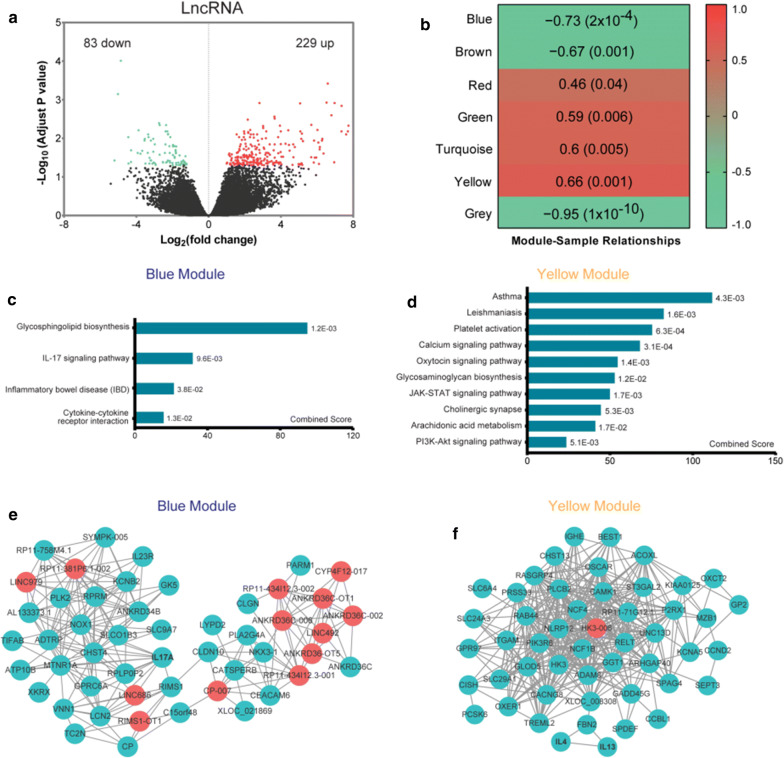
Fig. 5.
Differentially expressed lncRNAs and pathway analysis between CRSwNP + AS and CRSwNP-alone. a Volcano plots illustrating DE-lncRNAs of CRSwNP + AS versus CRSwNP-alone identified by RNA sequencing. b The correlation between modules and phenotype of CRSwNP + AS. Seven modules were identified by WGCNA based on expression of DE-mRNAs and DE-lncRNAs of CRSwNP + AS versus CRSwNP-alone. Pearson’s correlation coefficient between each module and phenotype of CRSwNP + AS and their associated P values are shown in the corresponding modules. The red and green colours show a strong positive and negative correlation, respectively. c, d All or top 10 KEGG pathways significantly enriched by genes in blue module c and yellow module (d). e, f Top 50 hub genes in blue module (e) and yellow module (f) visualized by cytoscape network. mRNAs or lncRNAs with high connectivity and edges with weight above a threshold of 0.1 were identified as hub genes. The red nodes denote lncRNAs, and the green nodes denote mRNAs. P < 0.05 were considered statistically significant. CRSwNP chronic rhinosinusitis with nasal polyps; AS: asthma, lncRNA long non-coding RNA, DE differentially expressed, KEGG Kyoto Encyclopedia of Genes and Genomes