller following manufacturers protocol. scRNA-Seq libraries were sequenced using Illumina (San Diego, CA, USA) NextSeq 500.
The Cell Ranger Single-Cell Software Suite (10x Genomics, Pleasanton, CA, USA) was used to perform sample demultiplexing, barcode processing, and single-cell 3′gene counting. Samples were aligned to the mouse genome (mm10) using “cellranger mkfastq” with default parameters. Unique molecular identifier (UMI) counts were generated using “cellranger count”. Further analysis was performed in R using the Seurat package [74]. First, cells with fewer than 500 detected genes per cell and genes that were expressed by fewer than 5 cells were filtered out. To remove noise from droplets containing more than one cell, cells with more 7800 measured genes were filtered out. Dead cells were excluded by retaining cells with less than 5% mitochondrial reads. After removing all the unwanted cells from the dataset, we normalized the data by employing a global-scaling normalization method “LogNormalize”. Subsequently, we identified the 2000 most variable genes in the dataset. The data was then scaled to a mean of 0 and variance of 1 and the dimensionality of the data was reduced by principal component analysis (PCA) using the previously determined 2000 variable genes. Subsequently, we constructed a K-nearest-neighbor (KNN) graph based on the Euclidean distance in PCA space using the “FindNeighbors” function (using dimensions 1 to 15) and applied Louvain algorithm to iteratively group cells together by “FindClusters” function (resolution = 0.3). A non-linear dimensional reduction was then performed via uniform manifold approximation and projection (UMAP) using the first 15 principle components. A total of 12 clusters were identified in the 4T1 scRNA-seq data. Raw count matrix is available online in Dryad data repository (https://datadryad.org, doi:10.6071/M3238R).
4.4. Single Cell Sequencing and Analysis of Stromal Cell-Enriched Fraction
4T1-Thy1.1 tumor from a female BALB/c mouse was processed as described above to obtain single cell suspensions. To enrich for stromal cells in the sample, immune cells and Thy1.1 expressing cancer cells were removed from the single cell suspension using magnetic cell separation with CD90.1 and CD45 MicroBeads (Miltenyi Biotec, Bergisch Gladbach, Germany; Catalog no. 130-121-273 and 130-052-301, respectively) depletion in combination with LS magnetic separation columns (Miltenyi Biotec, Bergisch Gladbach, Germany; Catalog no. 130-042-401). Cells were prepared for single cell sequencing using Chromium Single Cell 3′ GEM, Library & Gel Bead Kit v3 (10x Genomics, Pleasanton, CA, USA; Catalog no. 1000075) on a 10x Genomics Chromium Controller following manufacturers protocol and sequenced using Illumina (San Diego, CA, USA) NextSeq 500 at median sequencing depth of ~30,000 reads.
The scRNA-seq data was analyzed using Cell Ranger Single-Cell Software Suite (10x Genomics, Pleasanton, CA, USA) as described above to obtain UMI counts. Subsequent analysis was performed in Seurat [74] as described above and clusters of various cell types were identified. Clusters with high expression of Ptprc and Epcam were regarded as contaminants of immune cells and cancer cells, respectively and were discarded from subsequent analysis. Clusters with high expression of Pecam1 and Rgs5 were identified as endothelial cells and pericytes, respectively and were also discarded from further analysis. Resulting clusters showed enrichment for CAF markers Pdpn, Pdgfra, Thy1, Col1a1 and Dcn. Raw expression values were obtained for cells in these clusters and were further analyzed using Seurat as described above. To remove noise from droplets containing more than one cell, cells with more 5800 measured genes were filtered out. Subsequently, after performing a log-normalization we identified the 2000 most variable genes in the dataset. The data was then scaled, and the dimensionality of the data was reduced by PCA. Clusters of cells were identified on the basis of a shared-nearest neighbor graph between cells and the Louvain algorithm as described above. Subsequently, a dimensional reduction was performed via UMAP as described above and various CAF subtypes were identified. We subsequently repeated this analysis by varying the number of variable genes used for dimensionality reduction to further confirm the presence of these CAF subtypes in 4T1 tumors. Raw count matrix is available online in Dryad data repository (https://datadryad.org, doi:10.6071/M3238R).
