Abstract
Pseudogenes have been reported to exert oncogenic or tumor-suppressive functions in cancer. However, the expression, role, and mechanism of pseudogene-derived RNAs in breast cancer remain unclear. The RNA levels and prognostic values of pseudogenes in breast cancer were determined. The levels of RP11-480I12.5 in cell lines and clinical samples were validated by quantitative real-time PCR. In vitro effects of RP11-480I12.5 on cell growth were measured by cell counting kit-8 (CCK-8) assay, colony formation assay, cell counting assay, and flow cytometry analysis. Xenograft model was established to detect its in vivo effect. The potential mechanism of RP11-480I12.5 was also studied by a combination of bioinformatic analysis and experimental confirmation. Finally, the possible functional parental genes of RP11-480I12.5 in breast cancer were explored. After a series of bioinformatic analyses, RP11-480I12.5 was selected as the most potential pseudogene in breast cancer. RP11-480I12.5 expression was significantly upregulated in breast cancer cell lines and clinical breast cancer tissues. Knockdown of RP11-480I12.5 markedly suppressed cell proliferation and colony formation, induced cell apoptosis of breast cancer in vitro, and inhibited tumor growth in vivo. Four transcripts of RP11-480I12.5 (001/002/003/004) were identified. Only overexpression of RP11-480I12.5-004 significantly enhanced cell growth of breast cancer both in vitro and in vivo. RP11-480I12.5-004 mainly located in cytoplasm and increased AKT3 and CDK6 mRNA expression, at least in part, by competitively binding to miR-29c-3p. Six parental genes of RP11-480I12.5 were found, among which TUBA1B and TUBA1C were statistically linked to RP11-480I12.5 expression, possessed prognostic values, and were upregulated in breast cancer. Our findings suggested that pseudogene-derived long non-coding RNA (lncRNA) RP11-480I12.5-004 promoted growth and tumorigenesis of breast cancer via increasing AKT3 and CDK6 expression by competitively binding to miR-29c-3p.
Keywords: breast cancer, pseudogene, microRNA, miRNA, long non-coding RNA, lncRNA, RP11-480I12.5-004, miR-29c-3p, CDK6, AKT3
Graphical Abstract

By combination of in silico analysis and experimental validation, Lou et al. elucidated that pseudogene-derived lncRNA RP11-480I12.5-004 promoted growth and tumorigenesis of breast cancer via increasing AKT3 and CDK6 expression by competitively binding to miR-29c-3p.
Introduction
Breast cancer is the second most frequent cancer in females globally.1 It also ranks as the most common type of cancer along with high mortality in women, accounting for approximately 25% of cancer-associated deaths worldwide.1,2 Despite the huge improvements of therapeutic strategies targeting breast cancer that have been received, including surgery, chemotherapy, radiotherapy, and endocrine therapy, the prognosis and outcome of patients with breast cancer are still not satisfactory.3,4 Development of effective therapeutic targets for breast cancer may benefit from understanding of underlying mechanisms of breast cancer pathogenesis. Over the past decades, researchers and scholars have performed a growing number of molecular investigations about carcinogenesis of breast cancer, supporting that long non-coding RNAs (lncRNAs) play crucial roles in occurrence and progression of breast cancer.5,6
lncRNAs represent a group of RNA molecules with more than 200 nucleotides without protein-coding potential, among which pseudogene-derived lncRNAs have been widely concerned recently.7 Numerous studies have been well-demonstrated that lncRNAs, containing pseudogene-derived lncRNAs, act as critical regulators in a variety of biological processes, such as cell differentiation, autophagy, apoptosis, and death.8, 9, 10, 11 Dysregulation of pseudogene-derived lncRNAs links to development of multiple diseases, including cancer.11 For example, PDIA3P-derived lncRNA promotes proliferation and invasion of non-small cell lung cancer,12 PCNAP1-derived lncRNA modulates hepatitis B virus replication and enhances tumor growth of liver cancer,13 and KRT19P3-derived lncRNA inhibits proliferation and metastasis through COPS7A-mediated nuclear factor κB (NF-κB) pathway in gastric cancer.14 Several studies conducted by our team and other labs have also suggested that pseudogenes and their RNA transcripts participate in carcinogenesis of breast cancer and serve as promising biomarkers for patients with breast cancer.15,16 However, the knowledge of roles and mechanisms of pseudogene-derived lncRNA in breast cancer is still insufficient and needs to be further investigated.
In this study, we first screened differentially expressed pseudogenes (DEPs) between breast cancer tissues and normal breast tissues using dreamBase and GEPIA databases. Next, stage expression analysis and survival analysis for these DEPs indicated that RP11-480I12.5 was the most potential pseudogene, which may be involved in occurrence and progression of breast cancer. The high expression of RP11-480I12.5 was then experimentally confirmed in breast cancer cell lines and collected breast cancer samples. Subsequently, in vitro and in vivo functional assays were performed to determine the roles of RP11-480I12.5 in breast cancer. Then, the possible action mechanism that RP11-480I12.5 exerted its functions in breast cancer was explored. Finally, for the first time, we identified TUBA1B and TUBA1C as two potential parental genes of RP11-480I12.5 in breast cancer. The findings from this work suggest the expression, role, and mechanism of RP11-480I12.5 in breast cancer and provide key clues for targeting RP11-480I12.5 as a promising therapeutic measure in patients with breast cancer.
Results
RP11-480I12.5 Is Upregulated in Breast Cancer and Correlates with Poor Prognosis and Progression of Breast Cancer
In order to study roles of pseudogene in breast cancer, a series of procedures were conducted to identify potential pseudogenes (Figure 1A). DEPs were first found by dreamBase database. As presented in Table S2, a total of 630 DEPs, containing 264 upregulated and 366 downregulated DEPs, were screened. Next, expression levels of these DEPs were validated using GEPIA database. 12 of 264 upregulated DEPs and 38 of 366 downregulated DEPs were also significantly upregulated and downregulated in breast cancer compared with normal controls in GEPIA database. Expression difference of the 50 pseudogenes among various major stages were determined in breast cancer. Four upregulated DEPs (RP11-424C20.2, UBE2SP2, RP11-480I12.5, and SORD2P) and six downregulated DEPs (RP11-481H12.1, RP4-631H13.6, RRN3P1, UBE2Q2P1, RP11-798K23.5, and FAR2P2) presented statistical expression difference (Table 1). Then, the prognostic values of the 10 DEPs in breast cancer were evaluated using The Cancer Genome Atlas (TCGA) breast cancer data as shown in Figures 1B–1K. Among the 10 pseudogenes, only breast cancer patients with higher expression of RP11-480I12.5 had poorer prognosis. chi-square test revealed that RP11-480I12.5 expression was significantly linked to estrogen receptor status (p < 0.0001), progesterone receptor status (p < 0.0001), Her2 receptor status (p < 0.0001), N stage (p = 0.0017), M stage (p = 0.0443), and pathological stage (p = 0.0054; Table 2). As presented in Figures 1L and 1M, RP11-480I12.5 expression was significantly increased in breast cancer cell lines and clinical breast cancer tissues when compared with corresponding controls, respectively. Notably, expression of RP11-480I12.5 in triple-negative breast cancer (TNBC) cell lines (HCC1937, BT549, and MDA-MB-231) was generally higher than that in luminal-type cell lines (MCF-7, T47D, and BT474). Furthermore, the result from clinical tissue analysis also demonstrated that RP11-480I12.5 expression was markedly increased in TNBC samples compared with non-TNBC samples (Figure 1N). Taken together, pseudogene RP11-480I12.5 may be a crucial oncogene in tumorigenesis of breast cancer. Therefore, RP11-480I12.5 was selected for subsequent functional investigation.
Figure 1.
