Abstract
Shoot branching from axillary buds (AXBs) is regulated by a network of inhibitory and promotive forces, which includes hormones. In perennials, the dwarfed stature of the embryonic shoot inside AXBs is indicative of gibberellin (GA) deficiency, suggesting that AXB activation and outgrowth require GA. Nonetheless, the role of GA in branching has remained obscure. We here carried out comprehensive GA transcript and metabolite analyses in hybrid aspen, a perennial branching model. The results indicate that GA has an inhibitory as well as promotive role in branching. The latter is executed in two phases. While the expression level of GA2ox is high in quiescent AXBs, decapitation rapidly downregulated it, implying increased GA signaling. In the second phase, GA3ox2-mediated de novo GA-biosynthesis is initiated between 12 and 24 h, prior to AXB elongation. Metabolite analyzes showed that GA1/4 levels were typically high in proliferating apices and low in the developmentally inactive, quiescent AXBs, whereas the reverse was true for GA3/6. To investigate if AXBs are differently affected by GA3, GA4, and GR24, an analog of the branch-inhibitor hormone strigolactone, they were fed into AXBs of single-node cuttings. GA3 and GA4 had similar effects on GA and SL pathway genes, but crucially GA3 induced AXB abscission whereas GA4 promoted outgrowth. Both GA3 and GA4 strongly upregulated GA2ox genes, which deactivate GA1/4 but not GA3/6. Thus, the observed production of GA3/6 in quiescent AXBs targets GA1/4 for GA2ox-mediated deactivation. AXB quiescence can therefore be maintained by GA3/6, in combination with strigolactone. Our discovery of the distinct tasks of GA3 and GA4 in AXB activation might explain why the role of GA in branching has been difficult to decipher. Together, the results support a novel paradigm in which GA3/6 maintains high levels of GA2ox expression and low levels of GA4 in quiescent AXBs, whereas activation and outgrowth require increased GA1/4 signaling through the rapid reduction of GA deactivation and subsequent GA biosynthesis.
Keywords: gibberellin, axillary branching, GA2-oxidases, GA3-oxidases, GA20-oxidases, GID1, strigolactone, hormones
Introduction
Shoot branching is governed by a network of hormones that includes auxin, cytokinin (CK) and strigolactone (SL). How they interact to regulate axillary bud (AXB) activation and outgrowth still divides opinion (Ferguson and Beveridge, 2009; Hayward et al., 2009; Müller and Leyser, 2011; Puig et al., 2012; Rameau et al., 2015). Classic experiments established that a growing shoot apex can repress branching, a phenomenon known as apical dominance. The physiological explanation is that a proliferating apex produces a surplus of auxin that is send down the stem to inhibit AXB outgrowth, thereby promoting apical elongation. Removal of the apex releases AXBs from inhibition, triggering branching, but this can be prevented by supplying auxin to the cut stem (Thimann and Skoog, 1934; Phillips, 1975; Cline, 1991, 1997).
A current interpretation of these experiments is that the growing apex increases the relative amount of auxin in the polar auxin transport stream (PATS) of the main stem, thereby preventing AXBs from establishing their own auxin export path to the stem (Li and Bangerth, 1999; Bennett et al., 2006; Ongaro et al., 2008; Domagalska and Leyser, 2011). When auxin levels in the stem drop, export of auxin from the AXB to the stem is initiated, promoting AXB outgrowth. An alternative model proposes that auxin export is a consequence of AXB activation rather than a cause (Dun et al., 2006; Brewer et al., 2009; Ferguson and Beveridge, 2009). This is in line with the proposal of Cline (1997) that a fast initial enlargement of an AXB should be distinguished from the much slower outgrowth process. Experimental support comes from studies with garden pea (Pisum sativum L.), in which shoot decapitation triggers AXB enlargement ahead of the arrival of the auxin depletion front (Morris et al., 2005). Moreover, supplying auxin to the cut stem can prevent branching but not AXB enlargement. Finally, depleting stem auxin levels by auxin transport inhibitors does not affect initial AXB enlargement, but once AXBs have enlarged it promotes sustained outgrowth (Morris et al., 2005; Ferguson and Beveridge, 2009; Mason et al., 2014). In addition to the network of hormones, nutrients are important in AXB outgrowth in intact plants, as well as after decapitation when sugars are diverted to the larger AXBs, which are the strongest sinks (Mason et al., 2014; Kebrom, 2017).
The transcription factor BRANCHED1 (BRC1)/TEOSINTE BRANCHED1 (TB1) is an important branch-inhibitor (Aguilar-Martínez et al., 2007; Brewer et al., 2009; Dun et al., 2009; Leyser, 2009). Although BRC1 was originally identified as the target of SL, it is now recognized to be a hub for branch-regulating signals, including various hormones and developmental as well as environmental cues (Wang et al., 2019). In Arabidopsis (Arabidopsis thaliana), BRC1 inhibits AXB outgrowth, probably by suppressing cell proliferation (Schommer et al., 2014), but in some circumstances it cannot prevent outgrowth (Seale et al., 2017). In rice (Oryza sativa), SL also induces degradation of the branch-promoting hormone CK through transcriptional activation of CK-oxidases (Duan et al., 2019). In accordance with this, AXB outgrowth in pea is accompanied by a reduction in SL biosynthesis and an increase in CK biosynthesis (Tanaka et al., 2006; Ferguson and Beveridge, 2009). Auxin also suppresses CK biosynthesis (Nordström et al., 2004). Thus, CK-induced outgrowth of activated AXBs may require low stem levels of auxin and SL.
While auxin, CKs and SLs are implicated in the regulation of AXBs, the role of gibberellins (GA) has remained obscure. This is unexpected as GAs promote many developmental processes, including germination, elongation, floral transition as well as AXB formation and dormancy release (Hazebroek et al., 1993; Richards et al., 2001; Yamaguchi, 2008; Rinne et al., 2011, 2016; Claeys et al., 2014; Zhuang et al., 2015). GA is often viewed as a branch-inhibitor because GA-biosynthesis and -perception mutants in Arabidopsis, as well as GA-deficient transgenic plants of various species have branched phenotypes. However, a complicating factor is that GA-deficiency or lack of GA perception not only increases branching but also reduces apical dominance (Scott et al., 1967; Talon et al., 1990; Murfet and Reid, 1993; Silverstone et al., 1997; Olszewski et al., 2002; Busov et al., 2003; Agharkar et al., 2007; Lo et al., 2008; Mauriat et al., 2011; Zawaski and Busov, 2014; Rameau et al., 2015). In contrast to the above, some studies suggest that GA promotes branching. In perennial strawberry, AXB outgrowth is diminished in a GA-biosynthesis mutant, while GA supply rescues the phenotype (Tenreira et al., 2017). Similarly, in the woody species Jatropha (J. curcas L.) (Ni et al., 2015) and hybrid aspen (Populus tremula × P. tremuloides) (Rinne et al., 2011), GA application promotes AXB outgrowth, whereas in Rosa sp. outgrowth requires GA biosynthesis (Choubane et al., 2012).
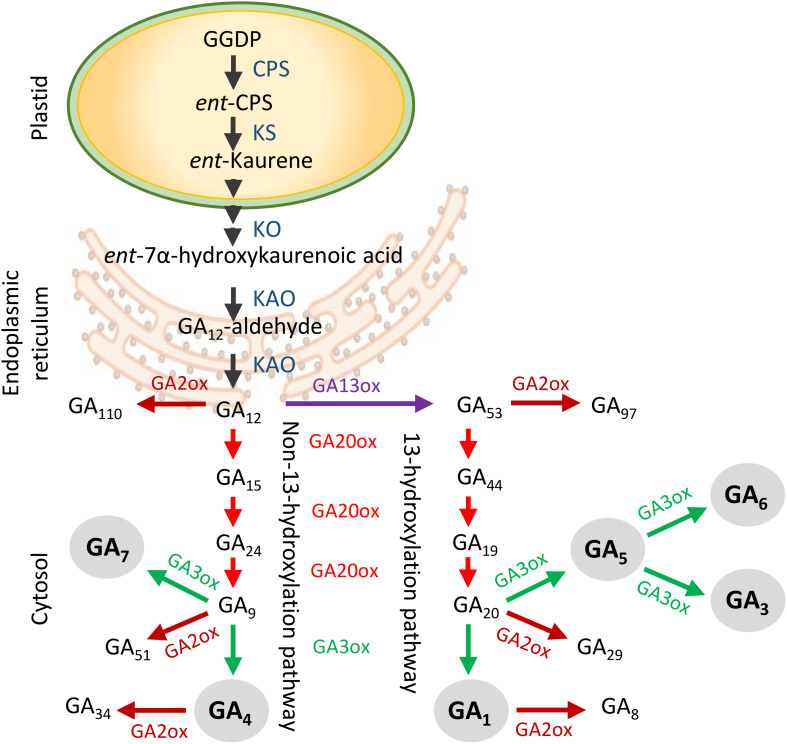
Only a small number of the more than 130 known GAs is biologically active, including GA1, GA3, GA4, GA5, GA6, and GA7 (King et al., 2001, 2003; Yamaguchi, 2008; Hedden and Sponsel, 2015). GA biosynthesis starts with plastid-localized geranylgeranyl diphosphate (GGDP), which is converted to ent-kaurene (Figure 1; Hedden and Phillips, 2000; Olszewski et al., 2002; Yamaguchi, 2008), and oxidized by cytochrome P450 mono-oxygenase in the endoplasmic reticulum to yield GA12 (Helliwell et al., 2001). From there, metabolites are shuttled through two parallel cytoplasmic pathways, the non-13-hydroxylation and 13-hydroxylation pathway, in which three groups of 2-oxoglutarate-dependent dioxygenases provide catalytic activity (Hedden and Phillips, 2000; Yamaguchi and Kamiya, 2000; Olszewski et al., 2002; Hedden and Thomas, 2012). These include GA20-oxidases (GA20oxs) that produce GA precursors, GA3-oxidases (GA3oxs) that produce bioactive GAs, and GA2-oxidases (GA2oxs) that irreversibly deactivate precursors and bioactive GAs by 2β-hydroxylation (Thomas et al., 1999; Lo et al., 2008; Rieu et al., 2008a). Which of the two pathways is dominant depends on species, developmental stage, and organ type. For example, in rice, GA1 dominates during vegetative growth but during anthesis it is GA4 (Kobayashi et al., 1989; Hirano et al., 2008), whereas in hybrid aspen GA4 regulates shoot elongation (Israelsson et al., 2004) and in Arabidopsis also flowering (Sponsel et al., 1997; Eriksson et al., 2006). GA signaling requires binding to the receptor GIBBERELLIN INSENSITIVE DWARF1 (GID1), which localizes to the cytoplasm and nucleus (Ueguchi-Tanaka et al., 2005; Willige et al., 2007; Hirano et al., 2008; Sun, 2010). Because GA4 has the highest affinity to GID1 (Nakajima et al., 2006), its effective concentration can be low. GA-GID1 binding enhances interaction with growth-repressor DELLA proteins, which are also present in the cytoplasm and nucleus (Sun, 2011; Davière and Achard, 2013). Subsequent interaction with ubiquitin E3 ligase complex SCFSLY1/GID2 leads to ubiquitination and degradation of DELLA (Peng et al., 1997; Silverstone et al., 1998; Bolle, 2004; Ueguchi-Tanaka et al., 2007).
FIGURE 1.
Generalized scheme of gibberellin (GA) biosynthesis and deactivation in higher plants. GA biosynthesis starts in the plastids and is followed by the production of GA12 in the endoplasmic reticulum. In the cytoplasm GA12 is processed by GA20ox and GA3ox enzymes in two separate branches to produce bioactive GAs (gray circles). The non-13-hydroxylation yields GA7 and GA4, whereas the 13-hydroxylation yields GA5, GA6, GA3, and GA1. The GA2ox enzymes deactivate precursors and bioactive GAs. Abbreviations: GGDP, geranylgeranyl diphosphate; CPS, ent-copalyl diphosphate synthase; KS, ent-kaurene synthase; KO, ent-kaurene oxidase; KAO, ent-kaurenoic acid oxidase.
It is uncertain if the herbaceous branching models can be transferred directly to woody perennials, considering their different shoot size, lifespan, and AXB composition. In hybrid aspen nodal bark tissue might contribute to the regulation of AXB behavior, perhaps compensating for the inefficiency of long-distance transfer of root-produced strigolactone precursors (Katyayini et al., 2019). In hybrid aspen, AXBs are elaborate structures with sturdy scales that enclose a dwarfed embryonic shoot (ES) that arises over a developmental time span of 10 to 12 plastochrons (Rinne et al., 2015). However, deciduous perennials can show strikingly distinct branching styles, suggesting that even within them regulation of AXB outgrowth can differ. In sylleptic species, AXBs grow out in the same season, producing plastic branching patterns in response to environmental conditions (Wu and Stettler, 1998; Wu and Hinckley, 2001), whereas in proleptic species AXBs do not grow out in the same year (Hallé et al., 1978; Barthélémy and Caraglio, 2007). In hybrid aspen, AXBs cease development at the bud maturation point (BMP) and remain inactive until the next growing season (Paul et al., 2014). The AXBs can therefore be viewed as containing side shoots in which phytomer development is temporarily decoupled from stem elongation, which is postponed until the next growing season. In spring, the elongating stem of the ES telescopes out of the opening bud, allowing subsequent neoformation of leaves. Despite being locked in a developmentally quiescent state, the current year AXBs have a high potential for outgrowth, as shoot decapitation induces rapid outgrowth.