4.5. Single-Cell Sequencing of Syngeneic mT3 Tumors and Data Analysis
To generate mT3 tumor, 10-week-old female C57BL/6 mice were injected with 25,000 mT3 cells subcutaneously (SQ) into the back flank in a 1:1 suspension of Matrigel (Corning, Corning, NY, USA; Catalog no. 354234) and PBS. At 3 weeks post injection the tumor was dissected and processed as described above to obtain single cell suspensions. Subsequently, immune cells and blood cells were removed by CD45+ magnetic bead-based depletion (Miltenyi Biotech, Bergisch Gladbach, Germany; Catalog no. 130-052-301) and ACK lysis buffer (ACK lysis buffer, ThermoFisher, Waltham, MA, USA catalog no. A1049201), respectively, following manufacturer’s guidelines. Remaining cells were prepared for single cell sequencing using Chromium Single Cell 3ʹ GEM, Library & Gel Bead Kit v3 (10x Genomics, Pleasanton, CA, USA; catalog no. 1000075) on a 10x Genomics Chromium Controller following manufacturers protocol and sequenced using Illumina (San Diego, CA, USA) NextSeq 500. The scRNA-seq data was demultiplexed and processed using Cell Ranger Single-Cell Software Suite (10x Genomics, Pleasanton, CA, USA) as described above to obtain UMI counts. Subsequent analysis was performed in Seurat [74] as described above and clusters of various cell types including cancer cells and CAFs were identified. Clusters expressing CAF markers Pdpn, Pdgfra, Thy1 and Dcn were extracted and further analyzed using Seurat as described above and various CAF subtypes were identified. Raw count matrix is available online in Dryad data repository (https://datadryad.org, doi:10.6071/M3238R).
4.6. Isolation and Sequencing of Fibroblasts from Normal Mammary Fat Pad
Fourth and fifth mammary fat pads were collected from four BALBc mice, aged approximately 12 weeks [55]. Tissue was minced and then digested in 50 mL RPMI with 10% heat inactivated (HI) FBS and Penicillin-Streptomycin and 1 mg/mL Collagenase IV (ThermoFisher, Waltham, MA, USA). After an hour, the suspension was strained using a 70 µm cell strainer and centrifuged at 500× g for 15 min. The supernatant was discarded, and the cells were resuspended in 1 mL ACK (ThermoFisher, Waltham, MA, USA) for five minutes. 10 mL of PBS with 1% HI FBS was added and the cells were spun down at 500× g for 5 min. The supernatant was discarded, and the cells were resuspended in 0.5 mL of PBS with 1% HI FBS. To identify fibroblasts, cells were stained with the following antibodies at a 1:100 dilution: anti-CD326 (FITC, clone G8.8, BioLegend, San Diego, CA, USA), anti-CD45 (APC-Cy7, clone 30-F11, BioLegend, San Diego, CA, USA), anti-CD140a (Super Bright 436, clone APA5, ThermoFisher, Waltham, MA, USA), and 7AAD to assess viability. Cells were then sorted on a BD Aria to gate for CD140a+ EPCAM− CD45− 7AAD− cells (fibroblasts). Cells were prepared for single cell sequencing using Chromium Single Cell 3ʹ GEM, Library & Gel Bead Kit v3 (10x Genomics, Pleasanton, CA, USA; catalog no. 1000075) on a 10x Genomics Chromium Controller following manufacturers protocol and sequenced using Illumina NextSeq 500.
The scRNA-seq data was demultiplexed and processed using Cell Ranger Single-Cell Software Suite (10x Genomics, Pleasanton, CA, USA) as described above to obtain UMI counts. Subsequent analysis was performed in Seurat [74]. To remove noise from droplets containing more than one cell, cells with more 5900 measured genes were filtered out. All the other filtering steps and data normalization were performed as described above for 4T1 tumors. Prior to PCA, we identified the 2000 most variable genes and PCA was performed in this gene space. Clusters of cells were identified on the basis of a shared-nearest neighbor graph between cells as described above. First 15 principal components were provided as an input for dimensionality reduction via UMAP. Raw count matrix is available online at in Dryad data repository (https://datadryad.org, doi:10.6071/M3238R).
Comparative analysis of normal mammary fibroblasts and 4T1tumor-derived CAFs was performed using Seurat. The data was pre-processed and normalized as described above and 2000 most variable features were identified. Subsequently, integration anchors were identified using ‘FindIntegrationAnchors’ function and both datasets were integrated to generate a new integrated dataset. The integrated data was scaled, dimensionality was reduced, and the data was further analyzed as described above to identify clusters of CAF-subtypes.
4.7. Analysis of Cancer-Associated Fibroblasts from Mouse Pancreatic Tumors
Fibroblast-enriched scRNA-seq data from PDAC tumors of 4 KPC mice were obtained from GEO (GSE129455) as log transformed gene-by-cell count matrix. Subsequent data analysis was carried out in R with Seurat [74] as described above and various cell types were identified. Clusters of cells expressing CAF markers Pdgfra, Pdpn, Thy1 and Col1a1 were further analyzed and various CAF subpopulations were identified. Comparative analysis of 4T1 tumor-derived and PDAC-derived CAFs were performed using Seurat as described above.
