Fig 7. Tepoxalin is active against ABCB1-high cancer cell lines via an ABCB1-mediated mechanism.
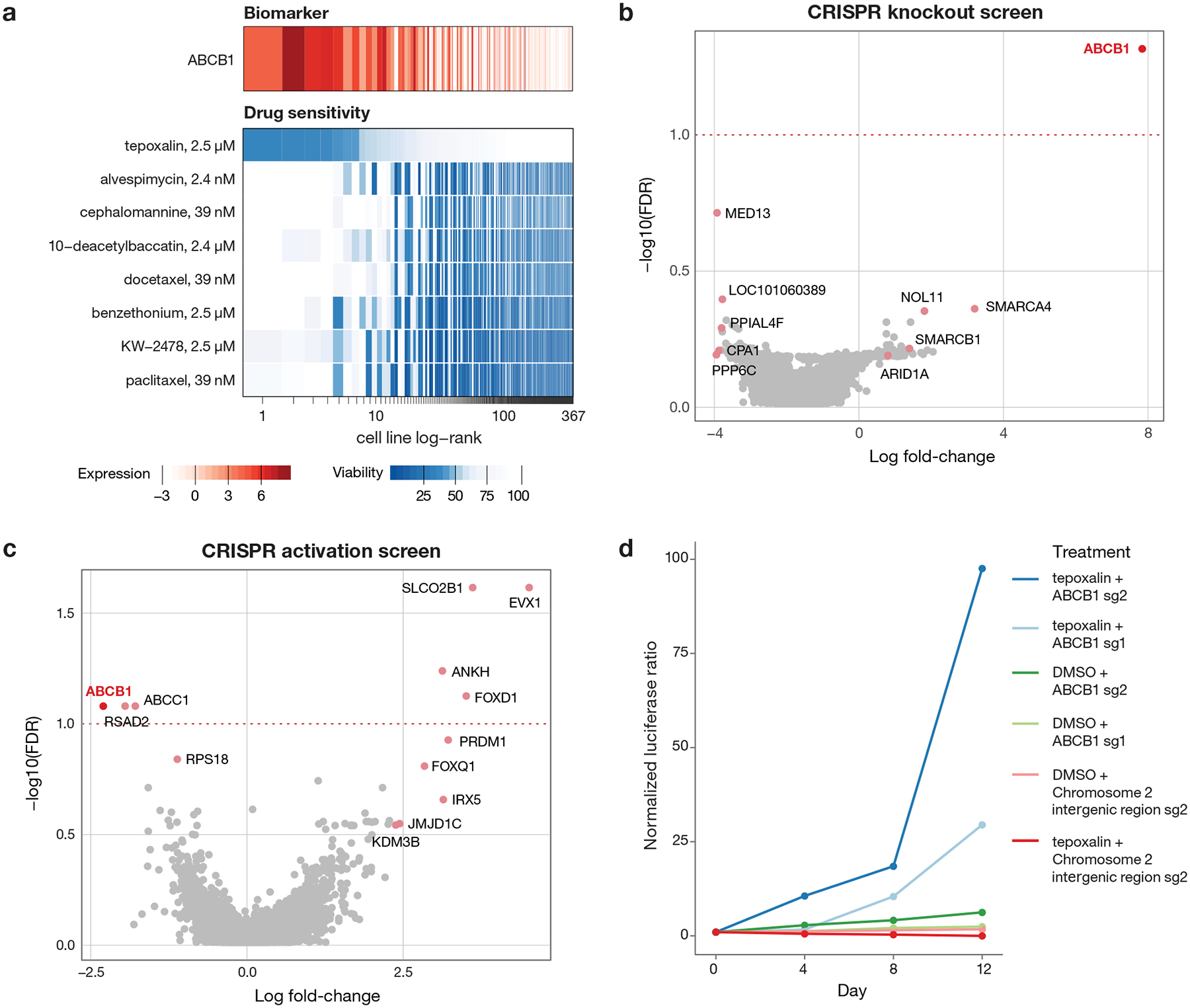
a, Drug sensitivity profiles of tepoxalin and seven additional compounds where ABCB1 RNA expression was the top predictive genomic feature of the PRISM activity profile (n = 489 cell lines). Tepoxalin was the only compound tested where high ABCB1 expression was associated with sensitivity rather than resistance to tepoxalin. Cell line viability is depicted with ABCB1 gene expression from CCLE RNAseq log2 RPKM data. Cell lines are ranked by mean viability with a ceiling at 100%. b, ABCB1 knockout is the top hit that rescues tepoxalin activity in a genome-wide CRISPR/Cas9 gene knockout screen. LS1034-Cas9 cells (pXPR_311) were infected with the Brunello sgRNA library, selected with puromycin, and treated with 16 μM tepoxalin versus vehicle control with two replicates in independent flasks. Cells were passaged every 3–4 days over a 30-day period. Two-sided p-values are computed using MAGeCK-MLE and shown versus gene-level log fold-changes. c, ABCB1 overexpression sensitizes to tepoxalin activity in a genome-wide CRISPR/dCas9 gene activation screen. LS1034-dCas9 cells (pXPR_109) were infected with the Calabrese sgRNA library, selected with puromycin, and treated with 16 μM tepoxalin versus vehicle control with two replicates in independent flasks. Cells were passaged every 3–4 days over a 14-day period. Two-sided p-values are computed using MAGeCK-MLE and shown versus gene-level log fold-changes. d, Tepoxalin cellular competition assay following ABCB1 knockout. LS1034-Cas9 cells (pXPR_311) were stably infected with Firefly luciferase and parental LS1034 cells (without Cas9) were stably infected with Renilla luciferase. Cells were mixed in 1:1 ratio an infected with the indicated sgRNA construct against ABCB1 or an intergenic region on chromosome 2 (negative control). Following puromycin selection, cell mixtures were treated with 16 μM tepoxalin. Firefly to Renilla luminescence ratio is plotted as log fold-change over time. Mean of three technical replicates is shown and results are representative of three independent experiments.