Identification of RP11-480I12.5 as a Potential Regulator in Breast Cancer
(A) The flow-process diagram of this study. (B) The prognostic value of RP11-424C20.2 in breast cancer. (C) The prognostic value of UBE2SP2 in breast cancer. (D) The prognostic value of RP11-480I12.5 in breast cancer. (E) The prognostic value of SORD2P in breast cancer. (F) The prognostic value of RP11-481H12.1 in breast cancer. (G) The prognostic value of RP4-631H13.6 in breast cancer. (H) The prognostic value of RRN3P1 in breast cancer. (I) The prognostic value of UBE2Q2P1 in breast cancer. (J) The prognostic value of RP11-798K23.5 in breast cancer. (K) The prognostic value of FAR2P2 in breast cancer. (L) The expression levels of RP11-480I12.5 in human breast cancer cell lines compared with normal breast cell. (M) The expression of RP11-480I12.5 in breast cancer samples (cancer) compared with control normal breast samples (normal). (N) The expression of RP11-480I12.5 in triple-negative breast cancer (TNBC) samples compared with non-TNBC samples. ∗p < 0.05.
Table 1.
Differential Expression of Pseudogene-Derived RNAs among Various Major Stages (Stage I, Stage II, Stage III, Stage IV, and Stage X) in Breast Cancer Determined by GEPIA
| Name | Expression (Tumor/Normal) | p Value | F-Value |
|---|---|---|---|
| AL162151.3 | upregulated | 0.452 | 0.919 |
| HIST1H1PS1 | upregulated | 0.835 | 0.363 |
| RP11-424C20.2 | upregulated | 0.037 | 2.570 |
| CYP2B7P | upregulated | 0.151 | 1.690 |
| RPLP0P2 | upregulated | 0.079 | 2.100 |
| UBE2SP2 | upregulated | 0.005 | 3.690 |
| UBE2SP1 | upregulated | 0.162 | 1.640 |
| RP11-480I12.5 | upregulated | 0.009 | 3.400 |
| HIST2H2BD | upregulated | 0.318 | 1.180 |
| SORD2P | upregulated | 0.047 | 2.420 |
| SNRPEP2 | upregulated | 0.346 | 1.120 |
| CTD-2583A14.11 | upregulated | 0.256 | 1.330 |
| RP11-258C19.4 | downregulated | 0.461 | 0.904 |
| FP325317.1 | downregulated | 0.062 | 2.250 |
| RP11-481H12.1 | downregulated | 0.046 | 2.430 |
| CPHL1P | downregulated | 0.224 | 1.420 |
| bP-21264C1.1 | downregulated | 0.644 | 0.626 |
| PKD1L2 | downregulated | 0.227 | 1.410 |
| TPTEP1 | downregulated | 0.057 | 2.300 |
| ZNF300P1 | downregulated | 0.241 | 1.370 |
| RPL23AP49 | downregulated | 0.183 | 1.560 |
| GGTA1P | downregulated | 0.173 | 1.600 |
| USP32P1 | downregulated | 0.170 | 1.610 |
| RPL23AP1 | downregulated | 0.285 | 1.260 |
| PABPC1P4 | downregulated | 0.092 | 2.010 |
| RP4-631H13.6 | downregulated | 0.030 | 2.700 |
| MT1L | downregulated | 0.293 | 1.240 |
| ZNF204P | downregulated | 0.202 | 1.490 |
| RP11-693N9.2 | downregulated | 0.151 | 1.690 |
| TSSC2 | downregulated | 0.212 | 1.460 |
| RP11-111F5.2 | downregulated | 0.127 | 1.800 |
| RRN3P1 | downregulated | 0.018 | 3.000 |
| GOLGA6L5P | downregulated | 0.065 | 2.220 |
| HERC2P3 | downregulated | 0.476 | 0.879 |
| CECR7 | downregulated | 0.610 | 0.674 |
| UBE2Q2P1 | downregulated | 0.032 | 2.640 |
| LRRC37A4P | downregulated | 0.096 | 1.970 |
| RP11-798K23.5 | downregulated | 0.002 | 4.220 |
| WHAMMP3 | downregulated | 0.382 | 1.050 |
| FAR2P2 | downregulated | 0.020 | 2.940 |
| RP11-79D8.2 | downregulated | 0.349 | 1.110 |
| FABP5P7 | downregulated | 0.192 | 1.530 |
| RP11-632K20.7 | downregulated | 0.163 | 1.640 |
| PPP4R1L | downregulated | 0.124 | 1.820 |
| OR2A20P | downregulated | 0.278 | 1.270 |
| HERC2P9 | downregulated | 0.235 | 1.390 |
| HERC2P2 | downregulated | 0.183 | 1.560 |
| HMGN2P15 | downregulated | 0.179 | 1.570 |
| RCN1P2 | downregulated | 0.636 | 0.638 |
| CDC42EP3 | downregulated | 0.249 | 1.350 |
Table 2.
Correlation of Pseudogene-Derived RNA RP11-480I12.5 Expression in TCGA Breast Cancer Tissue with Patients’ Clinicopathological Characteristics
|
RP11-480I12.5 Expression |
|||
|---|---|---|---|
| Variables | N = 1,096 | Low/High Expression Case (n) | p Value |
| Age at diagnosis | |||
| ≥60 | 510 | 361/149 | 0.0717 |
| <60 | 586 | 385/201 | |
| Estrogen receptor status | |||
| Positive | 806 | 584/222 | <0.0001 |
| Negative | 240 | 127/113 | |
| NAa | 50 | 35/15 | |
| Progesterone receptor status | |||
| Positive | 698 | 515/183 | <0.0001 |
| Negative | 345 | 194/151 | |
| NA | 53 | 37/16 | |
| Her2 receptor status | |||
| Positive | 161 | 81/80 | <0.0001 |
| Negative | 564 | 424/140 | |
| NA | 371 | 241/130 | |
| T stage | |||
| T1/T2 | 915 | 626/289 | 0.5783 |
| T3/T4 | 178 | 118/60 | |
| TX | 3 | 2/1 | |
| N stage | |||
| N0/N1 | 880 | 618/262 | 0.0017 |
| N2/N3 | 196 | 115/81 | |
| NX | 20 | 13/7 | |
| M stage | |||
| M0 | 910 | 622/288 | 0.0443 |
| M1 | 21 | 10/11 | |
| MX | 165 | 114/51 | |
| Pathologic stage | |||
| I/II | 806 | 566/240 | 0.0054 |
| III/IV | 267 | 163/104 | |
| NA | 23 | 17/6 | |
NA, not applicable
RP11-480I12.5 Contributes to Cell Proliferation In Vitro and Tumor Growth In Vivo
To study the effects of RP11-480I12.5 on tumorigenesis of breast cancer, we employed small interfering RNA (siRNA) targeting RP11-480I12.5 to suppress RP11-480I12.5 expression in MCF-7 (Figure 2A) and MDA-MB-231 (Figure 2B). Knockdown of RP11-480I12.5 significantly inhibited in vitro cell proliferation of MCF-7 (Figure 2C) and MDA-MB-231 (Figure 2D). Colonies of MCF-7 (Figure 2E) and MDA-MB-231 (Figure 2F) were also obviously reduced in siRNA-treated group compared with negative control (NC) group. Apoptosis suppression together with deregulated proliferation lead to the uncontrolled expansion of cancer.17 Thus, the role of RP11-480I12.5 in apoptosis of breast cancer cell was detected. Intriguingly, after decrease of RP11-480I12.5 expression, apoptosis rates of MCF-7 (Figures 2G–2I) and MDA-MB-231 (Figures 2J–2L) were markedly upregulated. In vivo assay also revealed that xenograft tumors from the group with RP11-480I12.5 inhibition had smaller tumor size, weight, and volume (Figures 2M–2P). Four RNA transcripts (001, 002, 003, and 004) of pseudogene RP11-480I12.5 were identified as shown in Figure 3A. Overexpression of RP11-480I12.5 transcripts in MCF-7 (Figure 3B) or MDA-MB-231 (Figure 3C) was achieved by transfecting corresponding overexpressed plasmids and the effect was determined by quantitative real-time PCR. Subsequently, the functions of the four lncRNAs in growth and tumorigenesis of breast cancer were successively studied by cell counting assay, colony formation assay, and cell apoptosis assay. As presented in Figures 3D–3F, among the four lncRNAs, overexpression of RP11-480I12.5-004 significantly facilitated cell growth (Figure 3D) and colony formation (Figure 3E) and impeded cell apoptosis (Figure 3F) of breast cancer in vitro. However, the other three lncRNA transcripts only exhibited extremely weak effects on these behaviors of breast cancer. In vivo tumor growth assay supported that RP11-480I12.5-004 overexpression induced tumor growth of breast cancer in vivo (Figures 3G–3J). All these findings suggest that RP11-480I12.5-004 enhances breast cancer cell proliferation in vitro and tumor growth in vivo.