4.8. Analysis of Normal Fibroblasts from Mouse Pancreas
From GEO (GSE125588) we obtained processed scRNA-seq files (barcodes.tsv, genes.tsv and matrix.mtx generated by Cell Ranger) associated with normal pancreas. The data was analyzed using Seurat [74] as described above to identify various cell types. Clusters of cells expressing fibroblast markers Pdgfra, Dcn and Col1a1 were extracted and further analyzed using Seurat to identify various fibroblast subpopulations.
4.9. Analysis of Mammary Tumors from MMTV-PyMT Mice
scRNA-seq data for fibroblasts isolated from mammary tumors of two MMTV-PyMT mice were obtained from GEO (GSE111229) as raw counts. The data was analyzed using Seurat as described above to identify various CAF subtypes [74]. vCAFs, mCAFs, cCAFs and dCAFs were identified based on previously established markers [16].
4.10. Trajectory Finding
Single-cell pseudotime trajectories of CAFs were constructed with Monocle [30]. Expression data, phenotype data, and feature data were extracted from the Seurat object and a Monocle “CellDataSet” object was constructed using the “newCellDataSet” function. Highly variable genes from Seurat object were used as ordering genes. Dimensionality reduction was performed using the DDRTree algorithm implemented Monocle via the “reduceDimension” function. Cells were ordered along the trajectory using the “orderCells” method with default parameters.
4.11. Flow Cytometric Analysis of Inflammatory and MHC Class II-Expressing Fibroblasts
Single cell suspensions of normal mammary fat pads (n = 6) or 4T1 tumor-bearing BALB/c fat pads (n = 5) were generated as described above. Antibody staining was performed by incubating the cells with 1:100 dilution of APC/Cyanine7 anti-mouse I-A/I-E (MHC class II), Brilliant Violet 421™ anti-mouse CD90.2, PerCP/Cyanine5.5 anti-rat CD90/mouse CD90.1 (Thy-1.1), PerCP/Cyanine5.5 anti-mouse CD45, FITC anti-mouse Ly-6C (Biolegend, San Diego, CA, USA; Catalog no. 107627, 105341, 202515, 103131, 128005, respectively) for 30 min prior to resuspension in PBS with 1% FBS and 7-AAD Viability Staining Solution (BioLegend, San Diego, CA, USA; Catalog No. 420403). Cells were then analyzed on a BD (San Jose, CA, USA) FACSMelody instrument. Viable fibroblasts populations were identified as 7AAD−/CD45−/CD90.1−/CD90.2+ cells. Data analysis was performed using FlowJo software (v10.6.2).
5. Conclusions
This study systematically examined CAF heterogeneity in TNBCs and uncovered several CAF subpopulations in mammary tumor microenvironment. A comparison of breast and pancreatic-tumor-derived CAFs revealed that myofibroblast-like CAFs, inflammatory CAFs and MHC class II-expressing CAFs share similar molecular profiles in both tumor types. Our study also suggests that inflammatory fibroblasts and MHC class II-expressing fibroblasts are endogenous to healthy tissues, suggesting that these fibroblast subtypes are not induced by the tumor microenvironment but, are likely recruited to the tumor during tumor development. Further studies are required determine the specific functions of these CAF subpopulations in tumor development and progression. Effectively targeting these CAF subtypes could prevent cancer progression.
Acknowledgments
This work was performed under the auspices of the U.S. Department of Energy by Lawrence Livermore National Laboratory under Contract DE-AC52-07NA27344.
Supplementary Materials
The following are available online at https://www.mdpi.com/2072-6694/12/5/1307/s1, Figure S1: Single-cell sequencing of immune depleted stromal cell fraction, Figure S2: Markers of various CAF subtypes, Figure S3: Comparative analysis of 4T1-derived CAFs (BC) and KPC -derived CAFs (PDAC), Figure S4: Expression profiles of tissue resident fibroblasts, Figure S5: Comparative analysis of breast cancer-derived CAFs and normal mammary fat pad-derived fibroblasts, Figure S6: Identification Crabp1high and Cd53high CAFs in multiple breast cancer datasets, Table S1:Genes enriched (fold change >1.25) in each 4T1 tumor-derived cluster compared to all other clusters, Table S2: Genes enriched (fold change >1.25) in each 4T1 tumor-derived CAF cluster compared to all other CAF clusters, Table S3: Genes enriched (fold change >1.25) in each normal mammary fibroblast cluster compared to all other clusters.
Author Contributions
Conceptualization, A.S. and G.G.L.; data acquisition, N.R.H., S.F.G., K.A.M., A.S.; formal analysis, A.S.; writing A.S. and G.G.L.; resource sharing S.W.B., I.P.; project administration, E.K.W. and M.A.C. All authors have read and agreed to the published version of the manuscript.
Funding
The research was supported by LLNL internal grants LDRD grant number 19-SI-003 and 17-ER-121. K.A.M. was supported by DOE-funded Livermore Graduate Scholar Program. G.G.L was also supported in part by a UC Davis Comprehensive Cancer Center Support Grant (CCCSG) awarded by the National Cancer Institute (NCI P30CA093373).
