Fig. 5.
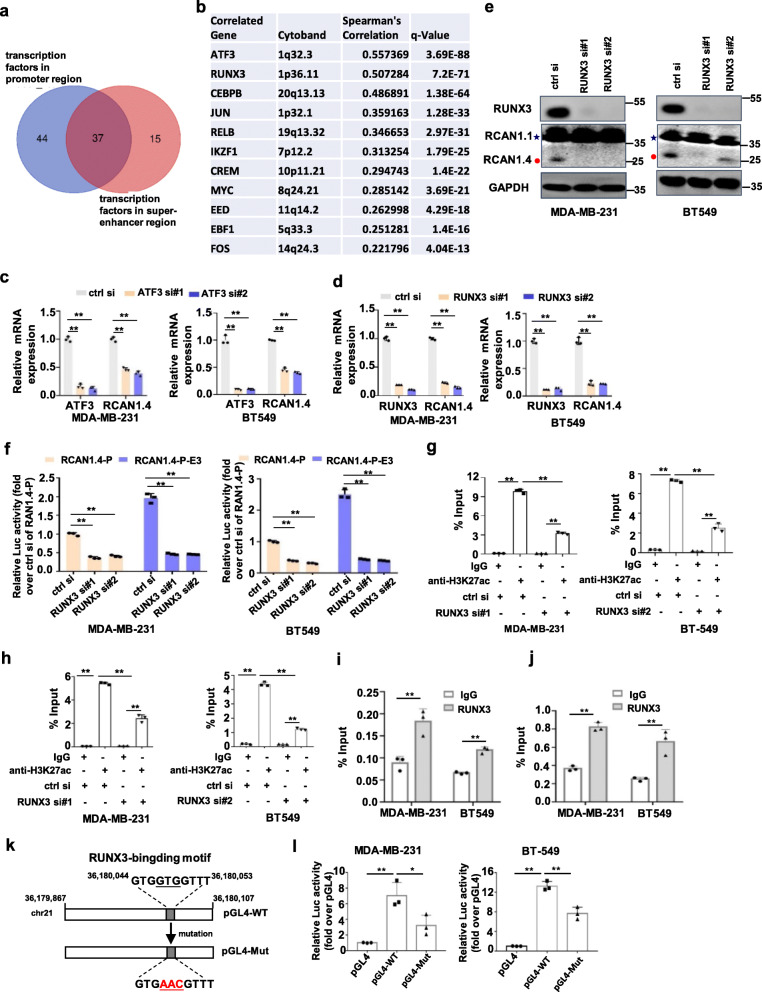
RUNX3 interacts with both the super-enhancer and promoter regions of RCAN1.4 and promotes its transcription. a Venn diagrams showing commonly binding transcription factors on the promote and super-enhancer regions of RCAN1.4. b The correlation of RCAN1.4 with 11 commonly binding transcription factors on the promoter and super-enhancer regions of RCAN1.4. The data were retrieved from breast invasive carcinoma (TCGA, Firehose Legacy) database using the Cbioportal website (https://www.cbioportal.org/). c-d MDA-MB-231 cells and BT549 cells were transfected with siRNAs targeting ATF3 (c) or RUNX3 (d). The mRNA levels were quantified using qRT–PCR. e MDA-MB-231 cells and BT549 cells were transfected with siRNAs targeting RUNX3. The cell lysates were prepared for immunoblots. Blue star, RCAN1.1; Red closed circle, RCAN1.4. f Luciferase reporter assay of RCAN1.4 promote activity and E3 super-enhancer activity in MDA-MB-231 and BT549 cells with RUNX3 knockdown. g-h The MDA-MB-231 and BT549 cells with RUNX3 knockdown were subjected to ChIP analysis using antibodies against H3K27ac. The association with the SE region (g) and promoter region (h) of RCAN1.4 was quantified by qPCR. i-j MDA-MB-231 cells and BT549 were subjected to ChIP analysis using antibodies against RUNX3. The association with the promoter region region (i) and SE (j) of RCAN1.4 was quantified by qPCR. k Schematic representation of a 241 bp region of the RCAN1.4 enhancer (from chr21:36179867–36,180,107) containing the wild-type RUNX3 motif binding sequence (from chr21:36180044–36,180,053) or the mutant alleles. l The MDA-MB-231 and BT549 cells were transfected with the indicated plasmids for 48 h. The levels of luciferase activity were normalized to pRL-TK luciferase activity. Error bars represent mean ± SD, n = 3 biological independent samples. * P < 0.05, **P < 0.01. The P value in c, d, f was determined by one-way analysis ANOVA with Dunnett’s multiple comparisons test, the P value in g, h, l was determined by one-way ANOVA with Tukey’s multiple comparisons test, no adjustments were made for multiple comparisons. Data were representative of three independent experiments