Figure 2.
Knockdown of RP11-480I12.5 Inhibits Growth of Breast Cancer In Vitro and In Vivo
(A) The knockdown effect of siRNA targeting RP11-480I12.5 in MCF-7. (B) The knockdown effect of siRNA targeting RP11-480I12.5 in MDA-MB-231. (C) Knockdown of RP11-480I12.5 suppresses proliferation of MCF-7. (D) Knockdown of RP11-480I12.5 suppresses proliferation of MDA-MB-231. (E) Knockdown of RP11-480I12.5 suppresses colony formation of MCF-7. (F) Knockdown of RP11-480I12.5 suppresses colony formation of MDA-MB-231. (G–I) Knockdown of RP11-480I12.5 promotes apoptosis of MCF-7. (J–L) Knockdown of RP11-480I12.5 promotes apoptosis of MDA-MB-231. (M) Knockdown of RP11-480I12.5 reduces tumor size of breast cancer in mice. (N) The expression of RP11-480I12.5 in tumor samples isolated from si-NC-treated group and si-RP11-480I12.5-treated group. (O) Knockdown of RP11-480I12.5 decreases tumor weight of breast cancer in mice. (P) Knockdown of RP11-480I12.5 inhibits tumor growth in vivo. “ns” represents no significant difference; ∗p < 0.05.
Figure 3.
Overexpression of RP11-480I12.5-004 Enhances Growth of Breast Cancer In Vitro and In Vivo
(A) The transcript isoforms of pseudogene RP11-480I12.5. (B) The overexpression effects of plasmids carrying four transcripts of pseudogene RP11-480I12.5 in MCF-7. (C) The overexpression effects of plasmids carrying four transcripts of pseudogene RP11-480I12.5 in MDA-MB-231. (D) Overexpression of RP11-480I12.5 transcripts influence cell growth of breast cancer in vitro. (E) Overexpression of RP11-480I12.5 transcripts influence colony formation of breast cancer in vitro. (F) Overexpression of RP11-480I12.5 transcripts influence cell apoptosis of breast cancer in vitro. (G) Overexpression of RP11-480I12.5-004 increases tumor size of breast cancer in mice. (H) The expression of RP11-480I12.5-004 in tumor samples isolated from OE-NC-treated group and OE-RP11-480I12.5-004-treated group. (I) Overexpression of RP11-480I12.5-004 largens tumor weight of breast cancer in mice. (J) Overexpression of RP11-480I12.5-004 facilitates tumor growth in vivo. “ns” represents no significant difference; ∗p < 0.05.
RP11-480I12.5-004 Directly Binds to miR-29c-3p in Breast Cancer
Next, we intended to uncover the possible mechanism of RP11-480I12.5-004 in promoting breast cancer cell proliferation and tumor growth. Other groups and our team previously showed that competing endogenous RNA (ceRNA) mechanism is a key action mode of pseudogene-derived RNA.15,18 To ascertain whether RP11-480I12.5-004 exerts its roles in breast cancer by ceRNA mechanism, we performed a series of studies. First of all, we investigated the subcellular location of RP11-480I12.5-004 using the lncLocator (http://www.csbio.sjtu.edu.cn/bioinf/lncLocator/). As depicted in Figure 4A, RP11-480I12.5-004 mainly located in cytoplasm (65.5%) and cytosol (14.3%). RP11-480I12.5-004 was also experimentally confirmed to be mainly located in cytoplasm (Figure 4B). Subsequently, 27 microRNAs (miRNAs) that may potentially bind to RP11-480I12.5-004 were predicted through starBase database and a RP11-480I12.5-004-miRNA network was established by Cytoscape software (Figure 4C). Expression correlation of RP11-480I12.5 and miRNA in breast cancer was assessed (Table 3). Based on ceRNA mechanism, there should be an inverse relationship between RP11-480I12.5 and miRNA. Among the 27 miRNAs, only three miR-29 family members, including miR-29a-3p, miR-29b-3p, and miR-29c-3p, were found to be significantly negatively associated with RP11-480I12.5 expression in breast cancer (Figures 4D–4F). Survival analysis for miR-29a-3p (Figure 4G; Figures S1A–S1D), miR-29b-3p (Figure 4H; Figures S1E–S1H), and miR-29c-3p (Figure 4I; Figures S1I–S1L) was conducted using data from four datasets (GSE19783, GSE40267, TCGA, and METABRIC). The results revealed that only high expression of miR-29c-3p indicated favorable prognosis of patients with breast cancer in all the four datasets, supporting that miR-29c-3p is the most potential binding miRNA of RP11-480I12.5-004. The binding sites of RP11-480I12.5-004 and miR-29c-3p were shown in Figure 4J. Wild-type (WT) and mutant (mut) 3′ UTR sequences of RP11-480I12.5-004 were cloned into psi-CHECK2 vector. Dual-luciferase reporter assay demonstrated that luciferase activity of the reporter with WT-RP11-480I12.5-004 was significantly decreased by miR-29c-3p mimic transfection while the activity of the reporter with mut-RP11-480I12.5-004 showed no statistical change (Figure 4K). These results indicate that RP11-480I12.5-004 interacts with miR-29c-3p by direct binding.
Figure 4.
miR-29c-3p Directly Binds to RP11-480I12.5-004 in Breast Cancer and Reverses the Effect of RP11-480I12.5-004 on Breast Cancer Cell Growth
(A and B) The cellular location of RP11-480I12.5-004. (C) The potential miRNAs binding to RP11-480I12.5-004. (D) The expression correlation of miR-29a-3p and RP11-480I12.5 in breast cancer. (E) The expression correlation of miR-29b-3p and RP11-480I12.5 in breast cancer. (F) The expression correlation of miR-29c-3p and RP11-480I12.5 in breast cancer. (G) The prognostic value of miR-29a-3p in breast cancer. (H) The prognostic value of miR-29b-3p in breast cancer. (I) The prognostic value of miR-29c-3p in breast cancer. (J) The potential binding sites of miR-29c-3p with RP11-480I12.5. (K) Dual-luciferase reporter assay confirms that miR-29c-3p directly binds to RP11-480I12.5. (L) The overexpression effect of miR-29c-3p mimic in MCF-7. (M) Overexpression of miR-29c-3p reverses the effect of RP11-480I12.5-004 on cell growth in breast cancer. (N) Overexpression of miR-29c-3p reverses the effect of RP11-480I12.5-004 on colony formation in breast cancer. (O) Overexpression of miR-29c-3p reverses the effect of RP11-480I12.5-004 on cell growth in breast cancer. (P) The knockdown effect of miR-29c-3p inhibitor in MDA-MB-231. (Q) Silencing miR-29c-3p reverses the effect of RP11-480I12.5 on cell growth in breast cancer. (R) Silencing miR-29c-3p reverses the effect of RP11-480I12.5 on colony formation in breast cancer. (S) Silencing miR-29c-3p reverses the effect of RP11-480I12.5 on apoptosis in breast cancer. “ns” represents no significant difference; ∗p < 0.05.
Table 3.