Conflicts of Interest
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
References
- 1.Zhang W., Huang P. Cancer-stromal interactions: Role in cell survival, metabolism and drug sensitivity. Cancer Biol. Ther. 2011;11:150–156. doi: 10.4161/cbt.11.2.14623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Werb Z., Lu P. The Role of Stroma in Tumor Development. Cancer J. 2015;21:250–253. doi: 10.1097/PPO.0000000000000127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Musa M. Single-cell analysis on stromal fibroblasts in the microenvironment of solid tumours. Adv. Med. Sci. 2020;65:163–169. doi: 10.1016/j.advms.2019.12.001. [DOI] [PubMed] [Google Scholar]
- 4.Gandellini P., Andriani F., Merlino G., D’Aiuto F., Roz L., Callari M. Complexity in the tumour microenvironment: Cancer associated fibroblast gene expression patterns identify both common and unique features of tumour-stroma crosstalk across cancer types. Semin. Cancer Biol. 2015;35:96–106. doi: 10.1016/j.semcancer.2015.08.008. [DOI] [PubMed] [Google Scholar]
- 5.Shiga K., Hara M., Nagasaki T., Sato T., Takahashi H., Takeyama H. Cancer-Associated Fibroblasts: Their Characteristics and Their Roles in Tumor Growth. Cancers. 2015;7:2443–2458. doi: 10.3390/cancers7040902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Gascard P., Tlsty T.D. Carcinoma-associated fibroblasts: Orchestrating the composition of malignancy. Genes Dev. 2016;30:1002–1019. doi: 10.1101/gad.279737.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Cirri P., Chiarugi P. Cancer associated fibroblasts: The dark side of the coin. Am. J. Cancer Res. 2011;1:482–497. [PMC free article] [PubMed] [Google Scholar]
- 8.Torre L.A., Islami F., Siegel R.L., Ward E.M., Jemal A. Global Cancer in Women: Burden and Trends. Cancer Epidemiol. Biomark. Prev. 2017;26:444–457. doi: 10.1158/1055-9965.EPI-16-0858. [DOI] [PubMed] [Google Scholar]
- 9.Podo F., Buydens L.M., Degani H., Hilhorst R., Klipp E., Gribbestad I.S., Van Huffel S., van Laarhoven H.W., Luts J., Monleon D., et al. Triple-negative breast cancer: Present challenges and new perspectives. Mol. Oncol. 2010;4:209–229. doi: 10.1016/j.molonc.2010.04.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kumar P., Aggarwal R. An overview of triple-negative breast cancer. Arch. Gynecol. Obstet. 2016;293:247–269. doi: 10.1007/s00404-015-3859-y. [DOI] [PubMed] [Google Scholar]
- 11.Bauer K.R., Brown M., Cress R.D., Parise C.A., Caggiano V. Descriptive analysis of estrogen receptor (ER)-negative, progesterone receptor (PR)-negative, and HER2-negative invasive breast cancer, the so-called triple-negative phenotype: A population-based study from the California cancer Registry. Cancer. 2007;109:1721–1728. doi: 10.1002/cncr.22618. [DOI] [PubMed] [Google Scholar]
- 12.Sugimoto H., Mundel T.M., Kieran M.W., Kalluri R. Identification of fibroblast heterogeneity in the tumor microenvironment. Cancer Biol. Ther. 2006;5:1640–1646. doi: 10.4161/cbt.5.12.3354. [DOI] [PubMed] [Google Scholar]
- 13.Ohlund D., Handly-Santana A., Biffi G., Elyada E., Almeida A.S., Ponz-Sarvise M., Corbo V., Oni T.E., Hearn S.A., Lee E.J., et al. Distinct populations of inflammatory fibroblasts and myofibroblasts in pancreatic cancer. J. Exp. Med. 2017;214:579–596. doi: 10.1084/jem.20162024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Costa A., Kieffer Y., Scholer-Dahirel A., Pelon F., Bourachot B., Cardon M., Sirven P., Magagna I., Fuhrmann L., Bernard C., et al. Fibroblast Heterogeneity and Immunosuppressive Environment in Human Breast Cancer. Cancer Cell. 2018;33:463–479.e10. doi: 10.1016/j.ccell.2018.01.011. [DOI] [PubMed] [Google Scholar]