Previous analyses of several GA pathway genes in hybrid aspen suggested that GA-deficiency could explain the dwarfed nature of the ES, and that GA biosynthesis would be required for decapitation-induced elongation (Rinne et al., 2015, 2016). ES elongation might require GA4 to regulate cell division and cell stretching, and to recruit GA4-inducible 1,3-β-glucanases that optimize symplasmic conduits for nutrient and sugar import (Rinne et al., 2011, 2016). While different GA forms can have different developmental effects, the basis of this has not been investigated. To our knowledge, it has remained unknown which GAs play a role during AXB quiescence and branching in hybrid aspen as well as other woody perennial species. The relative prominence of AXBs in hybrid aspen permitted us to carry out comprehensive analyzes of GA metabolite levels and GA-pathway transcripts.
The results support a novel paradigm of a dual role of GA in shoot branching, in which GA3/6 and GA1/4 have opposing tasks. AXBs produce GA3/6 to maintain quiescence by upregulating GA2ox genes, which deactivate GA1/4, keeping their levels low despite ongoing biosynthesis. AXB activation, in turn, is achieved by the instantaneous and strong downregulation of the GA2ox genes, boosting GA1/4-induced signaling. Subsequent elongation is followed by GA1/4 biosynthesis through GA3ox2 and supported by GA precursor import from the node.
Results
To understand the role of GA in shoot branching, we mapped the expression of all GA pathway genes in the major parts of intact plants, and in decapitation activated AXBs and associated nodes. The data were combined with analyses of GA intermediates and bioactive GAs. As GA and SL are thought to have opposite effects on AXB activation, we investigated how feeding of GA3, GA4 and the synthetic SL analog GR24 into AXBs of single-node cuttings influenced the expression of GA and SL pathway genes.
GA20oxs and GA3oxs Expression Is Organ- and Development-Related
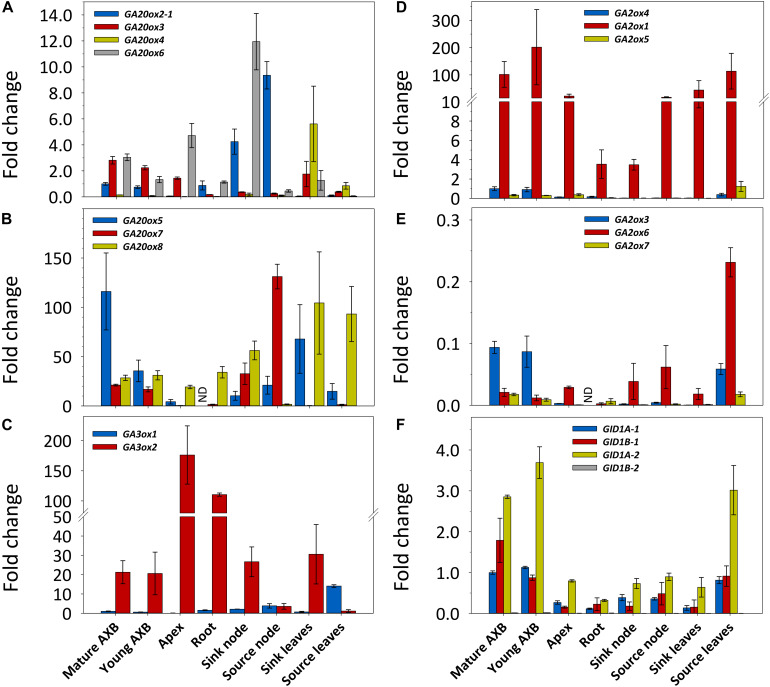
The genome of P. trichocarpa contains eight GA20ox and three GA3ox genes (Tuskan et al., 2006; Figure 1 and Supplementary Figure S1), but preliminary studies showed that GA20ox2-2 was not expressed in hybrid aspen. Transcripts of the seven remaining GA20ox genes were present in young (developing) and mature (developmentally quiescent) AXBs (Figure 2). In decreasing order, the highest transcript levels were found for GA20ox5, GA20ox8 and GA20ox7, whereas GA20ox6, GA20ox3 and GA20ox2-1 were little expressed, and GA20ox4 only in leaves (Figures 2A,B). Of the highly expressed genes of this family, GA20ox8 was the most generally expressed, but transcript levels were especially high in leaves. Whereas in bark tissue of nodes associated with sink leaves (denoted sink nodes) GA20ox8 expression was high, it was almost completely absent in bark tissue of nodes at source leaves (denoted source nodes). Except for the AXBs, all other plant parts expressed GA20ox genes selectively, suggesting the various paralogs might have tissue-specific roles. That all GA20ox family genes were expressed in AXBs makes sense as AXBs harbor a complete, albeit dwarfed shoot system. Combining the transcript levels of all GA20ox paralogs showed that GA-precursor production was highest in sink leaves, followed by source nodes and associated AXBs. In contrast, roots and apices had low transcript levels (Figures 2A,B). Although transcript levels in young AXBs were approximately half of those in the mature quiescent AXBs, they were still almost three times higher than in apices and root tips.
FIGURE 2.
Expression of gibberellin (GA) biosynthesis, deactivation and signaling genes in different plant parts in hybrid aspen. Relative expression (fold change) of GA20ox (A,B), GA3ox (C), GA2ox (D,E), and GID1 (F) family genes. The two larger gene families are depicted in two separate graphs with high (B,D) and little (A,E) expressed genes. Values represent the means of three biological replicates ± S.E. (n = six plants). ND, not detected. Fold changes are relative to reference gene expression in quiescent AXBs, set to 1. A moderately expressed gene within each family was selected for comparison: GA20ox2-1, GA3ox1, GA2ox4, and GID1A-1.
Mature as well as young developing AXBs expressed GA3ox1 and GA3ox2, but transcript levels of GA3ox1 were significantly lower than those of GA3ox2 (Figure 2C and Supplementary Figure S1). In apices, GA3ox1 was virtually absent, whereas it increased in nodes and leaves during their maturation, reaching the highest levels in source leaves. In stark contrast, the expression of GA3ox2 was very high in proliferating shoot apices, and high in growing root tips, sink nodes, sink leaves and AXBs. In the mature nodes and leaves GA3ox2 expression was considerably reduced. The expression ratio of GA3ox2/GA3ox1 showed that apices had the highest approximate ratio (1000), followed by tissues in sinks (20) and sources (0.25). Together the results reveal that, rather than being tissue specific, GA3ox1 and GA3ox2 are developmentally regulated, and that their physiological importance is reversed during tissue maturation. Thus, GA3ox2 expression supports cell proliferation and growth, whereas GA3ox1 is dominant in mature tissues.
In summary, the spatio-temporal expression patterns of the GA20ox and GA3ox family members show that source nodes and source leaves might stockpile GA precursors for delivery to AXBs, while AXBs themselves can produce precursors as well as bioactive GAs.
GA2ox Gene Expression Is Highest in AXBs and Source Leaves
In P. trichocarpa, the GA-deactivating GA2ox family is composed of seven genes (Gou et al., 2011; Supplementary Figure S1). GA2ox2 was not expressed at measurable amounts in shoot tissues of hybrid aspen (not shown) and therefore was not included in the analyses. In decreasing order, the highest transcript levels were found for GA2ox1, GA2ox4, GA2ox5, GA2ox6, GA2ox3, and GA2ox7 (Figures 2D,E). In all plant parts, GA2ox1 had by far the highest transcript levels of the entire GA2ox family. The little expressed genes, GA2ox5 and GA2ox6, were most highly expressed in source leaves. AXBs and source leaves stood apart by expressing most genes, and having the highest combined expression levels, around six times more than apices. Notably, the actively growing tissues, including apices, sink nodes and sink leaves, which expectedly are most active in GA signaling, all expressed GA2ox genes at a low level.
GID1 Receptor Gene Expression Is Highest in AXBs
We identified in hybrid aspen all four paralogs of the P. trichocarpa GID1 genes, and named them GID1A-1, GID1A-2, GID1B-1, and GID1B-2 (Supplementary Figure S2). In shoot tissues, transcript levels of GID1A-2 were the highest, followed by GID1B-1 and GID1-A1, whereas expression of GID1B-2 was very low (Figure 2F). The combined transcript levels of GID1 genes were clearly highest in AXBs and source leaves. In contrast, the expression was low in strong sinks, including proliferating apices, growing root tips, sink nodes and sink leaves. As growing tissues, but especially apices, expressed high levels of the proliferation-related GA-biosynthesis gene GA3ox2 (Figure 2C), the lower GID1 expression levels are expected to reflect high levels of bioactive GAs because receptor abundance correlates negatively with GA levels (Middleton et al., 2012).
GA20ox and GA3ox Genes Are Not Early Activators of AXBs
Even though AXBs in hybrid aspen become quiescent when they reach the BMP, they maintained elevated transcript levels of GID1 receptor genes (Figure 2F), indicating that they remain highly sensitive to GA even after completing development. Despite this, AXBs forestall outgrowth, likely through high expression of GA2ox genes to neutralize GA biosynthesis (Figures 2D,E).
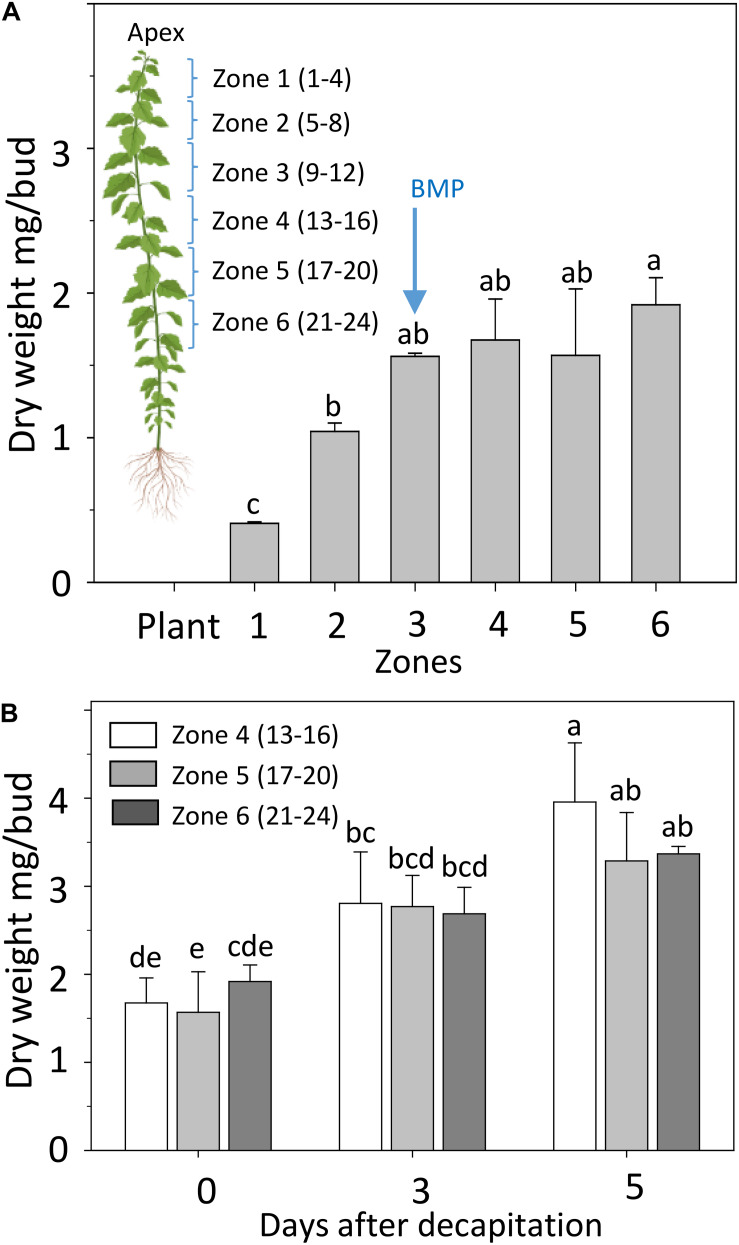
To investigate if and how deactivation and GA biosynthesis changes during AXB activation and outgrowth we decapitated plants at the BMP, recorded the growth of the proximal AXBs over a 5-day period, and analyzed the changes in gene expression that occurred during the critical first 48 h. The BMP has been assessed before, based on the number of embryonic leaves (Rinne et al., 2015). Here, we determined the AXB growth by monitoring the dry weight increment of AXBs along the stem. A plateau in weight gain was reached at the end of zone 3, which is around AXB 12 (Figure 3A), in agreement with the earlier assessment based on embryonic leaf number (Rinne et al., 2015). Precise weight measurements revealed that decapitation not only significantly increased the weight of the proximal AXBs (zone 4), but also of the lower AXBs (zone 5 and 6), showing that all AXBs were activated (Figure 3B). Nonetheless, only the uppermost AXBs (zone 4) grew out, indicating that AXB activation is distinct from outgrowth, as suggested earlier (Cline, 1997).
