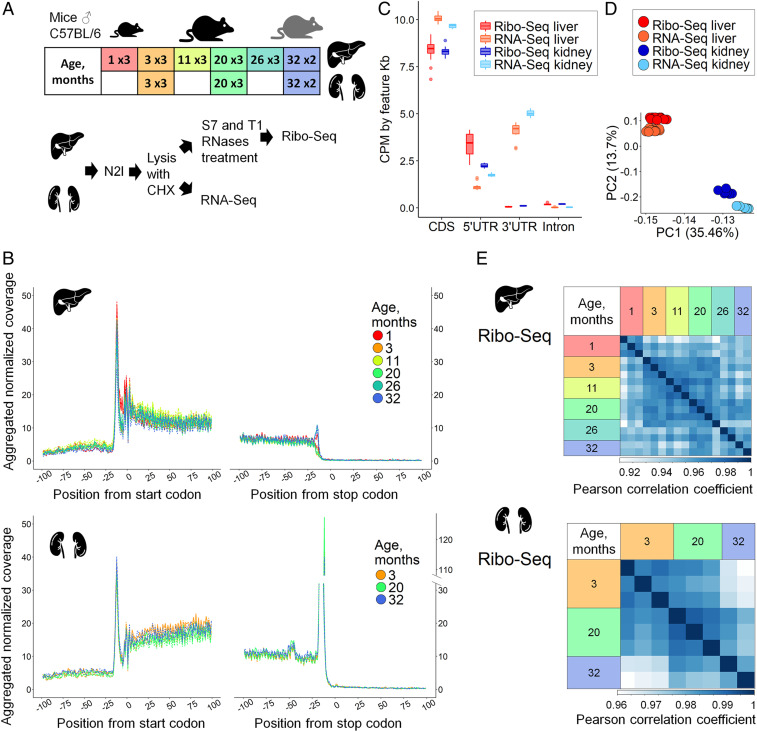
Fig. 1.
Ribo-seq and RNA-seq of aging mouse liver and kidney. (A) Overview of experimental design. Mouse livers representing six age groups (1-, 3-, 11-, 20-, 26-, and 32-mo-old) and kidneys representing three age groups (3-, 20-, and 32-mo-old) were used. For each age, three biological replicates were prepared (three C57BL/6 male mice), except for the 32-mo group (two mice). Ribo-seq and RNA-seq libraries were prepared from the same cytoplasmic cell lysate. (B) Metagene profiles of ribosomal footprint 5′ ends in 200-nt windows centered at start and stop codons built for 2,920 and 4,566 transcripts for liver and kidney, respectively. For each transcript, raw Ribo-seq coverage was normalized to the sum of transcript coverage divided by its length. Normalized transcript coverage in the window was then aggregated for all selected transcripts. (C) Distribution of Ribo-seq and RNA-seq coverage in different gene regions. (D) Principal-component analysis (PCA) of 8,562 genes in Ribo-seq and RNA-seq datasets of mouse liver and kidney. (E) Heatmaps of Pearson correlation coefficients for replicates of mouse liver and kidney analyzed by Ribo-seq. For PCA and calculation of Pearson correlation coefficients, and further in the study, Ribo-seq was analyzed together with the RNA-seq dataset, but separately for organs. In total, the number of genes covered in each sample was 8,992 in liver and 11,461 in kidney.