- 15.Elyada E., Bolisetty M., Laise P., Flynn W.F., Courtois E.T., Burkhart R.A., Teinor J.A., Belleau P., Biffi G., Lucito M.S., et al. Cross-Species Single-Cell Analysis of Pancreatic Ductal Adenocarcinoma Reveals Antigen-Presenting Cancer-Associated Fibroblasts. Cancer Discov. 2019;9:1102–1123. doi: 10.1158/2159-8290.CD-19-0094. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Bartoschek M., Oskolkov N., Bocci M., Lovrot J., Larsson C., Sommarin M., Madsen C.D., Lindgren D., Pekar G., Karlsson G., et al. Spatially and functionally distinct subclasses of breast cancer-associated fibroblasts revealed by single cell RNA sequencing. Nat. Commun. 2018;9:1–13. doi: 10.1038/s41467-018-07582-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Gorchs L., Hellevik T., Bruun J.A., Camilio K.A., Al-Saad S., Stuge T.B., Martinez-Zubiaurre I. Cancer-associated fibroblasts from lung tumors maintain their immunosuppressive abilities after high-dose irradiation. Front. Oncol. 2015;5:87. doi: 10.3389/fonc.2015.00087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Hu C., Wang Z., Zhai L., Yang M., Shan L., Chai C., Liu M., Wang L. Effects of cancer-associated fibroblasts on the migration and invasion abilities of SGC-7901 gastric cancer cells. Oncol. Lett. 2013;5:609–612. doi: 10.3892/ol.2012.1023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nagasaki T., Hara M., Nakanishi H., Takahashi H., Sato M., Takeyama H. Interleukin-6 released by colon cancer-associated fibroblasts is critical for tumour angiogenesis: Anti-interleukin-6 receptor antibody suppressed angiogenesis and inhibited tumour-stroma interaction. Br. J. Cancer. 2014;110:469–478. doi: 10.1038/bjc.2013.748. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Tang F., Lao K., Surani M.A. Development and applications of single-cell transcriptome analysis. Nat. Methods. 2011;8(Suppl. 4):S6–S11. doi: 10.1038/nmeth.1557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Baslan T., Hicks J. Unravelling biology and shifting paradigms in cancer with single-cell sequencing. Nat. Rev. Cancer. 2017;17:557–569. doi: 10.1038/nrc.2017.58. [DOI] [PubMed] [Google Scholar]
- 22.Hingorani S.R., Wang L., Multani A.S., Combs C., Deramaudt T.B., Hruban R.H., Rustgi A.K., Chang S., Tuveson D.A. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell. 2005;7:469–483. doi: 10.1016/j.ccr.2005.04.023. [DOI] [PubMed] [Google Scholar]
- 23.Boj S.F., Hwang C.I., Baker L.A., Chio I.I., Engle D.D., Corbo V., Jager M., Ponz-Sarvise M., Tiriac H., Spector M.S., et al. Organoid models of human and mouse ductal pancreatic cancer. Cell. 2015;160:324–338. doi: 10.1016/j.cell.2014.12.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Butler A., Hoffman P., Smibert P., Papalexi E., Satija R. Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat. Biotechnol. 2018;36:411–420. doi: 10.1038/nbt.4096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Liu T., Han C., Wang S., Fang P., Ma Z., Xu L., Yin R. Cancer-associated fibroblasts: An emerging target of anti-cancer immunotherapy. J. Hematol. Oncol. 2019;12:1–15. doi: 10.1186/s13045-019-0770-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Esteves C.L., Donadeu F.X. Pericytes and their potential in regenerative medicine across species. Cytom. A. 2018;93:50–59. doi: 10.1002/cyto.a.23243. [DOI] [PubMed] [Google Scholar]
- 27.Shen J., Shrestha S., Yen Y.H., Scott M.A., Soo C., Ting K., Peault B., Dry S.M., James A.W. The pericyte antigen RGS5 in perivascular soft tissue tumors. Hum. Pathol. 2016;47:121–131. doi: 10.1016/j.humpath.2015.09.013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Liu T., Zhou L., Li D., Andl T., Zhang Y. Cancer-Associated Fibroblasts Build and Secure the Tumor Microenvironment. Front. Cell Dev. Boil. 2019;7:60. doi: 10.3389/fcell.2019.00060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Canty E.G., Kadler K.E. Procollagen trafficking, processing and fibrillogenesis. J. Cell Sci. 2005;118 Pt 7:1341–1353. doi: 10.1242/jcs.01731. [DOI] [PubMed] [Google Scholar]
- 30.Trapnell C., Cacchiarelli D., Grimsby J., Pokharel P., Li S., Morse M., Lennon N.J., Livak K.J., Mikkelsen T.S., Rinn J.L. The dynamics and regulators of cell fate decisions are revealed by pseudotemporal ordering of single cells. Nat. Biotechnol. 2014;32:381–386. doi: 10.1038/nbt.2859. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lee P.Y., Wang J.X., Parisini E., Dascher C.C., Nigrovic P.A. Ly6 family proteins in neutrophil biology. J. Leukoc. Biol. 2013;94:585–594. doi: 10.1189/jlb.0113014. [DOI] [PubMed] [Google Scholar]
