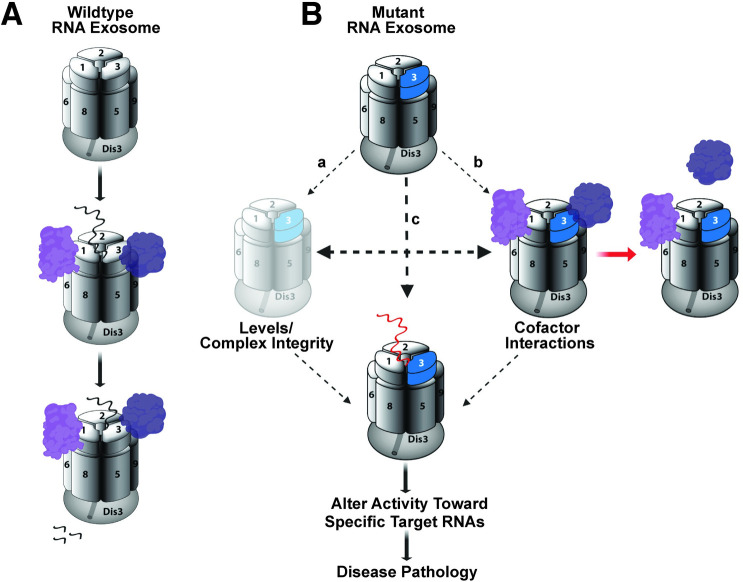
Fig 10. Mechanistic model for how amino acid substitutions could alter RNA exosome function.
(A) The Wildtype RNA exosome (top) associates with a variety of cofactors (purple) that can aid in target RNA recognition (middle) and decay/processing (bottom). (B) Amino acid substitutions that occur in the EXOSC3/Rrp40 cap subunit (shown in blue) could impact the function of the complex through multiple mechanisms, including: (a) decrease the steady-state level of the specific subunit or alter interaction of the subunit with the complex, leading to a decrease in overall Levels or altered Complex Integrity; (b) affect Cofactor Interactions, which could impact interactions with target RNAs; and/or (c) could directly alter interactions with target RNAs (red RNA indicates impaired interaction). Ultimately, any of these changes could Alter Activity Toward Specific Target RNAs. Changes in the steady state level or processing of RNAs could then cause downstream Disease Pathology.