FIGURE 3.
Development of AXBs. (A) AXBs in different zones along the stem in intact plants. The numbers in parenthesis of zones indicate the position of AXBs, counted from the top. (B) AXB enlargement after decapitation in the remaining zones 4–6. Values represent the means of three biological replicates ± S.E. (n = six plants). One-way ANOVA. Different letters indicate statistical differences between samples (Fischer’s LSD post hoc analysis; P-value at least < 0.05).
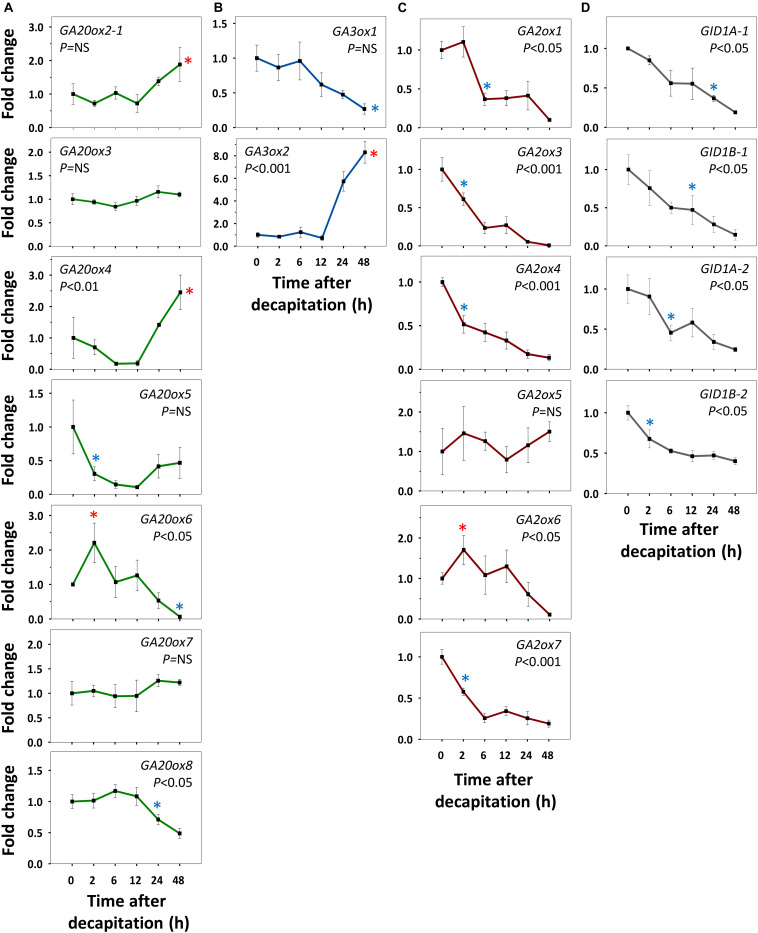
Refining our previous suggestion (Rinne et al., 2016), we show here that net GA-biosynthesis is not the first step in decapitation-induced AXB activation. Although GA20ox6 increased transiently at 2 h, and GA20ox2-1 and GA20ox4 at 48 h, these genes were little expressed in quiescent AXBs compared to GA20ox5 and GA20ox8, which significantly decreased by 2 and 24 h, respectively (Figure 4A). Strikingly, the proliferation-related gene GA3ox2, serving de novo biosynthesis of GA, became significantly upregulated only between 12 and 24 h, in parallel with the downregulation of maturation-related GA3ox1 (Figure 4B). In brief, de novo GA biosynthesis by GA20ox and GA3ox genes is not the initial factor that triggers AXB activation.
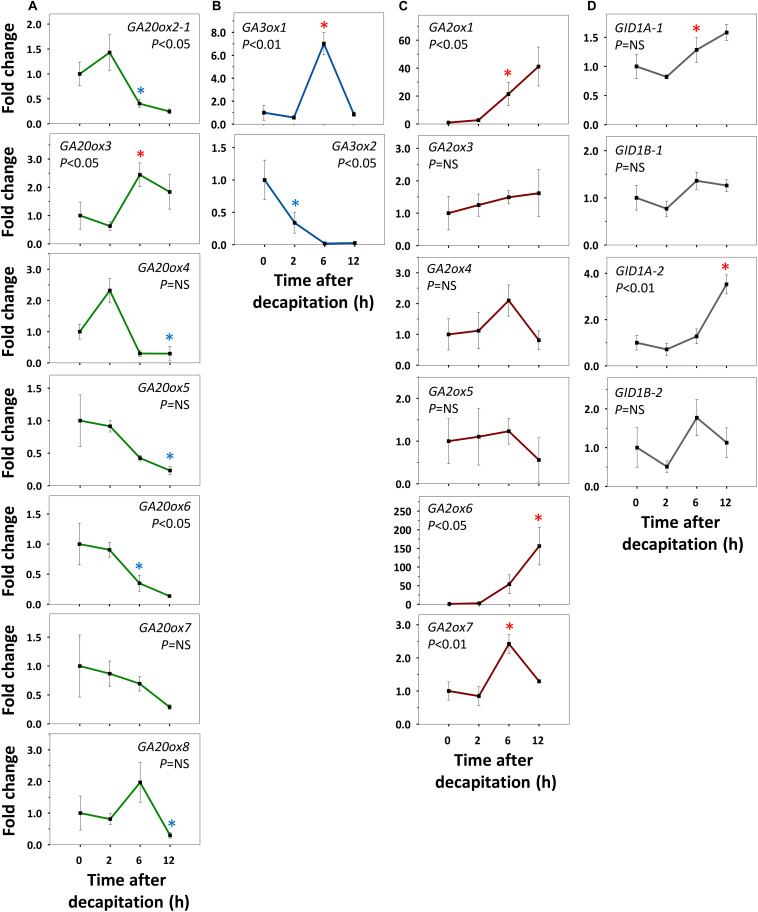
FIGURE 4.
Expression of gibberellin (GA) biosynthesis, deactivation and signaling genes in AXBs after activation by decapitation. (A) GA biosynthesis gene families encoding GA20-oxidases and (B) GA3-oxidases. (C) GA-deactivation gene family encoding GA2-oxidases. (D) GID1-receptor genes. Relative expression (fold change) was analyzed at indicated times after decapitation in three successive AXBs proximal to the decapitation point. Values represent the means of three biological replicates ± S.E. (n = six plants). One-way ANOVA (P-value; NS, not significant). Asterisk indicates the first significant decrease (blue) or increase (red), in gene expression in comparison to time 0 (Fischer’s LSD post hoc analysis, P-value at least < 0.05).
GA2ox Genes Are Early Responders During AXB Activation
All AXBs of intact plants expressed the GA-deactivating GA2ox genes, some at relatively high or very high levels (Figures 2D,E), but decapitation significantly downregulated them within a few hours (Figure 4C). This represented the first change induced by decapitation. The highly expressed gene GA2ox1 was strongly downregulated between 2 and 6h post-decapitation, whereas the little expressed genes GA2ox3, GA2ox4, and GA2ox7 were downregulated even earlier (Figure 4C). The remaining two little-expressed genes GA2ox6 and GA2ox5 responded later or not at all. This shows that the considerable levels of GA3ox1 and GA3ox2 expression in quiescent AXBs were counteracted by the high levels of GA2ox1 expression. In other words, deactivation neutralizes biosynthesis in quiescent AXBs, whereas decapitation increases bioactive GAs by strongly reducing GA deactivation (Figure 4C). The significant parallel reduction in the expression of the GID1 genes (Figure 4D) supports this conclusion, as it is well-known that transcription of GID1 is reduced when levels of bioactive GAs rise (Middleton et al., 2012). That the expression of GA20ox genes did not increase in AXBs after decapitation, while the expression of GA3ox2 was significantly elevated at 24 h, may indicate that additional GA precursors arrived from the nodes. In support of this, expression of GA20ox2-1, GA20ox5 and GA20ox7 in source nodes was high (Figures 2A,B), and decapitation transiently upregulated GA20ox2-1, GA20ox3, GA20ox4, and GA20ox8 (Figure 5A). The putative pool of precursors in the nodes is unlikely to serve the production of bioactive GA in the source node itself, because the proliferation-related gene GA3ox2 was little expressed, and further downregulated 2 h post-decapitation (Figure 5B). Although the maturation-related gene GA3ox1 was transiently upregulated in source nodes between 2 and 6 h (Figure 5B), this was offset by the dramatic upregulation of GA2ox1 and GA2ox6, the two major deactivating genes, as well as the little-expressed gene GA2ox7 (Figure 5C). Moreover, the expression of the GID1 receptor genes tended to increase in the nodes, suggesting a reduction in bioactive GA levels. Notably, the expression patterns of GA-biosynthesis, GA-deactivation and GID1 receptor genes were almost opposite in nodes and activated AXBs (Figures 4, 5).
FIGURE 5.
Expression of gibberellin (GA) biosynthesis, deactivation and signaling genes in nodal bark after decapitation. (A) GA biosynthesis gene families encoding GA20-oxidases and (B) GA3-oxidases. (C) GA-deactivation gene family encoding GA2-oxidases. (D) GID1-receptor genes. Relative expression (fold change) was analyzed at indicated times after decapitation in three successive nodes proximal to the decapitation point. Values represent the means of three biological replicates ± S.E. (n = six plants). One-way ANOVA (P-value; NS, not significant). Asterisk indicates the first significant decrease (blue) or increase (red), in gene expression in comparison to time 0 (Fischer’s LSD post hoc analysis, P-value at least < 0.05).
Collectively, the results support the idea that nodal bark acts as a storage of GA precursors. The time frame of the events suggests that AXB activation is based on diminished deactivation of bioactive GAs in AXBs, making them available for GA signaling, whereas outgrowth relies on biosynthesis, assisted by delivery of node-produced GA precursors.
Xylem-Fed GA3, GA4, and GR24 Modulate GA- and SL-Pathways
Although often functioning redundantly, GA3 and GA4 are produced in separate biosynthetic branches. A biologically meaningful distinction is that GA4 is deactivated by GA2oxs, whereas GA3 is protected by a double bond at the C2, preventing 2β-hydroxylation (Nakayama et al., 1990). In hybrid aspen, GA4 application to dormant AXBs triggers outgrowth, whereas GA3 fails to do so, and a high concentration induces AXB abscission (Rinne et al., 2011). Another factor that affects AXB activation is SL, which acts as an inhibitor of outgrowth in hybrid aspen (Katyayini et al., 2019).
To investigate possible interference of these three hormone pathways, we fed them separately into single-node cuttings, monitored AXB behavior, and analyzed the expression of GA- and SL-pathway genes. As the simple act of isolating the single-node cuttings already activates the AXBs, these experiments test possible interference during AXB elongation. Because preliminary tests with 1% methylene blue showed that it took more than 24 h before dye entered AXBs (not shown), the analyses were carried out at day 3 and day 5, well within the AXB elongation phase. AXB outgrowth tests showed that feeding of a relatively high concentration of GR24 neither inhibited nor promoted AXB burst, relative to the controls, while GA4 significantly accelerated it, and GA3 induced AXB abscission (Supplementary Figure S3). Gene expression analyses of AXBs showed that at the 3 d time point GA20ox genes were downregulated by both GA3 and GA4, except for the unresponsive GA20ox3, and there was no clear difference between the effects of GA3 and GA4 (Figure 6A). At the 5 d time point, the downregulated GA20ox2-1, GA20ox4, GA20ox6, and GA20ox7 were upregulated by both GA3 and GA4. GA20ox8 was unique in that it remained completely unaffected. Notably, it was downregulated by decapitation (Figure 4A). Overall, GA feeding showed that the expression of most GA20ox genes was under strong homeostatic control. Contrary to the downregulating effect of GA3 and GA4 at the 3 d time point, GR24 feeding tended to upregulate the expression of several GA20ox genes (Figure 6A) whereas at the 5 d time point GA20ox6, and GA20ox7 were significantly upregulated, similarly to GA3 and GA4.
