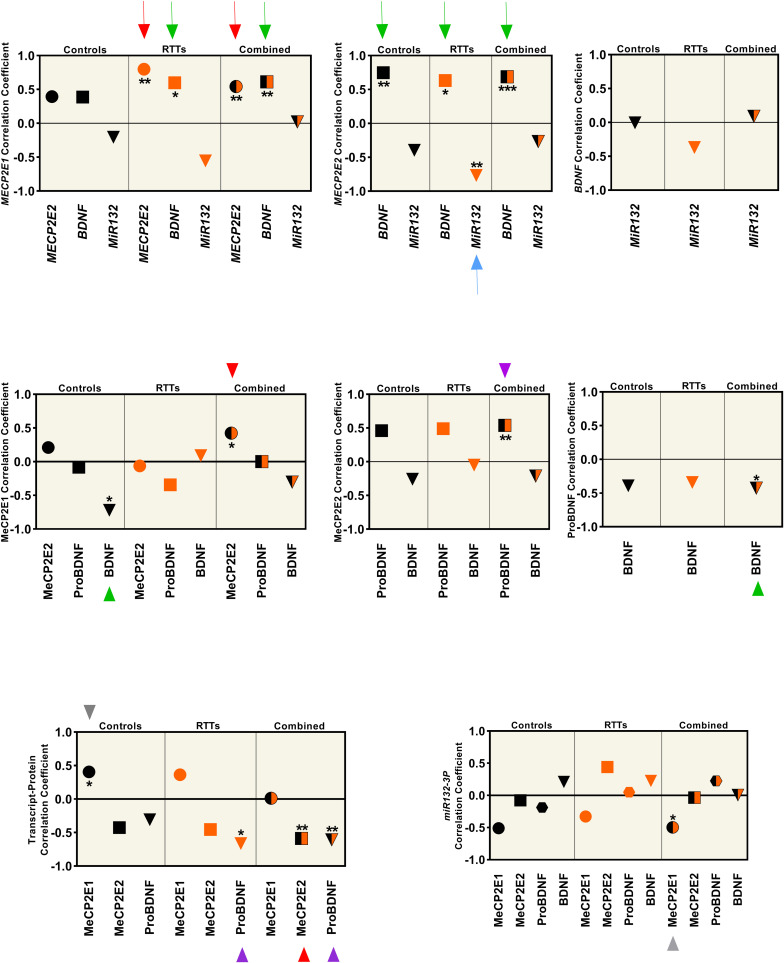
FIGURE 4.
Correlation analysis between all the studied parameters (MeCP2 isoforms, ProBDNF, BDNF, and miR132) from four brain regions (Frontal Cerebrum, Hippocampus, Amygdala, and Cerebellum) in transcript and protein level. All graphs represent the Pearson’s (for parametric variables), or Spearman’s (for non-parametric variables) correlation coefficient in controls (black), RTTs (orange), and controls and RTTs combined (black/orange). Statistical significance: ***P < 0.001; **P < 0.01; *P < 0.05; The charts are showing the data collected from four different brain regions (Frontal Cerebrum, Hippocampus, Amygdala and Cerebellum) of 3 patients and their age and sex matched controls (N = 3 controls + 3 RTTs). The two isoforms are positively correlated at the transcript level, which is significant for RTT and mixed data (red arrow), and MECP2E2 is negatively correlated with its protein, which is significant in mixed data (red arrowhead). Both isoforms show positive correlation with BDNF at the transcript level. Green arrows show the significant ones. BDNF protein has a significant negative correlation with MeCP2E1 in control samples (green arrowhead). ProBDNF is positively correlated with MeCP2E2, which is significant in mixed data (purple arrowhead). There is a negative correlation between ProBDNF and BDNF transcript in both RTT and mixed data (purple arrowhead). The only significant correlation of miR132 is a negative correlation with MECP2E2 in RTTs (blue arrow). Note that all miR132 transcripts in this Figure refer to miR132-3P strand.