- 32.Itano N., Sawai T., Atsumi F., Miyaishi O., Taniguchi S., Kannagi R., Hamaguchi M., Kimata K. Selective expression and functional characteristics of three mammalian hyaluronan synthases in oncogenic malignant transformation. J. Biol. Chem. 2004;279:18679–18687. doi: 10.1074/jbc.M313178200. [DOI] [PubMed] [Google Scholar]
- 33.Ansorge H.L., Meng X., Zhang G., Veit G., Sun M., Klement J.F., Beason D.P., Soslowsky L.J., Koch M., Birk D.E. Type XIV Collagen Regulates Fibrillogenesis: Premature collagen fibril growth and tissue dysfunction in null mice. J. Biol. Chem. 2009;284:8427–8438. doi: 10.1074/jbc.M805582200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Price Z.K., Lokman N.A., Ricciardelli C. Differing Roles of Hyaluronan Molecular Weight on Cancer Cell Behavior and Chemotherapy Resistance. Cancers. 2018;10:482. doi: 10.3390/cancers10120482. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hu X., Li Y.Q., Li Q.G., Ma Y.L., Peng J.J., Cai S.J. Osteoglycin (OGN) reverses epithelial to mesenchymal transition and invasiveness in colorectal cancer via EGFR/Akt pathway. J. Exp. Clin. Cancer Res. 2018;37:41. doi: 10.1186/s13046-018-0718-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Xu T., Zhang R., Dong M., Zhang Z., Li H., Zhan C., Li X. Osteoglycin (OGN) Inhibits Cell Proliferation and Invasiveness in Breast Cancer via PI3K/Akt/mTOR Signaling Pathway. Onco Targets Ther. 2019;12:10639–10650. doi: 10.2147/OTT.S222967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Li Y., Gong L., Qi R., Sun Q., Xia X., He H., Ren J., Zhu O., Zhuo D. Paeoniflorin suppresses pancreatic cancer cell growth by upregulating HTRA3 expression. Drug Des. Dev. Ther. 2017;11:2481–2491. doi: 10.2147/DDDT.S134518. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Hu J., Duan B., Jiang W., Fu S., Gao H., Lu L. Epidermal growth factor-containing fibulin-like extracellular matrix protein 1 (EFEMP1) suppressed the growth of hepatocellular carcinoma cells by promoting Semaphorin 3B(SEMA3B) Cancer Med. 2019;8:3152–3166. doi: 10.1002/cam4.2144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Yin X., Fang S., Wang M., Wang Q., Fang R., Chen J. EFEMP1 promotes ovarian cancer cell growth, invasion and metastasis via activated the AKT pathway. Oncotarget. 2016;7:47938–47953. doi: 10.18632/oncotarget.10296. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Sarkar A., Chanda A., Regmi S.C., Karve K., Deng L., Jay G.D., Jirik F.R., Schmidt T.A., Bonni S. Recombinant human PRG4 (rhPRG4) suppresses breast cancer cell invasion by inhibiting TGFbeta-Hyaluronan-CD44 signalling pathway. PLoS ONE. 2019;14:e0219697. doi: 10.1371/journal.pone.0219697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Oikawa K., Mizusaki A., Takanashi M., Ozaki T., Sato F., Kuroda M., Muragaki Y. PRG4 expression in myxoid liposarcoma maintains tumor cell growth through suppression of an antitumor cytokine IL-24. Biochem. Biophys. Res. Commun. 2017;485:209–214. doi: 10.1016/j.bbrc.2017.02.055. [DOI] [PubMed] [Google Scholar]
- 42.Hinz B., Phan S.H., Thannickal V.J., Galli A., Bochaton-Piallat M.L., Gabbiani G. The myofibroblast: One function, multiple origins. Am. J. Pathol. 2007;170:1807–1816. doi: 10.2353/ajpath.2007.070112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Guerrero-Juarez C.F., Dedhia P.H., Jin S., Ruiz-Vega R., Ma D., Liu Y., Yamaga K., Shestova O., Gay D.L., Yang Z., et al. Single-cell analysis reveals fibroblast heterogeneity and myeloid-derived adipocyte progenitors in murine skin wounds. Nat. Commun. 2019;10:1–17. doi: 10.1038/s41467-018-08247-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Wendt M.K., Allington T.M., Schiemann W.P. Mechanisms of the epithelial-mesenchymal transition by TGF-beta. Future Oncol. 2009;5:1145–1168. doi: 10.2217/fon.09.90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Aguiar D.P., de Farias G.C., de Sousa E.B., de Mattos Coelho-Aguiar J., Lobo J.C., Casado P.L., Duarte M.E., Abreu J.G., Jr. New strategy to control cell migration and metastasis regulated by CCN2/CTGF. Cancer Cell Int. 2014;14:61. doi: 10.1186/1475-2867-14-61. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Nagy J.A., Dvorak A.M., Dvorak H.F. VEGF-A and the induction of pathological angiogenesis. Annu. Rev. Pathol. 2007;2:251–275. doi: 10.1146/annurev.pathol.2.010506.134925. [DOI] [PubMed] [Google Scholar]