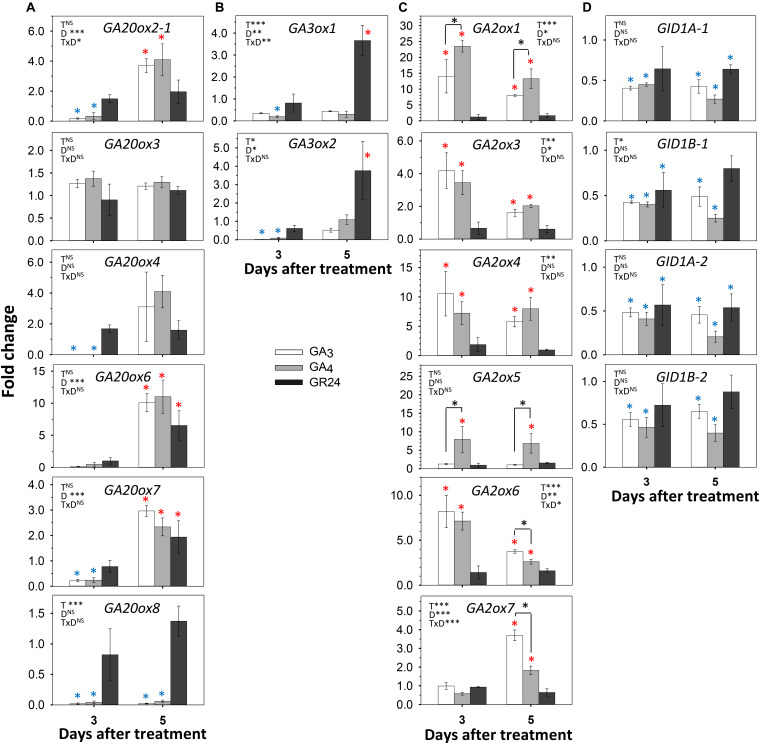
FIGURE 6.
Effect of GA3, GA4, and GR24 on expression of GA-pathway genes in AXBs. AXBs on single-node cuttings were fed with or without GA3, GA4 and GR24 at a concentration of 10 μM. (A) GA20-oxidase genes. (B) GA3-oxidase genes. (C) GA2-oxidase genes. (D) GID1 genes. Values are calculated relative to control and represent the means of three biological replicates ± S.E. (n = six plants). The significance of factors in two-way ANOVA (T, treatments; D, duration in days; TxD, interaction) are indicated by asterisk(s) (*P < 0.05; **P < 0.01; and ***P < 0.001). Asterisks above the bar indicate decrease (blue) or increase (red), relative to control, and above the hook differences between GA3- and GA4-treatments (Fischer’s LSD post hoc analysis; P-value at least < 0.05).
Of the GA3ox family genes, GA3ox1 was significantly downregulated in AXBs by both GA3 and GA4 (Figure 6B). That GA3ox1 expression remained low during the entire period was expected, as it was downregulated by decapitation and did not play a role in AXB activation (Figures 4B, 5B). In contrast, GA3ox2, which is characteristically expressed in proliferating apices and upregulated in activated AXBs (Figures 2C, 4B), was very strongly downregulated at day 3, although it recovered at day 5 (Figure 6B). Overall, GA feeding showed that GA3ox genes, especially GA3ox2, were homeostatically controlled. GR24 did not initially affect the expression of GA3ox genes, but at day 5 it significantly increased the expression of the maturation-related GA3ox1 as well as the proliferation-related GA3ox2.
The GA-deactivating GA2ox genes were strongly upregulated by both GAs. The GA4-induced upregulation of the major GA2ox1 gene was almost 25-fold at day 3, while GA3 was less effective (Figure 6C). In most cases, the expression levels decreased somewhat at day 5. However, GA2ox5 expression continued to rise during GA4 feeding, while this gene was unresponsive to GA3. In contrast, the minor genes GA2ox6 and GA2ox7, were more responsive to GA3 than to GA4 at day 5. The significant upregulation of GA2ox genes indicates that both GAs were effective, whereas GR24 had no effect, suggesting that GR24 does not promote GA deactivation in activated AXBs. GID1 genes were significantly downregulated by GA3 and GA4 (Figure 6D). Interestingly, GR24 also reduced GID1 expression almost to the same degree as the GAs, probably because it upregulated many GA biosynthesis genes (Figures 6A,B).
To assess if the reverse could also be the case, we tested how GA3 and GA4 affected expression of SL pathway genes (Figure 7). In the SL pathway, the gene MAX1 encodes an enzyme that converts plastid-produced carlactone to the SL precursor carlactonoic acid (Abe et al., 2014). Of the two hybrid aspen paralogs MAX1.1 and MAX1.2, the gene MAX1.2 was downregulated by GA3 and GA4, especially by GA3, both at day 3 and 5, whereas MAX1.1 was downregulated only by day 5 (Figure 7A). The genes that encode the SL receptor, D14a and D14b, were strongly upregulated by GA3 and GA4, while MAX2a and MAX2b were moderately upregulated at day 3 (Figures 7B,C). Together this indicates that the GA-induced reduction of SL levels caused upregulation of D14 and MAX2 signaling genes through SL homeostasis. This might have transiently increased expression of BRC1, a downstream target of SL. In contrast, BRC2 was slightly downregulated by both GAs (Figure 7D).
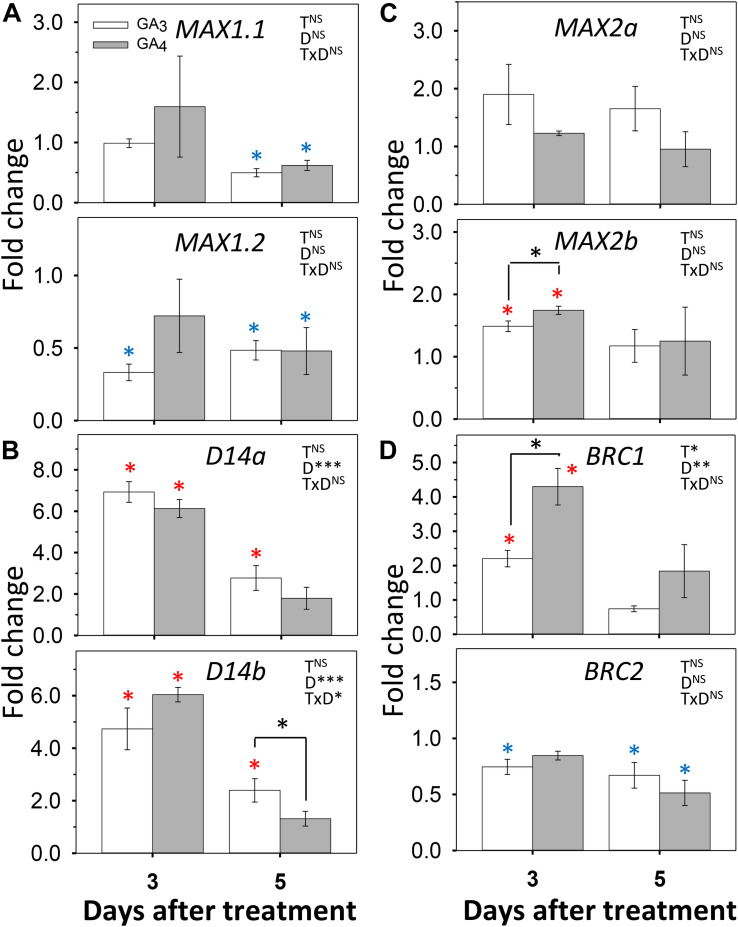
FIGURE 7.
Effect of GA3 and GA4 on selected SL-pathway genes in AXBs. AXBs on single-node cuttings were fed with or without GA3 and GA4 at concentration of 10 μM. Gene expression was analyzed after 3 and 5 days of treatment. (A) MAX1.1 and MAX1.2. (B) D14a and D14b. (C) MAX2a and MAX2b. (D) BRC1 and BRC2. Values are calculated relative to control and represent the means of three biological replicates ± S.E. (n = six plants). The significance of factors in two-way ANOVA (T, treatments; D, duration in days; TxD, interaction) are indicated by asterisks (*P < 0.05; **P < 0.01; and ***P < 0.001). Asterisks above the bar indicate decrease (blue) or increase (red), relative to control, and above the hook differences between GA3- and GA4-treatments (Fischer’s LSD post hoc analysis; P-value at least < 0.05).
AXB Activation Increases the Ratio of GA4/GA1 to GA3/GA6
Gibberellin metabolites, precursors and bioactive molecules in intact and decapitated plants were analyzed using an establish method (Urbanová et al., 2013). This revealed the presence of spatio-temporal patterns in apices and AXBs of distinct zones along the stem (Figure 8). A notable finding was that apices contained bioactive GA of both branches of the GA pathway, although GA1 was the dominant bioactive GA in apices, and levels of GA4, GA5, and GA7 were significantly lower, at least by a factor 20. GA6 was hardly detectable in apices, whereas GA3 was below the detection limit of the LC-MS/MS method used. Although GA1 and GA4 levels were higher in apices than AXBs, these differences were not reflected at the level of precursors. In the case of GA4, its immediate precursor, GA9 was under the detection limit in apices, in contrast to GA24, which was present at high levels. This could indicate that the pool of GA9 is very small due to its rapid conversion to GA4, GA7 and the deactivation product GA51. The GA20ox that produces GA9 from GA24 could therefore be a rate-limiting enzyme in apices, but not in AXBs where these genes were well expressed. GA1 levels in apices were about 40 times higher than the levels of its precursor GA20, even though GA1 was strongly deactivated to GA8. This suggests that the GA20 pool is in a state of rapid flux in apices.
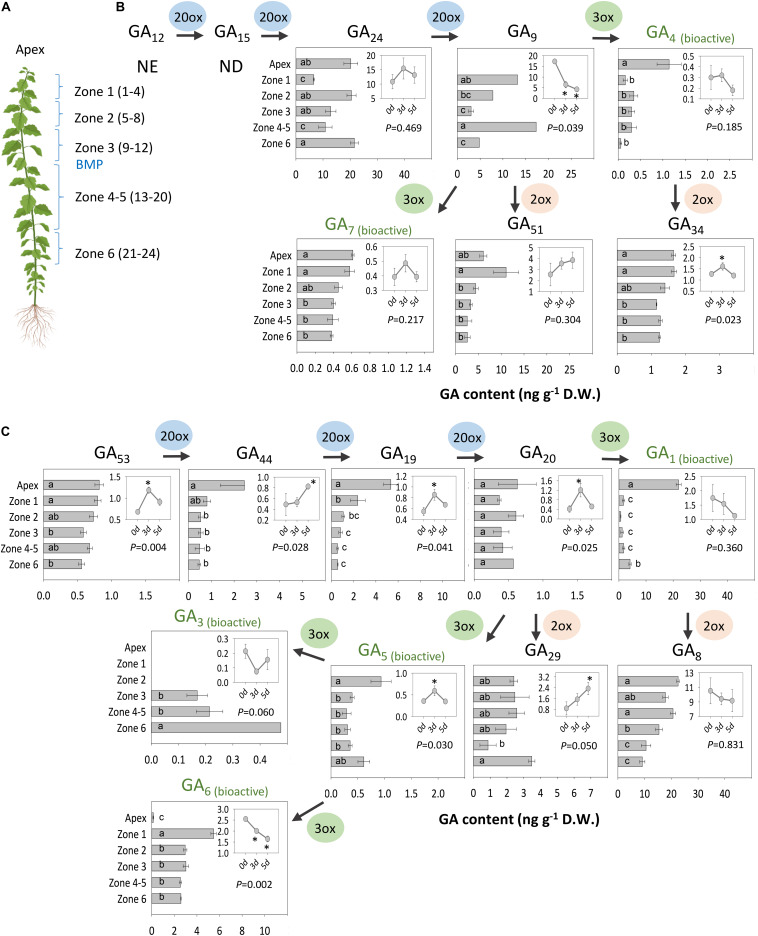
FIGURE 8.
Analysis of GAs. (A) Analyzed materials are indicated. The numbers in parenthesis of each zone refer to the position and the number of AXBs in each sample. GAs of the (B) non-13-hydroxylation and (C) the 13-hydroxylation pathway. Insets: Changes in GA levels 0, 3, and 5 days after decapitation in AXBs proximal to the decapitation point (P-value shown). Asterisks in insets indicate statistically significant change in GA levels. Values represent the means of three biological replicates ± S.E. (n = six plants). Different letters in bars indicate statistical differences in GA levels between the samples. NE, not estimated; ND, not detected. One-way ANOVA and pairwise post hoc analysis by Fischer’s LSD test (P-value at least < 0.05).