The Expression Correlation of RP11-480I12.5 and Predicted miRNAs Determined by starBase
| miRNA Name | R | p Value |
|---|---|---|
| hsa-miR-339-5p | 0.097 | 1.34E-03 |
| hsa-miR-148a-3p | 0.044 | 1.52E-01 |
| hsa-miR-152-3p | −0.07 | 2.02E-02 |
| hsa-miR-148b-3p | 0.044 | 1.43E-01 |
| hsa-miR-365a-3p | 0.128 | 2.46E-05 |
| hsa-miR-365b-3p | 0.128 | 2.40E-05 |
| hsa-miR-500a-3p | 0.216 | 7.02E-13 |
| hsa-miR-212-5p | −0.028 | 3.63E-01 |
| hsa-miR-1197 | 0.028 | 3.58E-01 |
| hsa-miR-3064-5p | 0.078 | 1.04E-02 |
| hsa-miR-6504-5p | −0.023 | 4.51E-01 |
| hsa-miR-29a-3p | -0.157 | 1.94E-07 |
| hsa-miR-29c-3p | -0.226 | 4.92E-14 |
| hsa-miR-29b-3p | -0.164 | 5.91E-08 |
| hsa-miR-670-5p | −0.021 | 4.88E-01 |
| hsa-miR-345-3p | 0.165 | 4.87E-08 |
| hsa-miR-653-5p | −0.032 | 2.99E-01 |
| hsa-miR-3918 | −0.009 | 7.62E-01 |
| hsa-miR-3622b-5p | −0.045 | 1.41E-01 |
| hsa-miR-378 g | −0.004 | 9.01E-01 |
| hsa-miR-1287-5p | 0.139 | 4.66E-06 |
| hsa-miR-2682-5p | 0.039 | 2.04E-01 |
| hsa-miR-34b-5p | −0.059 | 5.11E-02 |
| hsa-miR-449c-5p | 0.063 | 3.85E-02 |
| hsa-miR-214-3p | 0.03 | 3.25E-01 |
| hsa-miR-761 | 0 | 1.00E+00 |
| hsa-miR-3619-5p | 0.024 | 4.29E-01 |
miR-29c-3p Reverses the Effect of RP11-480I12.5-004 on Breast Cancer Growth through Targeting AKT3 and CDK6
miR-29c-3p has been confirmed as a direct target of RP11-480I12.5-004. Next, we further studied the function and potential mechanism of RP11-480I12.5/miR-29c-3p interaction in growth of breast cancer. The overexpression of miR-29c-3p in MCF-7 was achieved by transfection of miR-29c-3p mimic (Figure 4L). Increased miR-29c-3p expression reversed the effect of RP11-480I12.5-004 upregulation on breast cancer cell proliferation (Figure 4M) and colony formation (Figure 4N). As shown in Figure 4O, the inhibitory role of RP11-480I12.5-004 in apoptosis of breast cancer was also reversed by transfecting miR-29c-3p mimic. miR-29c-3p inhibitor was introduced to suppress miR-29c-3p expression in MDA-MB-231 (Figure 4P). Moreover, expectedly, the effects of RP11-480I12.5 knockdown on breast cancer cell proliferation (Figure 4Q), colony formation (Figure 4R), and cell apoptosis (Figure 4S) were reversed after inhibition of miR-29c-3p. To explore the downstream molecular mechanism of RP11-480I12.5/miR-29c-3p axis, we predicted the potential target genes of miR-29c-3p by miRNet and found 254 targets. The 254 genes were entered into Enrichr database for Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis. The top 10 enriched pathways ranked by p value were presented in Figure 5A. Three candidate pathways, including pathways in cancer, PI3K-Akt signaling pathway, and breast cancer, were selected for subsequent analysis. Four genes (PTEN, AKT2, AKT3, and CDK6) were commonly appeared in the three pathways. The predicted binding sites between miR-29c-3p and these targets were presented in Figure S2. As shown in Figure 5B, luciferase activities of the reporters with WT-AKT3 and WT-CDK6 were decreased by transfection of miR-29c-3p mimic whereas luciferase activities of the reporters with WT-PTEN and WT-AKT2 were not significantly changed. No statistical change of luciferase activity of the reporters with mut-AKT3 or mut-CDK6 was observed when miR-29c-3p expression was upregulated (Figure 5C), suggesting that, among the four genes, only AKT3 and CDK6 are direct targets of miR-29c-3p. Expression correlation analysis also demonstrated that miR-29c-3p expression was markedly negatively correlated with expression of AKT3 (Figure S3A) and CDK6 (Figure S3B). Moreover, the role of RP11-480I12.5-004/miR-29c-3p axis in regulating AKT3 or CDK6 expression was further evaluated. Increase of miR-29c-3p expression could reverse the upregulated effect of RP11-480I12.5-004 overexpression on AKT3 level (Figure 5D), and knockdown of miR-29c-3p reversed the inhibitory effect of RP11-480I12.5 suppression on AKT3 expression (Figure 5E). For CDK6, similar results were obtained as presented in Figures 5F and 5G. MS2-RNA immunoprecipitation (RIP) assay was also indicated that the ceRNA crosstalk between RP11-480I12.5-004 and CDK6 or AKT3 may be mediated by miR-29c-3p at least in part (Figure 6A). Based on all these findings, we supposed an action model, in which overexpressed RP11-480I12.5-004 acts as a ceRNA to bind to miR-29c-3p to relieve its suppression on AKT3 and CDK6 expression, thereby facilitating growth and tumorigenesis of breast cancer (Figure 6B).
Figure 5.
AKT3 and CDK6 Are Two Direct Targets of miR-29c-3p in Breast Cancer
(A) The top 10 enriched KEGG pathways of target genes of miR-29c-3p and four potential target genes (PTEN, AKT2, AKT3, and CDK6) are identified. (B) miR-29c-3p suppresses luciferase activity of the reporters containing the WT AKT3 and CDK6 but not PTEN and AKT2. (C) miR-29c-3p can’t inhibit luciferase activity of the reporters containing the mut type AKT3 and CDK6. (D) Overexpression of RP11-480I12.5 increases AKT3 expression, and upregulation of miR-29c-3p reverses the role of RP11-480I12.5 overexpression in increasing AKT3 expression in MCF-7. (E) Knockdown of RP11-480I12.5 reduces AKT3 expression and silencing miR-29c-3p reverses the role of RP11-480I12.5 inhibition in decreasing AKT3 expression in MDA-MB-231. (F) Overexpression of RP11-480I12.5 increases CDK6 expression, and upregulation of miR-29c-3p reverses the role of RP11-480I12.5 overexpression in increasing CDK6 expression in MCF-7. (G) Knockdown of RP11-480I12.5 reduces CDK6 expression and silencing miR-29c-3p reverses the role of RP11-480I12.5 inhibition in decreasing CDK6 expression in MDA-MB-231. “ns” represents no significant difference; ∗p < 0.05.
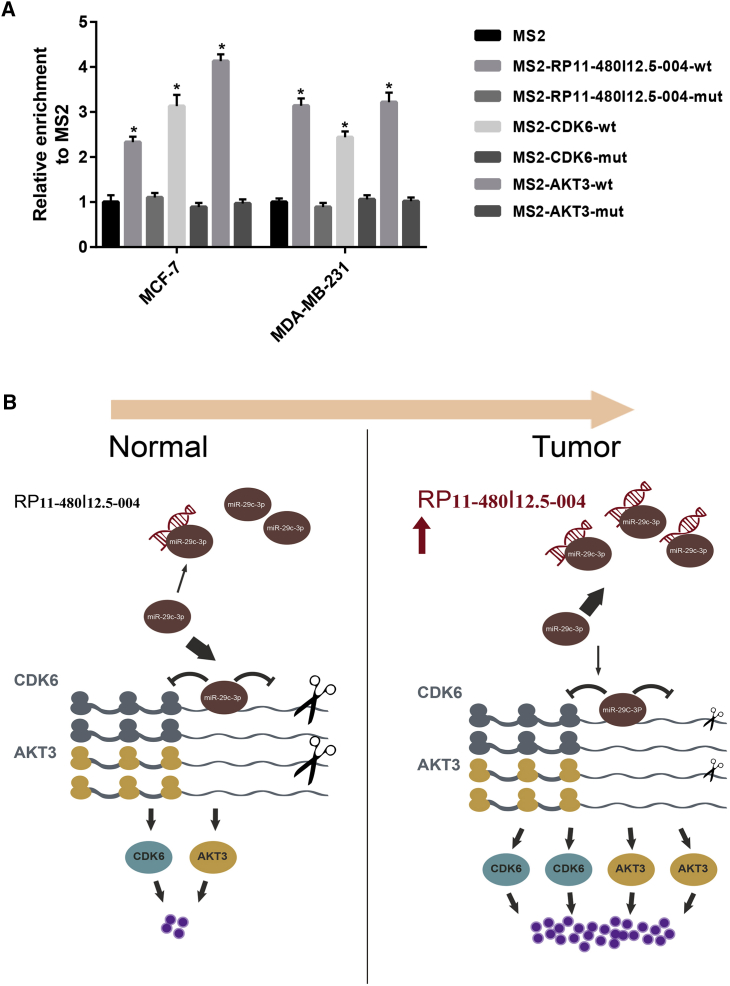
Figure 6.
