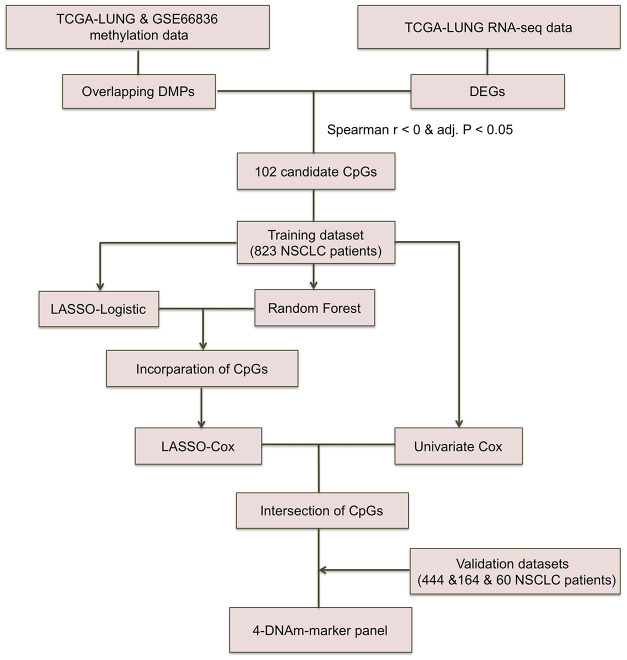
Figure 1.
Workflow chart of CpG marker selection. Two DNA methylation (DNAm) datasets and TCGA RNA-seq dataset were used for identifying 102 candidate CpG markers. Based on recurrence-free survival data of the training cohort (823 TCGA NSCLC patients), LASSO-Logistic and Random Forest methods were applied to identify recurrence associated CpG markers. With the incorporation of CpGs identified by two methods above, LASSO-Cox were implemented to select robust DNAm signatures. Using the CpGs overlapped in results of univariate Cox and LASSO-Cox models, the 4-DNAm-marker panel was finally identified and verified in validation cohorts. adj.P: Bonferroni corrected P.