In AXBs, GA1 levels were ca.10-fold lower than in apices, while the level of the bioactive GA4 was about 3- to 4-fold lower (Figures 8B,C). AXBs contained a considerable amount of GA6, while GA3 was produced at a much lower level, and only in mature AXBs (mature AXBs in zone 4-5) and aging (oldest AXBs in zone 6) (Figures 8A,C). GA-deactivation was especially prominent in the early 13-hydroxylation pathway, resulting in high levels of GA29 and, especially, GA8. Whereas GA1 content was low in AXBs, its deactivation product GA8, was almost at the same level as in apices. When the GA2ox genes, responsible for this conversion, are abruptly downregulated, as observed after decapitation in AXBs (Figure 4C), GA1 availability is expected to rise. In the non-13-hydroxylation pathway, most GA9 was de-activated to GA51, and comparatively little to the bioactive GA4 and GA7, both in apices and AXBs. Similarly, to GA8, the GA4-deactivation product GA34 was almost the same in AXBs and apices, suggesting that decapitation-induced downregulation of GA2ox expression in AXBs increases GA4 availability.
Shoot decapitation only slightly affected GA content during the AXB elongation phase at 3 d and 5 d post-decapitation (Figures 8B,C, insets). The changes in the 13-hydroxylation pathway (Figure 8C) were more often statistically significant than those in the non-13-hydroxylation pathway (Figure 8B). In the latter, only the deactivation product GA34 increased significantly. In the 13-hydroxylation pathway, GA1 also did not show any increase, even though all its precursors increased at day 3 and 5. The overall increase in precursors (GA53 to GA20) resulted in a significant increase of the deactivation product GA29. GA3 and GA6 were absent from apices, but were detected in AXBs, whereas decapitation lowered their contents, especially that of GA6.
Interestingly, in a separate experiment under suboptimal greenhouse conditions, where plants tended to cease growth, GA20 levels and their deactivation products GA29 and GA8 were higher in apices, while GA1 levels were very low (Supplementary Figure S4). In these plants GA3 was also detectable in apices, while GA5 and GA6 were under the detection limit. This highlights that GA3 is not unique to AXBs per se but can be produced to restrict proliferation.
Discussion
Shoot branching is regulated by a network of inhibitory and promotive forces. The present results obtained by combining gene expression profiling, metabolite quantitation and hormone treatments show that specific GAs promote branching, while others maintain AXBs in a quiescent state.
AXB Activation and Outgrowth Require Diminished GA-Deactivation
The differential expression of GA-pathway genes at the whole plant level appears to reflect the proleptic lifestyle of hybrid aspen, in which AXBs become quiescent once they reach maturity (Rinne et al., 2015). AXBs expressed most GA20ox genes at significantly higher levels than apices (Figures 2A,B). Nonetheless, the levels of bioactive GA1/4 were significantly lower in AXBs than in proliferating apices (Figure 8). The obvious reason for this is that GA2ox genes were strongly expressed in AXBs, about 6-fold relative to apices (Figures 2D,E). As the encoded GA2ox enzymes irreversibly deactivate bioactive GAs by 2β-hydroxylation (Thomas et al., 1999; Olszewski et al., 2002; Middleton et al., 2012), the high level of GA2ox expression in AXBs can keep them quiescent. This is strongly supported by the fact that during AXB activation several GA2ox genes were rapidly and significantly downregulated, and subsequently also the four GID1 receptor genes (Figures 4C,D). This indicates that GA availability had effectively increased because GID1 levels are known to diminish when GA levels increase due to homeostatic adjustment (Gallego-Giraldo et al., 2008; Hedden and Thomas, 2012; Middleton et al., 2012). Because bioactive GA levels reflect the balance between GA biosynthesis and deactivation (Phillips et al., 1995; Xu et al., 1999; Olszewski et al., 2002; Yamaguchi, 2008), the decapitation-induced reduction of GA deactivation increases its availability for signaling, even in the absence of increased biosynthesis.
The emerging picture is that quiescent AXBs are sensitized to GA, because relative to apices they have low levels of GA1 and GA4 despite the ongoing GA biosynthesis, but high levels of GID1 expression. Thus, regardless of GA biosynthesis, the dwarfed ES of AXBs is GA deficient. The high GA2ox expression levels in AXBs appear to be developmentally controlled to keep AXB activation at bay and safeguard the proleptic nature of the shoot system. GA3/6 can play a role in maintaining AXB quiescence (Figure 8C) by upregulating GA2ox genes, thereby deactivating GA1/4, but not of itself (and GA5/6) because it is not a substrate (Nakayama et al., 1990; Ito et al., 2017; Li et al., 2017). Thus, the specific presence of GA3/6 in quiescent AXBs can effectively maintain them in a GA4-deficient state. As GA4 is involved in promoting cell division, elongation and energy metabolism (Hedden and Sponsel, 2015; Zhuang et al., 2015) and has the highest binding activity to GID1 (Ueguchi-Tanaka et al., 2005), keeping GA4 low is necessary to prevent AXB activation and outgrowth. In addition, other factors may play a role in AXB quiescence, including SL (Katyayini et al., 2019) and BRC1-regulated ABA signaling (González-Grandío et al., 2017; Wang et al., 2019). After AXB activation, subsequent AXB elongation is supported by de novo biosynthesis of GA1 and GA4, initiated between 12 and 24 h through upregulation of GA3ox2 (Figure 4B). In support of this, a previous study showed that this gene, originally named GA3ox1, is characteristically expressed in growing shoot apices (Israelsson et al., 2004). In short, our data support a model in which branching is initiated by a strong reduction of GA deactivation that raises the bioactive GA1/4 pool to spearhead AXB activation, while additional GA1/4 biosynthesis supports subsequent AXB elongation, as illustrated in Figure 9.
FIGURE 9.
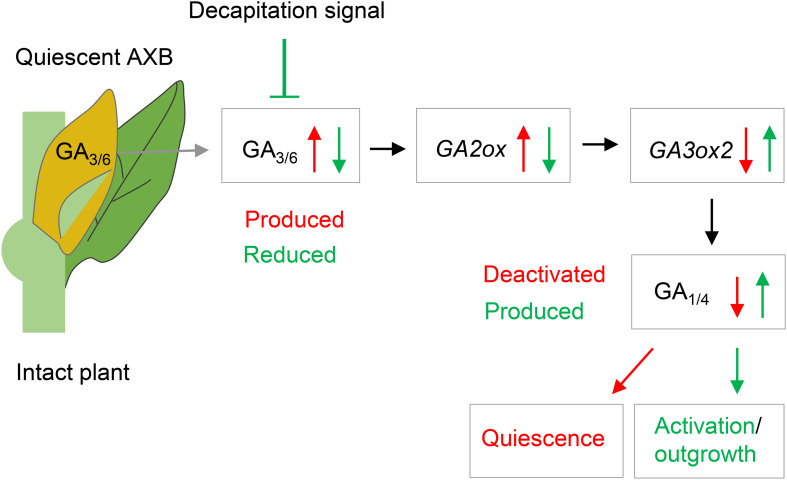
Model of axillary bud (AXB) quiescence and activation. During their formation, AXBs accumulate GA3/6, which upregulates GA2ox gene expression. GA2ox deactivates GA1/4, thereby depriving the AXBs of GA1/4-mediated signaling. Decapitation activates quiescent AXBs by rapidly reducing GA3/6 levels and downregulating GA2ox transcription, thereby elevating GA1/4 levels and signaling. Subsequently, the GA1/4 biosynthesis gene GA3ox2 is upregulated dramatically increasing GA levels and promoting AXB outgrowth. Red arrows and text indicate AXB inhibitory effects. Green arrows and text indicate AXB activating and growth promoting effects.
GA Biosynthesis Differs in Growing and Mature Tissues
The expression patterns of the GA biosynthesis genes were different for actively proliferating tissues (apices and roots), differentiated tissues (mature leaves), and developmentally inactive tissues with high growth potential (AXBs) (Figure 2). For example, apices expressed GA20ox genes less than other tissues, but they highly expressed GA3ox2, whereas GA3ox1 was hardly expressed. In contrast, quiescent AXBs expressed both GA3ox genes, whereas source leaves exclusively expressed GA3ox1 genes. Thus, GA3ox2 supports cell proliferation and growth at apices and root tips, whereas GA3ox1 reflects tissue maintenance in source nodes and leaves. The fact that quiescent AXBs expressed both GA3ox2 and GA3ox1 appears to reflect their opposing developmental tendencies, as AXBs combine developmental stasis with high growth potential. As indicated above, the high levels of GA deactivation, maintained by the GA2ox-insensitive GA3/6, are likely to be part of the developmental block that prevents AXB activation.
Although AXBs expressed all GA-pathway genes, their outgrowth is strongly dependent on a functional connection to the stem, especially nodal vascular tissue. The results suggest that nodal bark exported precursors to AXBs, because the GA20ox transcript levels in the AXBs were reduced soon after decapitation, whereas in the nodal bark they initially increased without increasing GA3ox2 expression (Figures 5A,B). Transport of precursors and bioactive GAs (GA3, GA4, GA9, GA12 and GA20) is known to be crucial in directing development (Proebsting et al., 1992; Eriksson et al., 2006; Yamaguchi, 2008; Ragni et al., 2011; Dayan et al., 2012; Lange and Lange, 2016; Regnault et al., 2016; Binenbaum et al., 2018). The GA quantitation data support the idea that precursors are transported from nodes to the AXBs, as their levels increased in AXBs after decapitation, for example in case of GA20, a key precursor of several bioactive forms of GA (Figure 8C). Such node-to-AXB delivery also plays a role in the SL-mediated control of AXB quiescence (Katyayini et al., 2019). Together, the analyses indicate that nodal bark tissue might affect AXBs by delivering SL and GA precursors.
GA and SL Pathways Are Buffered and Show Interference
During the AXB elongation phase, GA2ox genes responded strongly to GA feeding by upregulating their expression up to ≥20-fold at day 3. As the GID1 expression levels were only reduced by about 2-fold, the upregulated GA2ox must have been effective in deactivating part of the supplied GA. Feeding GR24 did not affect the expression of GA2ox genes, but it did increase the expression of GA biosynthesis genes at day 5 (Figures 6A,B). A putative increase in GA levels by GR24 could explain why GR24 feeding reduced GID1 expression levels to a similar degree as GA3 and GA4 (Figure 6D).
In hybrid aspen, SL pathway and perception genes are highly expressed in quiescent AXBs, but decapitation rapidly downregulated these genes as well as the downstream target gene BRC1 (Katyayini et al., 2019). While GA3/6, GA2ox as well as SL contribute to the quiescent state of AXBs in intact plants, their decrease in activated AXBs leads to elevated GA1/4 levels through a reduction of GA2ox activity. Subsequent outgrowth might require CK in addition (Ni et al., 2017; Duan et al., 2019).
As feeding GA3 and GA4 reduced the expression of both MAX1 genes (Figure 7A), GA represses SL biosynthesis, which supports earlier observations in other plant species (Ni et al., 2015; Ito et al., 2017; Marzec, 2017). Our data show that during the AXB elongation phase both GA3 and GA4 increased SL perception by upregulating D14 genes and MAX2b (Figures 7B,C). This increase in SL perception and signaling genes presumably is a homeostatic response to a GA-induced reduction in SL levels in the AXBs. In Arabidopsis, GA and GR24 converge on a large number of shared transcription targets (Lantzouni et al., 2017). However, in pea, SL can also independently of GA promote cell division in the stem (de Saint Germain et al., 2013). Here we found that GR24 increased the biosynthesis of GA during the AXB elongation phase. It is noteworthy that GR24 feeding can promote the elongation of the enclosed ES five to seven days post-decapitation (Katyayini et al., 2019), and the present data suggest this might involve GA. Whether these interferences between SL and GA pathways are direct or indirect remains to be established.
GA3 and GA6 Are Involved in AXB Development but Not in AXBs Outgrowth
In AXBs of intact plants, the gene GA3ox1 could be linked to presence of GA3 and GA6. After decapitation, GA3ox1 expression and GA3 and GA6 content decreased in AXBs (Figures 4B, 8C) and were absent from apices (Figure 2C). This indicates that GA3ox1 functions in the side branch of the 13-hydroxylation pathway that produces the deactivation-protected GA3, GA5 and GA6. In contrast, GA3ox2 converts precursors GA9 and GA20 to GA7, GA4 and GA1, in support of a previous study (Israelsson et al., 2004).
In apices GA1 was more abundant than GA4 (Figure 8), although GA4 more efficiently promotes shoot elongation (Israelsson et al., 2004). However, plants can switch between pathways, depending on developmental phase or environmental conditions (Rieu et al., 2008b). For example, in a grass species GA4 is produced during vegetative growth, while upon flowering it switched to GA5 and GA6 (King et al., 2001, 2003). That GA2oxs play a role in this, is supported by studies in Jatropha, where overexpression of GA2ox6 induced a switch from the non-13-hydroxylation pathway (GA4) to the 13-hydroxylation pathway (GA3), and led to dwarfing (Hu et al., 2017). Our data suggest that the GA precursor GA20 can be converted to the growth-promoting GA1 or the quiescence-related GA3/6 (Figure 8C) dependent on developmental cues as well as environmental conditions. GA3 accumulates in developing AXBs as well as in apices of stressed plants, while GA1 levels remain low (Supplementary Figure S4). The effect of these cues on GA metabolism, and the distinct responses of plants to different bioactive GAs (Elfving et al., 2011; Rinne et al., 2011; Ni et al., 2015) warrant further investigation.
