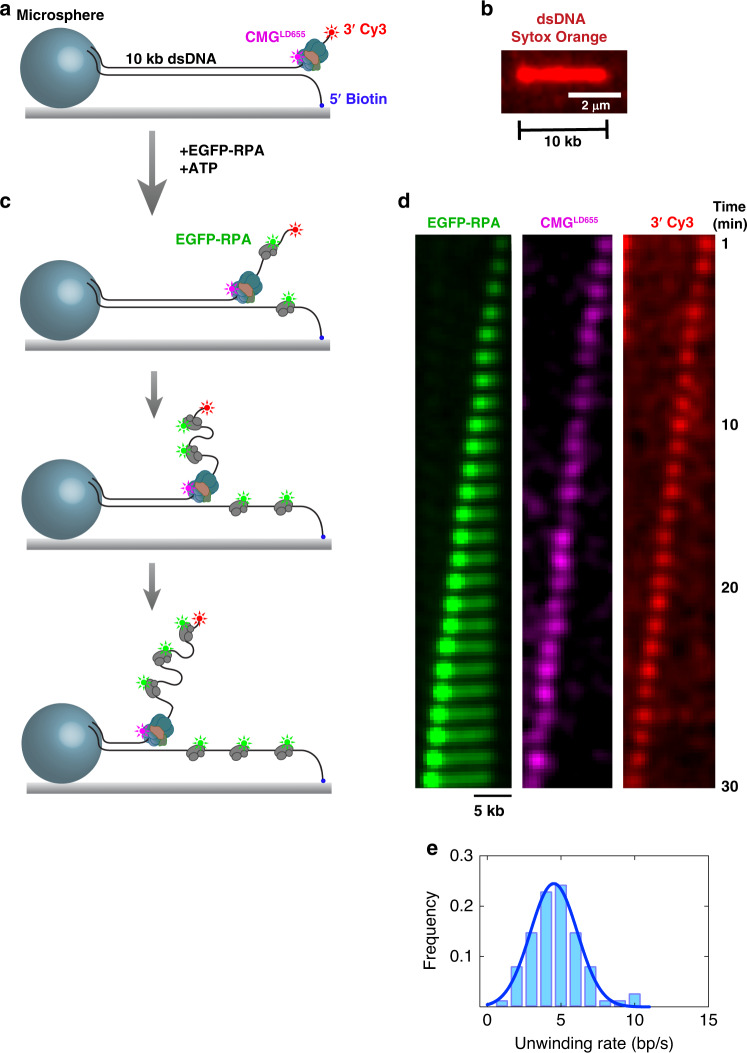
Fig. 1. Direct visualization of RPA-facilitated processive fork unwinding by individual CMG molecules.
a A 10-kb fragment of λ DNA was ligated to a short fork DNA substrate at one end and to a digoxigenin-modified DNA fragment on the opposite end. The 5′ tail of the forked end contained a biotin to attach the 10-kb substrate to biotin-functionalized glass through biotin-streptavidin binding. The digoxigenin-modified end was coupled to anti-digoxigenin-coated microsphere to stretch DNA molecules by buffer flow, and to subsequently attach this end to the surface (details are described in the Methods section). The DNA substrate was labeled with Cy3 at the 3′ dT40 ssDNA tail near the fork junction. LD655-labeled CMG was bound to the dT40 ssDNA in the presence of ATPγS. b A sample stretched 10-kb linear DNA stained with a fluorescent dsDNA intercalator Sytox Orange. Average length of stretched DNA molecules were determined by measuring the end-to-end distance of 717 individual DNA molecules (n = 2 independent experiments). c After binding CMG on the surface-immobilized DNA, EGFP-RPA, and ATP was introduced to initiate unwinding. While CMG unwinds DNA at the fork, EGFP-RPA binds both strands of unwound DNA. d Kymograph showing a representative 10-kb DNA being unwound entirely by a single CMG complex. EGFP-RPA (left panel), LD655-labeled CMG (center panel), and 3′ Cy3 (right panel) are imaged during unwinding under near-TIRF conditions. Images were acquired in the absence of buffer flow. e Histogram of CMG-catalyzed DNA-unwinding rates on stretched DNA (74 individual DNA molecules analyzed from two independent experiments). Source data are provided as a Source Data file.