RP11-480I12.5-004 Promotes Tumorigenesis of Breast Cancer by Relieving miR-29c-3p-Mediated AKT3 and CDK6 Degradation
(A) MS2-RIP assay confirms miR-29c-3p binding to RP11-480I12.5-004, CDK6, and AKT3 in breast cancer cells. (B) The model of the RP11-480I12.5-004/miR-29c-3p/CDK6/AKT3 axis and its role in tumorigenesis of breast cancer. ∗p < 0.05.
TUBA1B and TUBA1C Are Two Potential Cognate Genes of RP11-480I12.5 in Breast Cancer
Pseudogenes have been widely documented to interact with their cognate genes.19 In this study, we preliminarily investigated the cognate genes of RP11-480I12.5. Six cognate genes (TUBA1B, TUBA1C, TUBA3C, TUBA3D, TUBA4A, and TUBA4B) of RP11-480I12.5 were identified by basic local alignment search tool (BLAST) analysis (Figure 7A). Next, the expression correlation between RP11-480I12.5 and cognate gene was assessed using TCGA breast cancer data. As shown in Figure 7B, only two cognate genes (TUBA1B and TUBA1C) were significantly positively associated with expression of RP11-480I12.5 (R > 0.1 and p < 0.05). Survival analysis revealed that breast cancer patients with higher expression of TUBA1B and TUBA1C had poorer prognosis (Figure 7C). For the other 4 cognate genes of RP11-480I12.5, no significant effects on survival of breast cancer were observed. Subsequently, the mRNA expression levels of TUBA1B, TUBA1C, TUBA3C, TUBA3D, TUBA4A, and TUBA4B in breast cancer were determined by GEPIA database (Figure 7D). The result showed obvious upregulation of TUBA1B and TUBA1C in breast cancer samples when compared with normal breast samples. Furthermore, the protein expression levels of TUBA1B and TUBA1C in breast cancer tissue were markedly higher than that in normal controls (Figure 7E). Taken together, TUBA1B and TUBA1C are two potential functional cognate genes of RP11-480I12.5 in breast cancer. However, the regulatory effect of RP11-480I12.5 on TUBA1B or TUBA1C in breast cancer need to be further experimentally validated in the future.
Figure 7.
Exploration of the Relationship between RP11-480I12.5 and Its Cognate Genes in Breast Cancer
(A) The cognate genes (TUBA1B, TUBA1C, TUBA3C, TUBA3D, TUBA4A, and TUBA4B) of RP11-480I12.5. (B) The expression correlation of RP11-480I12.5 and its cognate genes in breast cancer. (C) The prognostic values of TUBA1B, TUBA1C, TUBA3C, TUBA3D, TUBA4A, and TUBA4B in breast cancer. (D) The mRNA expression of TUBA1B, TUBA1C, TUBA3C, TUBA3D, TUBA4A, and TUBA4B in breast cancer samples compared with normal breast samples determined by GEPIA. (E) The protein expression of TUBA1B and TUBA1C in breast cancer samples compared with normal breast samples determined by HumanProteinAtlas. Scale bar, 150 μm. ∗p < 0.05.
Discussion
Recently, pseudogene-expressed lncRNAs, a special class of lncRNAs, have been reported to serve as oncogenic and tumor-suppressive molecules in cancer.20,21 However, expression, functions, and mechanisms of pseudogene-expressed lncRNAs in breast cancer remain largely unknown. In this study, by in silico analysis, we first identified a novel pseudogene-expressed lncRNA RP11-480I12.5 that was upregulated in breast cancer and correlated with cancer progression and poor prognosis of patients with breast cancer.
It is known to all that uncontrolled cell proliferation/growth and suppression of cell apoptosis are two key hallmarks of cancer. Functional assays revealed that pseudogene-expressed lncRNA RP11-480I12.5-004 led to promotion of proliferation and inhibition of apoptosis of breast cancer, suggesting that RP11-480I12.5-004 plays oncogenic roles in carcinogenesis of breast cancer.
Next, we explored the possible mechanism of RP11-480I12.5-004 in breast cancer. ceRNA model is a key action mechanism of lncRNA, in which cytoplasm-located lncRNA competitively binds to miRNA, thereby relieving suppression of downstream targets.22, 23, 24 RP11-480I12.5-004 was mainly located in cytoplasm predicted by lncLocator, which is a subcellular localization predictor for lncRNAs based on a stacked ensemble classifier.25
Subsequently, using starBase, 27 miRNAs that potentially bind to RP11-480I12.5-004 were obtained, after which expression correlation between RP11-480I12.5-004 and miRNA in breast cancer was performed. According to ceRNA mechanism, RP11-480I12.5 in breast cancer, there should be a negative relationship between lncRNA and miRNA.26 The results revealed that only miR-29a-3p, miR-29b-3p, and miR-29c-3p were significantly negatively correlated with RP11-480I12.5. Furthermore, we also assessed the prognostic values of the 3 miRNAs in breast cancer by usage of four public datasets, finding that high expression of miR-29c-3p indicated favorable prognosis in all these datasets. Li et al.27 showed that miR-29c-3p inhibited proliferation, migration, invasion, and colony formation of breast cancer through targeting the TIMP3/STAT1/FOXO1 pathway, supporting that miR-29c-3p plays a suppressive role in breast cancer. However, several studies have demonstrated the oncogenic functions of miR-29a-3p and miR-29b-3p in breast cancer. For example, miR-29a-3p promoted proliferation and epithelial-mesenchymal transition (EMT) in breast cancer28 and miR-29b-3p enhanced progression of MDA-MB-231 TNBC cells.29 Taken together, miR-29c-3p may be the potential binding miRNA of RP11-480I12.5-004, which was subsequently confirmed by a dual-luciferase reporter assay. Functional experiments revealed that alteration of miR-29c-3p expression reversed the effects of RP11-480I12.5-004 on growth of breast cancer.
miRNAs are involved in biological processes via negative regulation of target gene expression in post-transcriptional levels.30,31 Through miRNet, 254 targets of miR-29c-3p were predicted. These targets were significantly enriched in some breast-cancer-associated pathways, such as pathways in cancer, PI3K-Akt signaling pathway, and breast cancer.32 Four genes, PTEN, AKT2, AKT3, and CDK6, commonly appeared in the three pathways and were selected for subsequent analysis. Dual-luciferase reporter assay indicated that AKT3 and CDK6 were two direct targets of miR-29c-3p. Both AKT3 and CDK6 have been previously reported to promote growth of breast cancer.33,34 MS2-RIP assay also showed miR-29c-3p-mediated the ceRNA crosstalk between RP11-480I12.5-004 and CDK6 or AKT3 in breast cancer cells.
Lots of studies have demonstrated that pseudogenes and pseudogene-expressed RNAs exert their oncogenic or tumor-suppressive roles in human cancers by regulating expression of parental genes.19 Therefore, we identified six parental genes of RP11-480I12.5 by sequence blast. Correlation analysis, survival analysis, and expression analysis together indicated that TUBA1B and TUBA1C might be two potential oncogenic parental genes of RP11-480I12.5-004 in breast cancer. TUBA1C has been confirmed to act as an oncogene in migration and proliferation of hepatocellular carcinoma.35 All these findings suggest that oncogenic roles of RP11-480I12.5-004 in breast cancer may partially result from regulation of TUBA1B and TUBA1C expression. However, more corresponding functional and mechanistic assays need to be further conducted to validate these results.