Although GA3 is often used as a generic GA, it is different from GA4 in important respects. The results show that in hybrid aspen GA3 and GA4 not only operate at distinct locations, their functions are also partly distinct. GA4 feeding promotes AXB outgrowth, whereas GA3 induces abscission in the non-dormant quiescent AXBs that form under long days (Supplementary Figure S3) as well as the AXBs that establish dormancy under short days (Rinne et al., 2011). GA3 and GA4 also induce different classes of 1,3-β-glucanases, destined for different subcellular locations (Rinne et al., 2011). Both GA3 and GA4 promote cell division, but GA4 function requires histone deacetylases to transcriptionally block GA2ox (Li et al., 2017). Although required for apical growth, in the vegetative in the meristem dome itself GA4 is absent, because its production is blocked, and a band of GA2ox expression below the meristem protects it from a damaging influx of GA4 (Sakamoto et al., 2001; Jasinski et al., 2005; King et al., 2008; Bolduc and Hake, 2009). As GA3 cannot be deactivated by GA2ox, GA3 (as well as GA5 and GA6) can enter the meristem and induce floral transition in grasses, whereas GA4 can only enter later, when the band of GA2ox expression is gone (King et al., 2003).
Because GA3 can significantly upregulate GA2ox genes (Figure 6C), its accumulation in quiescent AXBs results in low levels of GA1/4 due to deactivation, as both are substrates of GA2ox (Nakayama et al., 1990), thereby inhibiting GA4-mediated AXB activation and elongation. Our finding that GA3/6 were detected in quiescent AXBs and reduced by decapitation, matches our earlier finding that GA3, unlike GA4, cannot upregulate the growth-related α-clade 1,3-β-glucanases that optimize symplasmic conduits for transport to growing areas (Rinne et al., 2011).
Conclusion
A major finding was that hybrid aspen invests energy into producing and simultaneously deactivating GA1/4 in quiescent AXBs, although they remain developmentally inactive until the next year. This seemingly wasteful strategy is an effective way to keep AXBs ready for rapid outgrowth in case the shoot apex is damaged or lost, allowing a new shoot to form before winter arrives. The results support a model in which SL and GA3ox1-mediated accumulation of GA3/6 maintain AXBs in a quiescent state, with GA3/6 upregulating GA2ox genes that deactivate GA1/4. In turn, decapitation-induced AXB activation is triggered by a rapid downregulation of GA2ox genes, which shifts the balance between GA1/4 biosynthesis and deactivation, increasing the GA1/4 pool available for GA signaling. The initial GA1/4 pulse is followed by increased GA3ox2-mediated de novo GA biosynthesis, and subsequent elongation of the AXB. The dual, opposing roles of GA3/6 and GA1/4 can explain why the role of GA in branching has been ambiguous.
Materials and Methods
Plant Material and Sample Preparation
Hybrid aspen (Populus tremula × Populus tremuloides) clone T89 was micro-propagated in vitro and grown in a greenhouse under long days as previously described (Katyayini et al., 2019). When the plants were 80–100 cm tall, with stable leaf production and elongation rates, they were subdivided into three groups: (a) Intact plants for collection of tissues and organs for transcript analyses; (b) Decapitated plants (decapitated at the bud maturation point, ca. 40 cm below the apex), for transcript and GA analysis in AXBs, and transcript analysis of nodal bark; (c) Plants for xylem feeding of hormones into single-node cuttings. Samples for transcript and hormone analyzes were collected from six plants, with two plants pooled in three replicate samples. Position of sampled buds and tissues is indicated in Supplementary Figure S5.
Quantification of GAs With Liquid Chromatography-Mass Spectrometry (LC-MS/MS)
The samples (apices and AXBs) were harvested from different zones along the stem, as indicated in Figure 3A. For analysis, samples were immediately frozen in liquid nitrogen, and subsequently freeze dried. Sample preparation and quantitative analysis of GAs were performed by LC-MS/MS using 2H2-labeled GA internal standards as described (Urbanová et al., 2013).
AXB Burst Tests and Feeding of GA3, GA4, and GR24
To investigate the effects of GA3, GA4 and the synthetic strigolactone GR24 on AXB outgrowth and gene expression, we performed xylem-feeding experiments under forcing conditions in growth chambers (18 h of light with a PPFD of 160−200 μmol m–2 s–1, 20°C, and 60% relative humidity). Single-node cuttings were isolated from 6-week old plants. The internode base was punched through pores in a Styrofoam sheet, floated on water (control) or water supplemented with GA3, GA4 (Sigma-Aldrich) or racemic synthetic SL GR24 (Chiralix BV, Netherlands) at the effective 10 μM concentration (Katyayini et al., 2019). AXB burst was followed for 14 days and scored as Σ14-values, as explained in Supplementary Figure S3.
Experiment Design and Gene Selection
For analysis of GA-pathway, total RNA was extracted from different plant parts as indicated (Figure 2). Gene expression analysis included hybrid aspen homologs of P. trichocarpa GA-biosynthesis genes GA20ox2-1, GA20ox3, GA20ox4, GA20ox5, GA20ox6, GA20ox7, GA20ox8, GA3ox1, and GA3ox2; GA-catabolism genes GA2ox1, GA2ox3, GA2ox4, GA2ox5, GA2ox6, and GA2ox7; GA-signaling genes GID1A-1, GID1A-2, GID1B-1, and GID1B-2. For phylogenetic analysis, see Supplementary Figures S1, S2.
To assess decapitation-induced expression changes, AXBs proximal to the decapitation point of the BMP were collected 0, 2, 6, 12, 24, and 48 h post-decapitation. Sampling after day 1 and day 2 was carried out at the same time of day to avoid potential diurnal effects on gene expression. Nodal bark tissues were collected 0, 2, 6, and 12 h after decapitation.
The effects of 10 μM GA3, GA4 and GR24 on gene expression in AXBs were investigated after xylem feeding of the hormones into AXBs of single-node cuttings. Samples were collected after 0, 3, and 5 days. Gene expression analysis included GA-biosynthesis GA20ox2-1, GA20ox3, GA20ox4, GA20ox6, GA20ox7, GA20ox8, GA3ox1, and GA3ox2; GA-catabolism genes GA2ox1, GA2ox3, GA2ox4, GA2ox5, GA2ox6, and GA2ox7; GA-signaling genes GID1A-1, GID1A-2, GID1B-1 and GID1B-2. In addition, previously identified SL-biosynthesis and signaling genes MAX1.1, MAX1.2, D14a, D14b, MAX2a, and MAX2b, and the downstream target genes BRC1 and BRC2 (Katyayini et al., 2019) were analyzed after GA3 and GA4 feeding.
RNA Extraction and cDNA Preparation
Total RNA was extracted from 0.2 to 0.3 g of frozen tissue and grinded in a mortar with 500 μL extraction buffer (Qiagen RLT buffer containing 1% PVP-40), and further processed as described (Katyayini et al., 2019). The samples were transferred to RNeasy spin columns and further processed in accordance with instructions of the Qiagen Plant RNA isolation kit. Genomic DNA was eliminated using TURBOTM DNase kit (Invitrogen) treatment according to manufacturer’s instructions and cleaned using the total RNA purification system “Purelink RNA mini kit” (Invitrogen). RNA was quantified with NanoDrop 1000, and the RNA quality was assessed with the Agilent 2100 Bioanalyzer system. 1 μg of total RNA was reversely transcribed to cDNA with SuperScript® VILOTM reverse transcriptase (Invitrogen).
Quantitative RT-PCR (qRT) Analysis
The reaction setup (20 μl total volume) for qRT was prepared using SYBR® select PCR master mix (Applied Biosystems). As a template, 2 μl of the cDNA (200 ng) were added. Real-time qRT-PCR analyses were performed with the Applied Biosystems 7500 Fast Real-Time PCR system according to the manufacturer’s instruction. Thermocycling conditions were set to 50°C for 2 min, 95°C for 2 min, 45 cycles of 15 s at 95°C and 60 s at 60°C. Each PCR reaction included a negative control to check for potential genomic DNA contamination. For a complete list of primers and genes used for quantitative real time PCR (qRT-PCR) see Supplementary Table S1.
Statistical Analysis and Bioinformatics
Statistical analyses were carried out using analysis of variance (one- or two-way ANOVA) in combination with Fisher LSD post hoc test to determine significant differences between the subgroups. Computation was performed using Microsoft Excel data analysis1 and Minitab Statistical Software version 18.1.2
BLAST searches in GenBank, Populus trichocarpa genome v3.0 and Populus tremula × Populus tremuloides (T89) v3.0 databases3, 4, 5 were used to identify GA-biosynthesis, -catabolism and -signaling genes. Gene specific primer sequences for qPCR analysis were designed using Primer3.6
Data Availability Statement
All datasets generated for this study are included in the article/Supplementary Material.
Author Contributions
NK, PR, and CS designed the research. NK, PR, and DT performed the experiments. NK, PR, and CS analyzed and interpreted the data. NK designed the illustration. All authors participated in writing and revising the manuscript.
Conflict of Interest
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Acknowledgments
We thank Ms. Marit Siira for excellent help with the plants, and Dr. Manikandan Veerabagu for discussions.
Funding. The work was funded by a grant of the Norwegian University of Life Sciences to NK, a FRIPRO grant of the Norwegian Research Council (No. 263117), the Czech Science Foundation (No. 18-10349S), and the European Regional Development Fund Project “Centre for Experimental Plant Biology” (No. CZ.02.1.01/0.0/0.0/16_019/0000738).
