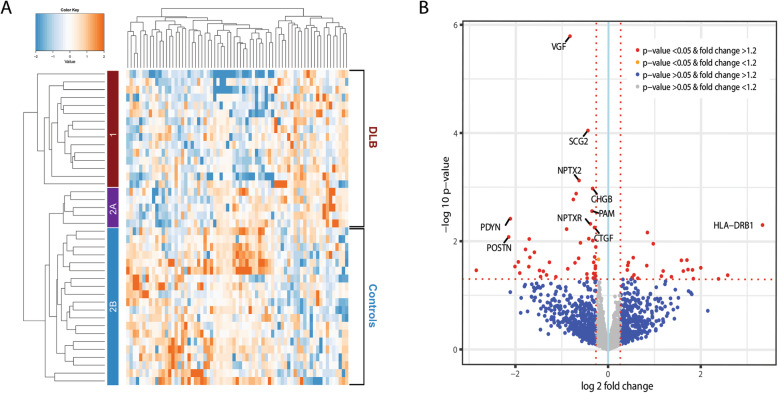
Fig. 2.
Results of discovery proteomics. a Heatmap and cluster analysis of differentially expressed proteins (n = 69) in cohort 1. The heatmap shows distinct patterns of up- and downregulated proteins in the clinical groups. The branching pattern of the dendrogram shows almost complete separation of patients with DLB from cognitively normal controls (35/40 (87.5%) were clustered correctly). Fifteen DLB patients were assigned to cluster 1 (red) and five DLB patients and 20 controls were assigned to cluster 2. The five DLB patients in cluster 2 clustered together in a small subgroup (cluster 2A, purple) and the controls clustered together in another subgroup (cluster 2B, blue). b Volcano plot representing the top biomarker candidates discriminating DLB from controls. The horizontal axis indicates log2 fold change. The vertical axis indicates − 10 log p-values. Each point represents a protein. Points at the far right- and left-hand sides of the plot have the largest fold changes, while those along the top of the plot are the most statistically significant. The non-axial red dotted vertical lines denote fold change thresholds of 1.2. The non-axial red dotted horizontal line denotes p-value threshold of 0.05. Proteins in red have a fold change > 1.2 and p-value < 0.05. The top-10 biomarker candidates are highlighted in the plot