Taken together, pseudogene-expressed lncRNA RP11-480I12.5-004, upregulated in breast cancer and associated with cancer progression and poor prognosis, promotes growth and tumorigenesis of breast cancer by increasing AKT3 and CDK6 expression through competitively binding to miR-29c-3p.
Conclusions
In a word, we validate RP11-480I12.5-004 as an oncogenic pseudogene-expressed lncRNA that is increased in breast cancer and acts as molecular sponge of miR-29c-3p to enhance AKT3 and CDK6 expression, consequently promoting growth and tumorigenesis of breast cancer. In addition, we also identify that TUBA1B and TUBA1C are two potential parental genes of RP11-480I12.5 in breast cancer. Targeting RP11-480I12.5-004/miR-29c-3p/AKT3/CDK6 axis may represent a promising therapeutic method for patients with breast cancer in the future.
Materials and Methods
dreamBase Analysis
dreamBase, an integrated platform for analyzing regulatory features of pseudogenes from multi-dimensional high-throughput sequencing data, was used to obtain the DEPs between breast cancer samples and normal samples.36 Fold change (tumor/normal) >2 was set as the criterion for selecting DEPs.
GEPIA Analysis
GEPIA is a newly developed interactive web tool for analyzing the RNA sequencing expression data from the TCGA and the GTEx projects, providing customizable functions, such as tumor/normal differential expression analysis, profiling according to pathological stages, patient survival analysis, and correlation analysis.37 GEPIA was employed to validate the expression of DEPs identified by dreamBase. Fold change (tumor/normal) >2 and p <0.05 were set as the thresholds for identifying potential pseudogenes. The expression difference of DEPs among various major stage in breast cancer and the prognostic values of DEPs in breast cancer were also determined by GEPIA. GEPIA was finally introduced to perform expression correlation of RP11-480I12.5 and its cognate genes. p <0.05 was considered as significant.
starBase Analysis
starBase is an open-source database for studying miRNA-ceRNA, miRNA-ncRNA, and protein-RNA interaction networks from large-scale CLIP-seq (crosslinking-immunprecipitation and high-throughput) data.38 starBase was utilized to predict the potential miRNAs that may bind to RP11-480I12.5. Moreover, expression correlation of RP11-480I12.5 with miRNA was also detected by starBase. R <−0.1 and p <0.05 were regarded as statistically significant.
Kaplan-Meier Plotter Analysis
The prognostic values of miRNAs and genes in breast cancer were assessed through Kaplan-Meier plotter, which is capable of accessing the effect of miRNAs and genes on survival in 21 cancer types.39 For miRNA analysis, four datasets containing GSE19783, GSE40267, TCGA, and METABRIC were included. Log rank p <0.05 was considered as statistically significant.
Enrichr Analysis
Target genes of miR-29c-3p were predicted by miRNet, which is a comprehensive database for miRNA-related studies.40 The KEGG pathway enrichment analysis for the target genes of miR-29c-3p was conducted by Enrichr database as we previously described.41 The bar plot of the top 10 enriched KEGG pathways was automatically generated and the enriched genes of each KEGG pathways were directly downloaded from the webpage.
HumanProteinAtlas Analysis
The protein expression levels of TUBA1B and TUBA1C were assessed using HumanProteinAtlas database, which is a proteomic resource for studying protein expression and prognosis.42
Cell Culture and Clinical Samples
Human breast cell line (HBL-100) and breast cancer cell lines (MCF-7, T47D, BT474, HCC1937, BT549, and MDA-MB-231) were purchased from the cell bank of Chinese Scientific Academy (Shanghai, China). MCF-7, T47D, HCC1937, and BT549 were maintained in Roswell Park Memorial Institute (RPMI) 1640 medium (GIBCO) supplemented with 10% fetal bovine serum (FBS; Biological Industries) and HBL-100, BT474, and MDA-MB-231 were cultured in Dulbecco’s modified Eagle’s medium (DMEM; GIBCO) containing 10% FBS under a humidified atmosphere of 5% CO2 at 37°C. 60 paired breast cancer tissues and their adjacent non-cancerous tissues were collected from patients who underwent surgery at First Affiliated Hospital, College of Medicine, Zhejiang University (Hangzhou, China). Informed consent was signed by every patient. This study has been approved by the Ethics Committee of First Affiliated Hospital, College of Medicine, Zhejiang University.
siRNA, miRNA Mimic, Inhibitor, Plasmids, and Cell Transfection
siRNA targeting RP11-480I12.5 and NC, the plasmids and corresponding NC vectors, miRNA mimic, inhibitor, and their NCs used in this study were designed and purchased from RiboBio (Guangzhou, China). RP11-480I12.5 suppression and overexpression were achieved by transfecting siRNA targeting RP11-480I12.5 and overexpression vectors carrying RP11-480I12.5 using Lipofectamine 3000 (Invitrogen, USA) at final concentrations of 50 μM and 2 μg/mL, respectively. miRNA mimic (50 μM) or inhibitor (100 μM) were transfected into cells to increase or decrease miRNA expression in accordance with the manufacturer’s instructions. The sequence of siRNA targeting RP11-480I12.5 was shown in Table S1.
RNA Extraction and Quantitative Real-Time PCR
Total RNAs of cells and tissues were extracted using Trizol reagent (Invitrogen, USA). Quantitative real-time PCR was performed as we previously described.43, 44, 45 GAPDH and U6 were used as endogenous controls for pseudogene/gene and miRNA, respectively. The primers used in this study were listed in Table S1.
Cell Counting Assay
In vitro cell growth was evaluated through a cell counting assay. First, MCF-7 and MDA-MB-231 cells were transfected as mentioned above. Subsequently, 5 × 104 of pre-transfected cells were re-plated into six-well plates. After culturing for 72 h, cell number of each well was counted. The results were shown as mean ± SD.
CCK-8 Assay
2 × 103 of MCF-7 and MDA-MB-231 cells were seeded into 96-well plates. 12 h later, cells were transfected as mentioned above. 12 h later, medium of each well was replaced with refresh medium. At 24, 48, 72, and 96 h post infection, 0.02 mL CCK-8 solution was added into each well and incubated for 2 (MCF-7) or 4 (MDA-MB-231) h at 37°C. The absorbance of each well was determined at 450 nm through a microplate reader (Bio-Rad, USA).
Colony Formation Assay
After transfected for 12 h, 1 × 103 of MCF-7 and MDA-MB-231 cells were re-seeded into six-well plates and cultured for 10 days (MCF-7) and 2 weeks (MDA-MB-231). At the end of culture, six-well plates were washed using PBS two times. Subsequently, colonies were fixed in methanol for 15–30 min and stained with 0.1% crystal violet solution for 15 min. The visible colonies of individual wells were carefully counted.
Apoptosis Assay
Pre-transfected cells were re-seeded into six-well plates and cultured for 48–72 h. Then, an Annexin fluorescein isothiocyanate/propidium iodide (FITC/PI) apoptosis detection kit (Beyotime, China) was employed to perform FITC and PI double labeling according to the manufacturer’s instructions. Finally, the percentage of apoptotic cells was measured by a flow cytometry (BD Biosciences, USA).
Dual-Luciferase Reporter Assay
1 × 104 of 293T cells were seeded into 96-well plates. 12 h later, these cells were co-transfected with miR-29c-3p mimic or NC (50 nM) and 100 ng of reporter plasmid using Lipofectamine 3000 (Invitrogen, USA) in accordance with the manufacturer’s instructions. 48 h later, dual-luciferase reporter assay was performed by a Dual-Luciferase Reporter Assay Kit (Promega, USA) according to the protocol with a Varioskan Flash Spectral Scanning Multimode Reader (Thermo Fisher Scientific, USA).