Supplementary Material
The Supplementary Material for this article can be found online at: https://www.frontiersin.org/articles/10.3389/fpls.2020.00736/full#supplementary-material
References
- Abe S., Sado A., Tanaka K., Kisugi T., Asami K., Ota S., et al. (2014). Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro. Proc. Natl. Acad. Sci. U.S.A. 111 18084–18089. 10.1073/pnas.1410801111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Agharkar M., Lomba P., Altpeter F., Zhang H., Kenworthy K., Lange T. (2007). Stable expression of AtGA2ox1 in a low-input turfgrass (Paspalum notatum Flugge) reduces bioactive gibberellin levels and improves turf quality under field conditions. Plant Biotechnol. 5 791–801. 10.1111/j.1467-7652.2007.00284.x [DOI] [PubMed] [Google Scholar]
- Aguilar-Martínez J. A., Poza-Carrión C., Cubas P. (2007). Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell 19 458–472. 10.1105/tpc.106.048934 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barthélémy D., Caraglio Y. (2007). Plant architecture: a dynamic, multilevel and comprehensive approach to plant form, structure and ontogeny. Ann. Bot. 99 375–407. 10.1093/aob/mcl260 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bennett T., Sieberer T., Willett B., Booker J., Luschnig C., Leyser O. (2006). The Arabidopsis MAX pathway controls shoot branching by regulating auxin transport. Curr. Biol. 16 553–563. 10.1016/j.cub.2006.01.058 [DOI] [PubMed] [Google Scholar]
- Binenbaum J., Weinstain R., Shani E. (2018). Gibberellin localization and transport in plants. Trends Plant Sci. 23 410–421. 10.1016/j.tplants.2018.02.005 [DOI] [PubMed] [Google Scholar]
- Bolduc N., Hake S. (2009). The maize transcription factor KNOTTED1 directly regulates the gibberellin catabolism gene ga2ox1. Plant Cell 21 1647–1658. 10.1105/tpc.109.068221 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolle C. (2004). The role of GRAS proteins in plant signal transduction and development. Planta 218 683–692. 10.1007/s00425-004-1203-z [DOI] [PubMed] [Google Scholar]
- Brewer P. B., Dun E. A., Ferguson B. J., Rameau C., Beveridge C. A. (2009). Strigolactone acts downstream of auxin to regulate bud outgrowth in pea and Arabidopsis. Plant Physiol. 150 482–493. 10.1104/pp.108.134783 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Busov V. B., Meilan R., Pearce D. W., Ma C., Rood S. B., Strauss S. H. (2003). Activation tagging of a dominant gibberellin catabolism gene (GA 2-oxidase) from poplar that regulates tree stature. Plant Physiol. 132 1283–1291. 10.1104/pp.103.020354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choubane D., Rabot A., Mortreau E., Legourrierec J., Péron T., Foucher F., et al. (2012). Photocontrol of bud burst involves gibberellin biosynthesis in Rosa sp. Plant Physiol. 169 1271–1280. 10.1016/j.jplph.2012.04.014 [DOI] [PubMed] [Google Scholar]
- Claeys H., De Bodt S., Inzé D. (2014). Gibberellins and DELLAs: central nodes in growth regulatory networks. Trends Plant Sci. 19 231–239. 10.1016/j.tplants.2013.10.001 [DOI] [PubMed] [Google Scholar]
- Cline M. G. (1991). Apical dominance. Bot. Rev. 57 318–358. 10.1007/bf02858771 [DOI] [Google Scholar]
- Cline M. G. (1997). Concepts and terminology of apical dominance. Am. J. Bot. 84 1064–1069. 10.2307/2446149 [DOI] [PubMed] [Google Scholar]
- Davière J.-M., Achard P. (2013). Gibberellin signaling in plants. Development 140 1147–1151. 10.1242/dev.087650 [DOI] [PubMed] [Google Scholar]
- Dayan J., Voronin N., Gong F., Sun T.-P., Hedden P., Fromm H., et al. (2012). Leaf-induced gibberellin signaling is essential for internode elongation, cambial activity, and fiber differentiation in tobacco stems. Plant Cell 24 66–79. 10.1105/tpc.111.093096 [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Saint Germain A., Ligerot Y., Dun E. A., Pillot J.-P., Ross J. J., Beveridge C. A., et al. (2013). Strigolactones stimulate internode elongation independently of gibberellins. Plant Physiol. 163 1012–1025. 10.1104/pp.113.220541 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Domagalska M. A., Leyser O. (2011). Signal integration in the control of shoot branching. Nat. Rev. Mol. Cell Biol. 12 211–221. 10.1038/nrm3088 [DOI] [PubMed] [Google Scholar]
- Duan J., Yu H., Yuan K., Liao Z., Meng X., Jing Y., et al. (2019). Strigolactone promotes cytokinin degradation through transcriptional activation of CYTOKININ OXIDASE/DEHYDROGENASE 9 in rice. Proc. Natl. Acad. Sci. U.S.A. 116 14319–14324. 10.1073/pnas.1810980116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dun E. A., Brewer P. B., Beveridge C. A. (2009). Strigolactones: discovery of the elusive shoot branching hormone. Trends Plant Sci. 14 364–372. 10.1016/j.tplants.2009.04.003 [DOI] [PubMed] [Google Scholar]
- Dun E. A., Ferguson B. J., Beveridge C. A. (2006). Apical dominance and shoot branching. Divergent opinions or divergent mechanisms? Plant Physiol. 142 812–819. 10.1104/pp.106.086868 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elfving N., Davoine C., Benlloch R., Blomberg J., Brännström K., Müller D., et al. (2011). The Arabidopsis thaliana Med25 mediator subunit integrates environmental cues to control plant development. Proc. Natl. Acad. Sci. U.S.A. 108 8245–8250. 10.1073/pnas.1002981108 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Eriksson S., Böhlenius H., Moritz T., Nilsson O. (2006). GA4 is the active gibberellin in the regulation of LEAFY transcription and Arabidopsis floral initiation. Plant Cell 18 2172–2181. 10.1105/tpc.106.042317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferguson B. J., Beveridge C. A. (2009). Roles for auxin, cytokinin, and strigolactone in regulating shoot branching. Plant Physiol. 149 1929–1944. 10.1104/pp.109.135475 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallego-Giraldo L., Ubeda-Tomas S., Gisbert C., García-Martínez J. L., Moritz T., López-Díaz I. (2008). Gibberellin homeostasis in tobacco is regulated by gibberellin metabolism genes with different gibberellin sensitivity. Plant Cell Physiol. 49 679–690. 10.1093/pcp/pcn042 [DOI] [PubMed] [Google Scholar]
- González-Grandío E., Pajoro A., Franco-Zorrilla J. M., Tarancón C., Immink R. G. H., Cubas P. (2017). Abscisic acid signaling is controlled by a BRANCHED1/HD-ZIP I cascade in Arabidopsis axillary buds. Proc. Natl. Acad. Sci. U.S.A. 114 E245–E254. 10.1073/pnas.1613199114 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gou J., Ma C., Kadmiel M., Gai Y., Strauss S., Jiang X., et al. (2011). Tissue-specific expression of Populus C19 GA 2-oxidases differentially regulate above-and below-ground biomass growth through control of bioactive GA concentrations. New Phytol. 192 626–639. 10.1111/j.1469-8137.2011.03837.x [DOI] [PubMed] [Google Scholar]
- Hallé F., Oldeman R. A., Tomlinson P. B. (1978). Tropical Trees and Forests. An Architectural Analysis. New York, NY: Springer, 444. [Google Scholar]
- Hayward A., Stirnberg P., Beveridge C., Leyser O. (2009). Interactions between auxin and strigolactone in shoot branching control. Plant Physiol. 151 400–412. 10.1104/pp.109.137646 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hazebroek J. P., Metzger J. D., Mansager E. R. (1993). Thermoinductive regulation of gibberellin metabolism in Thlaspi arvense L. II. Cold Induction of enzymes in gibberellin biosynthesis. Plant Physiol. 102 547–552. 10.1104/pp.102.2.547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedden P., Phillips A. L. (2000). Gibberellin metabolism: new insights revealed by the genes. Trends Plant Sci. 5 523–530. 10.1016/s1360-1385(00)01790-8 [DOI] [PubMed] [Google Scholar]
- Hedden P., Sponsel V. (2015). A century of gibberellin research. J. Plant Growth Regul. 34 740–760. 10.1007/s00344-015-9546-1 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedden P., Thomas S. G. (2012). Gibberellin biosynthesis and its regulation. Biochem. J. 444 11–25. 10.1042/BJ20120245 [DOI] [PubMed] [Google Scholar]
- Helliwell C. A., Chandler P. M., Poole A., Dennis E. S., Peacock W. J. (2001). The CYP88A cytochrome P450, ent-kaurenoic acid oxidase, catalyzes three steps of the gibberellin biosynthesis pathway. Proc. Natl. Acad. Sci. U.S.A. 98 2065–2070. 10.1073/pnas.041588998 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirano K., Ueguchi-Tanaka M., Matsuoka M. (2008). GID1-mediated gibberellin signaling in plants. Trends Plant Sci. 13 192–199. 10.1016/j.tplants.2008.02.005 [DOI] [PubMed] [Google Scholar]
- Hu Y.-X., Tao Y.-B., Xu Z.-F. (2017). Overexpression of Jatropha gibberellin 2-oxidase 6 (JcGA2ox6) induces dwarfism and smaller leaves, flowers and fruits in Arabidopsis and Jatropha. Front. Plant Sci. 8:2103. 10.3389/fpls.2017.02103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Israelsson M., Mellerowicz E., Chono M., Gullberg J., Moritz T. (2004). Cloning and overproduction of gibberellin 3-oxidase in hybrid aspen trees. Effects on gibberellin homeostasis and development. Plant Physiol. 135 221–230. 10.1104/pp.104.038935 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ito S., Yamagami D., Umehara M., Hanada A., Yoshida S., Sasaki Y., et al. (2017). Regulation of strigolactone biosynthesis by gibberellin signaling. Plant Physiol. 174 1250–1259. 10.1104/pp.17.00301 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jasinski S., Piazza P., Craft J., Hay A., Woolley L., Rieu I., et al. (2005). KNOX action in Arabidopsis is mediated by coordinate regulation of cytokinin and gibberellin activities. Curr. Biol. 15 1560–1565. 10.1016/j.cub.2005.07.023 [DOI] [PubMed] [Google Scholar]
- Katyayini N. U., Rinne P. L. H., van der Schoot C. (2019). Strigolactone-based node-to-bud signaling may restrain shoot branching in hybrid aspen. Plant Cell Physiol. 60 2797–2811. 10.1093/pcp/pcz170 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kebrom T. H. (2017). A growing stem inhibits bud outgrowth–the overlooked theory of apical dominance. Front. Plant Sci. 8:1874. 10.3389/fpls.2017.01874 [DOI] [PMC free article] [PubMed] [Google Scholar]
- King R. W., Evans L. T., Mander L. N., Moritz T., Pharis R. P., Twitchin B. (2003). Synthesis of gibberellin GA6 and its role in flowering of Lolium temulentum. Phytochemistry 62 77–82. 10.1016/s0031-9422(02)00447-8 [DOI] [PubMed] [Google Scholar]
- King R. W., Mander L. N., Asp T., MacMillan C. P., Blundell C. A., Evans L. T. (2008). Selective deactivation of gibberellins below the shoot apex is critical to flowering but not to stem elongation of Lolium. Mol. Plant 1 295–307. 10.1093/mp/ssm030 [DOI] [PubMed] [Google Scholar]
- King R. W., Moritz T., Evans L. T., Junttila O., Herlt A. (2001). Long-day induction of flowering in Lolium temulentum involves sequential increases in specific gibberellins at the shoot apex. Plant Physiol. 127 624–632. 10.1104/pp.010378 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kobayashi M., Sakurai A., Saka H., Takahashi N. (1989). Quantitative analysis of endogenous gibberellins in normal and dwarf cultivars of rice. Plant Cell Physiol. 30 963–969. 10.1093/oxfordjournals.pcp.a077841 [DOI] [Google Scholar]
- Lange M. J. P., Lange T. (2016). Ovary-derived precursor gibberellin A9 is essential for female flower development in cucumber. Development 143 4425–4429. 10.1242/dev.135947 [DOI] [PubMed] [Google Scholar]
- Lantzouni O., Klermund C., Schwechheimer C. (2017). Largely additive effects of gibberellin and strigolactone on gene expression in Arabidopsis thaliana seedlings. Plant J. 92 924–938. 10.1111/tpj.13729 [DOI] [PubMed] [Google Scholar]
- Leyser O. (2009). The control of shoot branching: an example of plant information processing. Plant Cell Environ. 32 694–703. 10.1111/j.1365-3040.2009.01930.x [DOI] [PubMed] [Google Scholar]
- Li C.-J., Bangerth F. (1999). Autoinhibition of indoleacetic acid transport in the shoots of two-branched pea (Pisum sativum) plants and its relationship to correlative dominance. Physiol. Plant. 106 415–420. 10.1034/j.1399-3054.1999.106409.x 11841302 [DOI] [Google Scholar]
- Li H., Torres-Garcia J., Latrasse D., Benhamed M., Schilderink S., Zhou W., et al. (2017). Plant-specific histone deacetylases HDT1/2 regulate GIBBERELLIN 2-OXIDASE2 expression to control Arabidopsis root meristem cell number. Plant Cell 29 2183–2196. 10.1105/tpc.17.00366 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lo S.-F., Yang S.-Y., Chen K.-T., Hsing Y.-I., Zeevaart J. A., Chen L.-J., et al. (2008). A novel class of gibberellin 2-oxidases control semidwarfism, tillering, and root development in rice. Plant Cell 20 2603–2618. 10.1105/tpc.108.060913 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marzec M. (2017). Strigolactones and gibberellins: a new couple in the phytohormone world? Trends Plant Sci. 22 813–815. 10.1016/j.tplants.2017.08.001 [DOI] [PubMed] [Google Scholar]
- Mason M. G., Ross J. J., Babst B. A., Wienclaw B. N., Beveridge C. A. (2014). Sugar demand, not auxin, is the initial regulator of apical dominance. Proc. Natl. Acad. Sci. U.S.A. 111 6092–6097. 10.1073/pnas.1322045111 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mauriat M., Sandberg L. G., Moritz T. (2011). Proper gibberellin localization in vascular tissue is required to control auxin-dependent leaf development and bud outgrowth in hybrid aspen. Plant J. 67 805–816. 10.1111/j.1365-313X.2011.04635.x [DOI] [PubMed] [Google Scholar]