- 47.Dewerchin M., Carmeliet P. Placental growth factor in cancer. Expert Opin. Ther. Targets. 2014;18:1339–1354. doi: 10.1517/14728222.2014.948420. [DOI] [PubMed] [Google Scholar]
- 48.Zeng R., Huang J., Zhong M.Z., Li L., Yang G., Liu L., Wu Y., Yao X., Shi J., Wu Z. Multiple Roles of WNT5A in Breast Cancer. Med. Sci. Monit. 2016;22:5058–5067. doi: 10.12659/MSM.902022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Paulin D., Li Z. Desmin: A major intermediate filament protein essential for the structural integrity and function of muscle. Exp. Cell Res. 2004;301:1–7. doi: 10.1016/j.yexcr.2004.08.004. [DOI] [PubMed] [Google Scholar]
- 50.Rick J.W., Chandra A., Dalle Ore C., Nguyen A.T., Yagnik G., Aghi M.K. Fibronectin in malignancy: Cancer-specific alterations, protumoral effects, and therapeutic implications. Semin. Oncol. 2019;46:284–290. doi: 10.1053/j.seminoncol.2019.08.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Kumra H., Reinhardt D.P. Fibronectin-targeted drug delivery in cancer. Adv. Drug Deliv. Rev. 2016;97:101–110. doi: 10.1016/j.addr.2015.11.014. [DOI] [PubMed] [Google Scholar]
- 52.Wang J.P., Hielscher A. Fibronectin: How Its Aberrant Expression in Tumors May Improve Therapeutic Targeting. J. Cancer. 2017;8:674–682. doi: 10.7150/jca.16901. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53.Nangia-Makker P., Hogan V., Raz A. Galectin-3 and cancer stemness. Glycobiology. 2018;28:172–181. doi: 10.1093/glycob/cwy001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Brauer R., Beck I.M., Roderfeld M., Roeb E., Sedlacek R. Matrix metalloproteinase-19 inhibits growth of endothelial cells by generating angiostatin-like fragments from plasminogen. BMC Biochem. 2011;12:38. doi: 10.1186/1471-2091-12-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Sharon Y., Alon L., Glanz S., Servais C., Erez N. Isolation of normal and cancer-associated fibroblasts from fresh tissues by Fluorescence Activated Cell Sorting (FACS) J. Vis. Exp. 2013;71:e4425. doi: 10.3791/4425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Hosein A.N., Huang H., Wang Z., Parmar K., Du W., Huang J., Maitra A., Olson E., Verma U., Brekken R.A. Cellular heterogeneity during mouse pancreatic ductal adenocarcinoma progression at single-cell resolution. JCI Insight. 2019 doi: 10.1172/jci.insight.129212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Li H., Courtois E.T., Sengupta D., Tan Y., Chen K.H., Goh J.J.L., Kong S.L., Chua C., Hon L.K., Tan W.S., et al. Reference component analysis of single-cell transcriptomes elucidates cellular heterogeneity in human colorectal tumors. Nat. Genet. 2017;49:708–718. doi: 10.1038/ng.3818. [DOI] [PubMed] [Google Scholar]
- 58.Evans R.A., Tian Y.C., Steadman R., Phillips A.O. TGF-beta1-mediated fibroblast-myofibroblast terminal differentiation-the role of Smad proteins. Exp. Cell Res. 2003;282:90–100. doi: 10.1016/S0014-4827(02)00015-0. [DOI] [PubMed] [Google Scholar]
- 59.Biffi G., Oni T.E., Spielman B., Hao Y., Elyada E., Park Y., Preall J., Tuveson D.A. IL1-Induced JAK/STAT Signaling Is Antagonized by TGFbeta to Shape CAF Heterogeneity in Pancreatic Ductal Adenocarcinoma. Cancer Discov. 2019;9:282–301. doi: 10.1158/2159-8290.CD-18-0710. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Li M.O., Wan Y.Y., Sanjabi S., Robertson A.K., Flavell R.A. Transforming growth factor-beta regulation of immune responses. Annu. Rev. Immunol. 2006;24:99–146. doi: 10.1146/annurev.immunol.24.021605.090737. [DOI] [PubMed] [Google Scholar]