MS2-RIP Assay
MS2-RIP assay was employed to determine whether endogenous miR-29c-3p could bind to RP11-480I12.5-004, CDK6, and AKT3 according to the previous reports using EZ-Magna RIP kit (Millipore) in accordance with the manufacturers’ instructions.18,46
Animal Studies
The female BALB/C nude mice (4–5 weeks old) were purchased from the Shanghai Laboratory Animal Center (Shanghai, China) and maintained in a specific-pathogen-free facility. 5 × 106 of pre-transfected MCF-7 cells and 2 × 106 of pre-transfected MDA-MB-231 cells were subcutaneously injected into the right back of nude mice (5 mice/group). Tumor volumes were calculated as the formula of 0.5 × length × width2 at 6, 9, 12, 15, 18, 21, 24, 27, and 30 days post-injection. Finally, these mice were sacrificed and the tumors were isolated and weighted. All animal experiments in this study were approved by the Ethics Committee of College of Medicine, Zhejiang University (Hangzhou, China).
Statistical Analysis
The statistical analysis of bioinformatics analyses was directly done by the online tools. GraphPad Prism 7 software was used to analyze all experimental data. Chi-square test and two-tailed Student’s t test were applied to analyze the statistical significance for qualitative data and quantitative data, respectively. p <0.05 was considered as significant.
Author Contributions
W.L. and P.F. designed this work. W.L. and B.D. performed experiments, analyzed data, and drafted the manuscript; G.Z. performed some experiments; W.F. revised the manuscript; and J.Y. collected clinical samples. All authors read and approved the final version of the manuscript.
Conflicts of Interest
The authors declare no competing interests.
Acknowledgments
This work was supported by grants from the National Natural Science Foundation of China (81874225) and the Natural Science Foundation of Zhejiang Province (LQ20H160030). The data in this work are available from the corresponding author on reasonable request.
Footnotes
Supplemental Information can be found online at https://doi.org/10.1016/j.omtn.2020.07.022.
Contributor Information
Weiyang Lou, Email: 11718264@zju.edu.cn.
Peifen Fu, Email: Peifen_Fu@163.com.
Supplemental Information
References
- 1.Harbeck N., Gnant M. Breast cancer. Lancet. 2017;389:1134–1150. doi: 10.1016/S0140-6736(16)31891-8. [DOI] [PubMed] [Google Scholar]
- 2.Li Y., Zeng Q., Qiu J., Pang T., Xian J., Zhang X. Long non-coding RNA UCA1 promotes breast cancer by upregulating PTP1B expression via inhibiting miR-206. Cancer Cell Int. 2019;19:275. doi: 10.1186/s12935-019-0958-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.de la Mare J.A., Contu L., Hunter M.C., Moyo B., Sterrenberg J.N., Dhanani K.C., Mutsvunguma L.Z., Edkins A.L. Breast cancer: current developments in molecular approaches to diagnosis and treatment. Recent Patents Anticancer Drug Discov. 2014;9:153–175. doi: 10.2174/15748928113086660046. [DOI] [PubMed] [Google Scholar]
- 4.Zhao P., Guan H., Dai Z., Ma Y., Zhao Y., Liu D. Long noncoding RNA DLX6-AS1 promotes breast cancer progression via miR-505-3p/RUNX2 axis. Eur. J. Pharmacol. 2019;865:172778. doi: 10.1016/j.ejphar.2019.172778. [DOI] [PubMed] [Google Scholar]
- 5.Tang T., Guo C., Xia T., Zhang R., Zen K., Pan Y., Jin L. LncCCAT1 Promotes Breast Cancer Stem Cell Function through Activating WNT/β-catenin Signaling. Theranostics. 2019;9:7384–7402. doi: 10.7150/thno.37892. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zheng S., Li M., Miao K., Xu H. lncRNA GAS5-promoted apoptosis in triple-negative breast cancer by targeting miR-378a-5p/SUFU signaling. J. Cell Biochem. 2019;121:2225–2235. doi: 10.1002/jcb.29445. [DOI] [PubMed] [Google Scholar]
- 7.Bhan A., Soleimani M., Mandal S.S. Long Noncoding RNA and Cancer: A New Paradigm. Cancer Res. 2017;77:3965–3981. doi: 10.1158/0008-5472.CAN-16-2634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Zhu L., Xu P.C. Downregulated LncRNA-ANCR promotes osteoblast differentiation by targeting EZH2 and regulating Runx2 expression. Biochem. Biophys. Res. Commun. 2013;432:612–617. doi: 10.1016/j.bbrc.2013.02.036. [DOI] [PubMed] [Google Scholar]
- 9.Qian C., Ye Y., Mao H., Yao L., Sun X., Wang B., Zhang H., Xie L., Zhang H., Zhang Y. Downregulated lncRNA-SNHG1 enhances autophagy and prevents cell death through the miR-221/222 /p27/mTOR pathway in Parkinson’s disease. Exp. Cell Res. 2019;384:111614. doi: 10.1016/j.yexcr.2019.111614. [DOI] [PubMed] [Google Scholar]
- 10.Zhan S., Wang K., Xiang Q., Song Y., Li S., Liang H., Luo R., Wang B., Liao Z., Zhang Y. lncRNA HOTAIR upregulates autophagy to promote apoptosis and senescence of nucleus pulposus cells. J. Cell Physiol. 2019;235:2195–2208. doi: 10.1002/jcp.29129. [DOI] [PubMed] [Google Scholar]
- 11.Xiao-Jie L., Ai-Mei G., Li-Juan J., Jiang X. Pseudogene in cancer: real functions and promising signature. J. Med. Genet. 2015;52:17–24. doi: 10.1136/jmedgenet-2014-102785. [DOI] [PubMed] [Google Scholar]
- 12.Yang X., Yang B. lncRNA PDIA3P regulates cell proliferation and invasion in non-small cell lung cancer. Exp. Ther. Med. 2019;18:3184–3190. doi: 10.3892/etm.2019.7882. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Feng J., Yang G., Liu Y., Gao Y., Zhao M., Bu Y., Yuan H., Yuan Y., Yun H., Sun M. LncRNA PCNAP1 modulates hepatitis B virus replication and enhances tumor growth of liver cancer. Theranostics. 2019;9:5227–5245. doi: 10.7150/thno.34273. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Zheng J., Zhang H., Ma R., Liu H., Gao P. Long non-coding RNA KRT19P3 suppresses proliferation and metastasis through COPS7A-mediated NF-kappaB pathway in gastric cancer. Oncogene. 2019;38:7073–7088. doi: 10.1038/s41388-019-0934-z. [DOI] [PubMed] [Google Scholar]
- 15.Lou W., Ding B., Fan W. High Expression of Pseudogene PTTG3P Indicates a Poor Prognosis in Human Breast Cancer. Mol. Ther. Oncolytics. 2019;14:15–26. doi: 10.1016/j.omto.2019.03.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Wang R., Zhang T., Yang Z., Jiang C., Seng J. Long non-coding RNA FTH1P3 activates paclitaxel resistance in breast cancer through miR-206/ABCB1. J. Cell Mol. Med. 2018;22:4068–4075. doi: 10.1111/jcmm.13679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Huang J.L., Cao S.W., Ou Q.S., Yang B., Zheng S.H., Tang J., Chen J., Hu Y.W., Zheng L., Wang Q. The long non-coding RNA PTTG3P promotes cell growth and metastasis via up-regulating PTTG1 and activating PI3K/AKT signaling in hepatocellular carcinoma. Mol. Cancer. 2018;17:93. doi: 10.1186/s12943-018-0841-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Wang M.Y., Chen D.P., Qi B., Li M.Y., Zhu Y.Y., Yin W.J., He L., Yu Y., Li Z.Y., Lin L. Pseudogene RACGAP1P activates RACGAP1/Rho/ERK signalling axis as a competing endogenous RNA to promote hepatocellular carcinoma early recurrence. Cell Death Dis. 2019;10:426. doi: 10.1038/s41419-019-1666-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.An Y., Furber K.L., Ji S. Pseudogenes regulate parental gene expression via ceRNA network. J. Cell. Mol. Med. 2017;21:185–192. doi: 10.1111/jcmm.12952. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Sun M., Nie F.Q., Zang C., Wang Y., Hou J., Wei C., Li W., He X., Lu K.H. The Pseudogene DUXAP8 Promotes Non-small-cell Lung Cancer Cell Proliferation and Invasion by Epigenetically Silencing EGR1 and RHOB. Mol. Ther. 2017;25:739–751. doi: 10.1016/j.ymthe.2016.12.018. [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- 21.Zhang G., Xu Y., Zou C., Tang Y., Lu J., Gong Z., Ma G., Zhang W., Jiang P. Long noncoding RNA ARHGAP27P1 inhibits gastric cancer cell proliferation and cell cycle progression through epigenetically regulating p15 and p16. Aging (Albany NY) 2019;11:9090–9110. doi: 10.18632/aging.102377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Wang X., Su R., Guo Q., Liu J., Ruan B., Wang G. Competing endogenous RNA (ceRNA) hypothetic model based on comprehensive analysis of long non-coding RNA expression in lung adenocarcinoma. PeerJ. 2019;7:e8024. doi: 10.7717/peerj.8024. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wang W., Lou W., Ding B., Yang B., Lu H., Kong Q., Fan W. A novel mRNA-miRNA-lncRNA competing endogenous RNA triple sub-network associated with prognosis of pancreatic cancer. Aging (Albany NY) 2019;11:2610–2627. doi: 10.18632/aging.101933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Zhong G., Lou W., Yao M., Du C., Wei H., Fu P. Identification of novel mRNA-miRNA-lncRNA competing endogenous RNA network associated with prognosis of breast cancer. Epigenomics. 2019;11:1501–1518. doi: 10.2217/epi-2019-0209. [DOI] [PubMed] [Google Scholar]
- 25.Cao Z., Pan X., Yang Y., Huang Y., Shen H.B. The lncLocator: a subcellular localization predictor for long non-coding RNAs based on a stacked ensemble classifier. Bioinformatics. 2018;34:2185–2194. doi: 10.1093/bioinformatics/bty085. [DOI] [PubMed] [Google Scholar]
- 26.Jia X., Shi Y., Zhu Y., Meng W., He L., Jia Y., Tong Z. Integrated Analysis of mRNA-miRNA-lncRNA ceRNA Network in Human HR+/Her-2- Breast Cancer and Triple Negative Breast Cancer. J. Comput. Biol. 2019;27:1055–1066. doi: 10.1089/cmb.2019.0152. [DOI] [PubMed] [Google Scholar]
- 27.Li W., Yi J., Zheng X., Liu S., Fu W., Ren L., Li L., Hoon D.S.B., Wang J., Du G. miR-29c plays a suppressive role in breast cancer by targeting the TIMP3/STAT1/FOXO1 pathway. Clin. Epigenetics. 2018;10:64. doi: 10.1186/s13148-018-0495-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Pei Y.F., Lei Y., Liu X.Q. MiR-29a promotes cell proliferation and EMT in breast cancer by targeting ten eleven translocation 1. Biochim. Biophys. Acta. 2016;1862:2177–2185. doi: 10.1016/j.bbadis.2016.08.014. [DOI] [PubMed] [Google Scholar]
- 29.Zhang B., Shetti D., Fan C., Wei K. miR-29b-3p promotes progression of MDA-MB-231 triple-negative breast cancer cells through downregulating TRAF3. Biol. Res. 2019;52:38. doi: 10.1186/s40659-019-0245-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Lou W., Liu J., Gao Y., Zhong G., Chen D., Shen J., Bao C., Xu L., Pan J., Cheng J. MicroRNAs in cancer metastasis and angiogenesis. Oncotarget. 2017;8:115787–115802. doi: 10.18632/oncotarget.23115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lou W., Liu J., Gao Y., Zhong G., Ding B., Xu L., Fan W. MicroRNA regulation of liver cancer stem cells. Am. J. Cancer Res. 2018;8:1126–1141. [PMC free article] [PubMed] [Google Scholar]
- 32.Guerrero-Zotano A., Mayer I.A., Arteaga C.L. PI3K/AKT/mTOR: role in breast cancer progression, drug resistance, and treatment. Cancer Metastasis Rev. 2016;35:515–524. doi: 10.1007/s10555-016-9637-x. [DOI] [PubMed] [Google Scholar]
- 33.Hu X., Wang J., He W., Zhao P., Ye C. MicroRNA-433 targets AKT3 and inhibits cell proliferation and viability in breast cancer. Oncol. Lett. 2018;15:3998–4004. doi: 10.3892/ol.2018.7803. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Zhang Y., Zhang H., Kang H., Huo W., Zhou Y., Zhang Y. Knockdown of long non-coding RNA HOST2 inhibits the proliferation of triple negative breast cancer via regulation of the let-7b/CDK6 axis. Int. J. Mol. Med. 2019;43:1049–1057. doi: 10.3892/ijmm.2018.3995. [DOI] [PubMed] [Google Scholar]
- 35.Wang J., Chen W., Wei W., Lou J. Oncogene TUBA1C promotes migration and proliferation in hepatocellular carcinoma and predicts a poor prognosis. Oncotarget. 2017;8:96215–96224. doi: 10.18632/oncotarget.21894. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Zheng L.L., Zhou K.R., Liu S., Zhang D.Y., Wang Z.L., Chen Z.R., Yang J.H., Qu L.H. dreamBase: DNA modification, RNA regulation and protein binding of expressed pseudogenes in human health and disease. Nucleic Acids Res. 2018;46:D85–D91. doi: 10.1093/nar/gkx972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Tang Z., Li C., Kang B., Gao G., Li C., Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–W102. doi: 10.1093/nar/gkx247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Li J.H., Liu S., Zhou H., Qu L.H., Yang J.H. starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res. 2014;42:D92–D97. doi: 10.1093/nar/gkt1248. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Lánczky A., Nagy Á., Bottai G., Munkácsy G., Szabó A., Santarpia L., Győrffy B. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res. Treat. 2016;160:439–446. doi: 10.1007/s10549-016-4013-7. [DOI] [PubMed] [Google Scholar]
- 40.Fan Y., Siklenka K., Arora S.K., Ribeiro P., Kimmins S., Xia J. miRNet - dissecting miRNA-target interactions and functional associations through network-based visual analysis. Nucleic Acids Res. 2016;44:W135-41. doi: 10.1093/nar/gkw288. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Lou W., Liu J., Ding B., Jin L., Xu L., Li X., Chen J., Fan W. Five miRNAs-mediated PIEZO2 downregulation, accompanied with activation of Hedgehog signaling pathway, predicts poor prognosis of breast cancer. Aging (Albany NY) 2019;11:2628–2652. doi: 10.18632/aging.101934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Pontén F., Schwenk J.M., Asplund A., Edqvist P.H. The Human Protein Atlas as a proteomic resource for biomarker discovery. J. Intern. Med. 2011;270:428–446. doi: 10.1111/j.1365-2796.2011.02427.x. [DOI] [PubMed] [Google Scholar]
- 43.Lou W., Chen J., Ding B., Chen D., Zheng H., Jiang D., Xu L., Bao C., Cao G., Fan W. Identification of invasion-metastasis-associated microRNAs in hepatocellular carcinoma based on bioinformatic analysis and experimental validation. J. Transl. Med. 2018;16:266. doi: 10.1186/s12967-018-1639-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lou W., Liu J., Ding B., Chen D., Xu L., Ding J., Jiang D., Zhou L., Zheng S., Fan W. Identification of potential miRNA-mRNA regulatory network contributing to pathogenesis of HBV-related HCC. J. Transl. Med. 2019;17:7. doi: 10.1186/s12967-018-1761-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Lou W., Liu J., Ding B., Xu L., Fan W. Identification of chemoresistance-associated miRNAs in breast cancer. Cancer Manag. Res. 2018;10:4747–4757. doi: 10.2147/CMAR.S172722. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Gong C., Maquat L.E. lncRNAs transactivate STAU1-mediated mRNA decay by duplexing with 3′ UTRs via Alu elements. Nature. 2011;470:284–288. doi: 10.1038/nature09701. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.