- Middleton A. M., Úbeda-Tomás S., Griffiths J., Holman T., Hedden P., Thomas S. G., et al. (2012). Mathematical modeling elucidates the role of transcriptional feedback in gibberellin signaling. Proc. Natl. Acad. Sci. U.S.A. 109 7571–7576. 10.1073/pnas.1113666109 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morris S. E., Cox M. C., Ross J. J., Krisantini S., Beveridge C. A. (2005). Auxin dynamics after decapitation are not correlated with the initial growth of axillary buds. Plant Physiol. 138 1665–1672. 10.1104/pp.104.058743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller D., Leyser O. (2011). Auxin, cytokinin and the control of shoot branching. Ann. Bot. 107 1203–1212. 10.1093/aob/mcr069 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Murfet I., Reid J. (1993). Peas: genetics, molecular biology and biotechnology. Seed Sci. Res. 4 165–216. [Google Scholar]
- Nakajima M., Shimada A., Takashi Y., Kim Y. C., Park S. H., Ueguchi-Tanaka M., et al. (2006). Identification and characterization of Arabidopsis gibberellin receptors. Plant J. 46 880–889. 10.1111/j.1365-313X.2006.02748.x [DOI] [PubMed] [Google Scholar]
- Nakayama I., Miyazawa T., Kobayashi M., Kamiya Y., Abe H., Sakurai A. (1990). Effects of a new plant growth regulator prohexadione calcium (BX-112) on shoot elongation caused by exogenously applied gibberellins in rice (Oryza sativa L.) seedlings. Plant Cell Physiol. 31 195–200. 10.1093/oxfordjournals.pcp.a077892 [DOI] [Google Scholar]
- Ni J., Gao C., Chen M.-S., Pan B.-Z., Ye K., Xu Z.-F. (2015). Gibberellin promotes shoot branching in the perennial woody plant Jatropha curcas. Plant Cell Physiol. 56 1655–1666. 10.1093/pcp/pcv089 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni J., Zhao M.-L., Chen M.-S., Pan B.-Z., Tao Y.-B., Xu Z.-F. (2017). Comparative transcriptome analysis of axillary buds in response to the shoot branching regulators gibberellin A3 and 6-benzyladenine in Jatropha curcas. Sci. Rep. 7:11417. 10.1038/s41598-017-11588-0 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nordström A., Tarkowski P., Tarkowska D., Norbaek R., Åstot C., Dolezal K., et al. (2004). Auxin regulation of cytokinin biosynthesis in Arabidopsis thaliana: a factor of potential importance for auxin–cytokinin-regulated development. Proc. Natl. Acad. Sci. U.S.A. 101 8039–8044. 10.1073/pnas.0402504101 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olszewski N., Sun T.-P., Gubler F. (2002). Gibberellin signaling: biosynthesis, catabolism, and response pathways. Plant Cell 14 S61–S80. 10.1105/tpc.010476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ongaro V., Bainbridge K., Williamson L., Leyser O. (2008). Interactions between axillary branches of Arabidopsis. Mol. Plant. 1 388–400. 10.1093/mp/ssn007 [DOI] [PubMed] [Google Scholar]
- Paul L. K., Rinne P. L., van der Schoot C. (2014). Shoot meristems of deciduous woody perennials: self-organization and morphogenetic transitions. Curr. Opin. Plant Biol. 17 86–95. 10.1016/j.pbi.2013.11.009 [DOI] [PubMed] [Google Scholar]
- Peng J., Carol P., Richards D. E., King K. E., Cowling R. J., Murphy G. P., et al. (1997). The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev. 11 3194–3205. 10.1101/gad.11.23.3194 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips A. L., Ward D. A., Uknes S., Appleford N. E., Lange T., Huttly A. K., et al. (1995). Isolation and expression of three gibberellin 20-oxidase cDNA clones from Arabidopsis. Plant Physiol. 108 1049–1057. 10.1104/pp.108.3.1049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Phillips I. D. J. (1975). Apical dominance. Annu. Rev. Plant Physiol. 26 341–367. 10.1146/annurev.pp.26.060175.002013 [DOI] [Google Scholar]
- Proebsting W. M., Hedden P., Lewis M. J., Croker S. J., Proebsting L. N. (1992). Gibberellin concentration and transport in genetic lines of pea: effects of grafting. Plant Physiol. 100 1354–1360. 10.1104/pp.100.3.1354 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puig J., Pauluzzi G., Guiderdoni E., Gantet P. (2012). Regulation of shoot and root development through mutual signaling. Mol. Plant 5 974–983. 10.1093/mp/sss047 [DOI] [PubMed] [Google Scholar]
- Ragni L., Nieminen K., Pacheco-Villalobos D., Sibout R., Schwechheimer C., Hardtke C. S. (2011). Mobile gibberellin directly stimulates Arabidopsis hypocotyl xylem expansion. Plant Cell 23 1322–1336. 10.1105/tpc.111.084020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rameau C., Bertheloot J., Leduc N., Andrieu B., Foucher F., Sakr S. (2015). Multiple pathways regulate shoot branching. Front. Plant Sci. 5:741. 10.3389/fpls.2014.00741 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Regnault T., Davière J.-M., Achard P. (2016). Long-distance transport of endogenous gibberellins in Arabidopsis. Plant Signal. Behav. 11:e1110661. 10.1080/15592324.2015.1110661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Richards D. E., King K. E., Ait-Ali T., Harberd N. P. (2001). How gibberellin regulates plant growth and development: a molecular genetic analysis of gibberellin signaling. Annu. Rev. Plant Physiol. Plant Mol. Biol. 52 67–88. 10.1146/annurev.arplant.52.1.67 [DOI] [PubMed] [Google Scholar]
- Rieu I., Eriksson S., Powers S. J., Gong F., Griffiths J., Woolley L., et al. (2008a). Genetic analysis reveals that C19-GA 2-oxidation is a major gibberellin inactivation pathway in Arabidopsis. Plant Cell 20 2420–2436. 10.1105/tpc.108.058818 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rieu I., Ruiz-Rivero O., Fernandez-Garcia N., Griffiths J., Powers S. J., Gong F., et al. (2008b). The gibberellin biosynthetic genes AtGA20ox1 and AtGA20ox2 act, partially redundantly, to promote growth and development throughout the Arabidopsis life cycle. Plant J. 53 488–504. 10.1111/j.1365-313X.2007.03356.x [DOI] [PubMed] [Google Scholar]
- Rinne P. L., Paul L. K., Vahala J., Kangasjärvi J., van der Schoot C. (2016). Axillary buds are dwarfed shoots that tightly regulate GA pathway and GA-inducible 1,3-β-glucanase genes during branching in hybrid aspen. J. Exp. Bot. 67 5975–5991. 10.1093/jxb/erw352 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinne P. L., Paul L. K., Vahala J., Ruonala R., Kangasjärvi J., van der Schoot C. (2015). Long and short photoperiod buds in hybrid aspen share structural development and expression patterns of marker genes. J. Exp. Bot. 66 6745–6760. 10.1093/jxb/erv380 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rinne P. L., Welling A., Vahala J., Ripel L., Ruonala R., Kangasjärvi J., et al. (2011). Chilling of dormant buds hyperinduces FLOWERING LOCUS T and recruits GA-inducible 1,3-β-glucanases to reopen signal conduits and release dormancy in Populus. Plant Cell 23 130–146. 10.1105/tpc.110.081307 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakamoto T., Kamiya N., Ueguchi-Tanaka M., Iwahori S., Matsuoka M. (2001). KNOX homeodomain protein directly suppresses the expression of a gibberellin biosynthetic gene in the tobacco shoot apical meristem. Genes Dev. 15 581–590. 10.1101/gad.867901 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schommer C., Debernardi J. M., Bresso E. G., Rodriguez R. E., Palatnik J. F. (2014). Repression of cell proliferation by miR319-regulated TCP4. Mol. Plant 7 1533–1544. 10.1093/mp/ssu084 [DOI] [PubMed] [Google Scholar]
- Scott T. K., Case D. B., Jacobs W. P. (1967). Auxin-gibberellin interaction in apical dominance. Plant Physiol. 42 1329–1333. 10.1104/pp.42.10.1329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seale M., Bennett T., Leyser O. (2017). BRC1 expression regulates bud activation potential, but is not necessary or sufficient for bud growth inhibition in Arabidopsis. Development 144 1661–1673. 10.1242/dev.145649 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silverstone A. L., Ciampaglio C. N., Sun T. (1998). The Arabidopsis RGA gene encodes a transcriptional regulator repressing the gibberellin signal transduction pathway. Plant Cell 10 155–169. 10.1105/tpc.10.2.155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silverstone A. L., Mak P. Y. A., Martinez E. C., Sun T. (1997). The new RGA locus encodes a negative regulator of gibberellin response in Arabidopsis thaliana. Genetics 146 1087–1099. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sponsel V. M., Schmidt F. W., Porter S. G., Nakayama M., Kohlstruk S., Estelle M. (1997). Characterization of new gibberellin-responsive semidwarf mutants of Arabidopsis. Plant Physiol. 115 1009–1020. 10.1104/pp.115.3.1009 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun T. (2010). Gibberellin-GID1-DELLA: a pivotal regulatory module for plant growth and development. Plant Physiol. 154 567–570. 10.1104/pp.110.161554 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun T. (2011). The molecular mechanism and evolution of the GA–GID1–DELLA signaling module in plants. Curr. Biol. 21 R338–R345. 10.1016/j.cub.2011.02.036 [DOI] [PubMed] [Google Scholar]
- Talon M., Koornneef M., Zeevaart J. A. (1990). Accumulation of C19-gibberellins in the gibberellin-insensitive dwarf mutant gai of Arabidopsis thaliana (L.) Heynh. Planta 182 501–505. 10.1007/BF02341024 [DOI] [PubMed] [Google Scholar]
- Tanaka M., Takei K., Kojima M., Sakakibara H., Mori H. (2006). Auxin controls local cytokinin biosynthesis in the nodal stem in apical dominance. Plant J. 45 1028–1036. 10.1111/j.1365-313X.2006.02656.x [DOI] [PubMed] [Google Scholar]
- Tenreira T., Lange M. J. P., Lange T., Bres C., Labadie M., Monfort A., et al. (2017). A specific gibberellin 20-oxidase dictates the flowering-runnering decision in diploid strawberry. Plant Cell 29 2168–2182. 10.1105/tpc.16.00949 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thimann K. V., Skoog F. (1934). On the inhibition of bud development and other functions of growth substance in Vicia faba. Proc. R. Soc. Lond. B Biol. Sci. 114 317–339. 10.1098/rspb.1934.0010 [DOI] [Google Scholar]
- Thomas S. G., Phillips A. L., Hedden P. (1999). Molecular cloning and functional expression of gibberellin 2-oxidases, multifunctional enzymes involved in gibberellin deactivation. Proc. Natl. Acad. Sci. U.S.A. 96 4698–4703. 10.1073/pnas.96.8.4698 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tuskan G. A., Difazio S., Jansson S., Bohlmann J., Grigoriev I., Hellsten U., et al. (2006). The genome of black cottonwood, Populus trichocarpa (Torr. & Gray). Science 313 1596–1604. 10.1126/science.1128691 [DOI] [PubMed] [Google Scholar]
- Ueguchi-Tanaka M., Ashikari M., Nakajima M., Itoh H., Katoh E., Kobayashi M., et al. (2005). GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 437 693–698. 10.1038/nature04028 [DOI] [PubMed] [Google Scholar]
- Ueguchi-Tanaka M., Nakajima M., Motoyuki A., Matsuoka M. (2007). Gibberellin receptor and its role in gibberellin signaling in plants. Annu. Rev. Plant Biol. 58 183–198. 10.1146/annurev.arplant.58.032806.103830 [DOI] [PubMed] [Google Scholar]
- Urbanová T., Tarkowská D., Novák O., Hedden P., Strnad M. (2013). Analysis of gibberellins as free acids by ultra performance liquid chromatography–tandem mass spectrometry. Talanta 112 85–94. 10.1016/j.talanta.2013.03.068 [DOI] [PubMed] [Google Scholar]
- Wang M., Le Moigne M. A., Bertheloot J., Crespel L., Perez-Garcia M. D., Ogé L., et al. (2019). BRANCHED1: a key hub of shoot branching. Front. Plant Sci. 10:76. 10.3389/fpls.2019.00076 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Willige B. C., Ghosh S., Nill C., Zourelidou M., Dohmann E. M., Maier A., et al. (2007). The DELLA domain of GA INSENSITIVE mediates the interaction with the GA INSENSITIVE DWARF1A gibberellin receptor of Arabidopsis. Plant Cell 19 1209–1220. 10.1105/tpc.107.051441 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu R., Hinckley T. M. (2001). Phenotypic plasticity of sylleptic branching: genetic design of tree architecture. Crit. Rev. Plant Sci. 20 467–485. 10.1080/07352689.2001.10131827 [DOI] [Google Scholar]
- Wu R., Stettler R. (1998). Quantitative genetics of growth and development in Populus. III. Phenotypic plasticity of crown structure and function. Heredity 81 299–310. 10.1046/j.1365-2540.1998.00397.x [DOI] [Google Scholar]
- Xu Y. L., Li L., Gage D. A., Zeevaart J. A. (1999). Feedback regulation of GA5 expression and metabolic engineering of gibberellin levels in Arabidopsis. Plant Cell 11 927–935. 10.1105/tpc.11.5.927 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaguchi S. (2008). Gibberellin metabolism and its regulation. Annu. Rev. Plant Biol. 59 225–251. 10.1146/annurev.arplant.59.032607.092804 [DOI] [PubMed] [Google Scholar]
- Yamaguchi S., Kamiya Y. (2000). Gibberellin biosynthesis: its regulation by endogenous and environmental signals. Plant Cell Physiol. 41 251–257. 10.1093/pcp/41.3.251 [DOI] [PubMed] [Google Scholar]
- Zawaski C., Busov V. B. (2014). Roles of gibberellin catabolism and signaling in growth and physiological response to drought and short-day photoperiods in Populus trees. PLoS One 9:e86217. 10.1371/journal.pone.0086217 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhuang W., Gao Z., Wen L., Huo X., Cai B., Zhang Z. (2015). Metabolic changes upon flower bud break in Japanese apricot are enhanced by exogenous GA4. Hortic. Res. 2:15046. 10.1038/hortres.2015.46 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Data Availability Statement
All datasets generated for this study are included in the article/Supplementary Material.