- 61.Ozdemir B.C., Pentcheva-Hoang T., Carstens J.L., Zheng X., Wu C.C., Simpson T.R., Laklai H., Sugimoto H., Kahlert C., Novitskiy S.V., et al. Depletion of carcinoma-associated fibroblasts and fibrosis induces immunosuppression and accelerates pancreas cancer with reduced survival. Cancer Cell. 2014;25:719–734. doi: 10.1016/j.ccr.2014.04.005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Tsuyada A., Chow A., Wu J., Somlo G., Chu P., Loera S., Luu T., Li A.X., Wu X., Ye W., et al. CCL2 mediates cross-talk between cancer cells and stromal fibroblasts that regulates breast cancer stem cells. Cancer Res. 2012;72:2768–2779. doi: 10.1158/0008-5472.CAN-11-3567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Zhang J., Chen L., Xiao M., Wang C., Qin Z. FSP1+ fibroblasts promote skin carcinogenesis by maintaining MCP-1-mediated macrophage infiltration and chronic inflammation. Am. J. Pathol. 2011;178:382–390. doi: 10.1016/j.ajpath.2010.11.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64.Shintani Y., Fujiwara A., Kimura T., Kawamura T., Funaki S., Minami M., Okumura M. IL-6 Secreted from Cancer-Associated Fibroblasts Mediates Chemoresistance in NSCLC by Increasing Epithelial-Mesenchymal Transition Signaling. J. Thorac. Oncol. 2016;11:1482–1492. doi: 10.1016/j.jtho.2016.05.025. [DOI] [PubMed] [Google Scholar]
- 65.Ebbing E.A., van der Zalm A.P., Steins A., Creemers A., Hermsen S., Rentenaar R., Klein M., Waasdorp C., Hooijer G.K.J., Meijer S.L., et al. Stromal-derived interleukin 6 drives epithelial-to-mesenchymal transition and therapy resistance in esophageal adenocarcinoma. Proc. Natl. Acad. Sci. USA. 2019;116:2237–2242. doi: 10.1073/pnas.1820459116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Sun X., Mao Y., Wang J., Zu L., Hao M., Cheng G., Qu Q., Cui D., Keller E.T., Chen X., et al. IL-6 secreted by cancer-associated fibroblasts induces tamoxifen resistance in luminal breast cancer. Oncogene. 2014 doi: 10.1038/onc.2014.158. [DOI] [PubMed] [Google Scholar]
- 67.Wu X., Tao P., Zhou Q., Li J., Yu Z., Wang X., Li J., Li C., Yan M., Zhu Z., et al. IL-6 secreted by cancer-associated fibroblasts promotes epithelial-mesenchymal transition and metastasis of gastric cancer via JAK2/STAT3 signaling pathway. Oncotarget. 2017;8:20741–20750. doi: 10.18632/oncotarget.15119. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Jeannin P., Duluc D., Delneste Y. IL-6 and leukemia-inhibitory factor are involved in the generation of tumor-associated macrophage: Regulation by IFN-gamma. Immunotherapy. 2011;3(Suppl. 4):23–26. doi: 10.2217/imt.11.30. [DOI] [PubMed] [Google Scholar]
- 69.Shen J.X., Liu J., Zhang G.J. Interleukin-33 in Malignancies: Friends or Foes? Front. Immunol. 2018;9:3051. doi: 10.3389/fimmu.2018.03051. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Meng W., Xue S., Chen Y. The role of CXCL12 in tumor microenvironment. Gene. 2018;641:105–110. doi: 10.1016/j.gene.2017.10.015. [DOI] [PubMed] [Google Scholar]
- 71.Dominguez C.X., Muller S., Keerthivasan S., Koeppen H., Hung J., Gierke S., Breart B., Foreman O., Bainbridge T.W., Castiglioni A., et al. Single-Cell RNA Sequencing Reveals Stromal Evolution into LRRC15(+) Myofibroblasts as a Determinant of Patient Response to Cancer Immunotherapy. Cancer Discov. 2020;10:232–253. doi: 10.1158/2159-8290.CD-19-0644. [DOI] [PubMed] [Google Scholar]
- 72.Hum N.R., Sebastian A., Gilmore S.F., He W., Martin K.A., Hinckley A., Dubbin K.R., Moya M.L., Wheeler E.K., Coleman M.A., et al. Comparative Molecular Analysis of Cancer Behavior Cultured In Vitro, In Vivo, and Ex Vivo. Cancers. 2020;12:690. doi: 10.3390/cancers12030690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73.Sobral-Filho R.G., DeVorkin L., Macpherson S., Jirasek A., Lum J.J., Brolo A.G. Ex Vivo Detection of Circulating Tumor Cells from Whole Blood by Direct Nanoparticle Visualization. ACS Nano. 2018;12:1902–1909. doi: 10.1021/acsnano.7b08813. [DOI] [PubMed] [Google Scholar]
- 74.Stuart T., Butler A., Hoffman P., Hafemeister C., Papalexi E., Mauck W.M., 3rd, Hao Y., Stoeckius M., Smibert P., Satija R. Comprehensive Integration of Single-Cell Data. Cell. 2019;177:1888–1902. doi: 10.1016/j.cell.2019.05.031. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.