Figure 2.
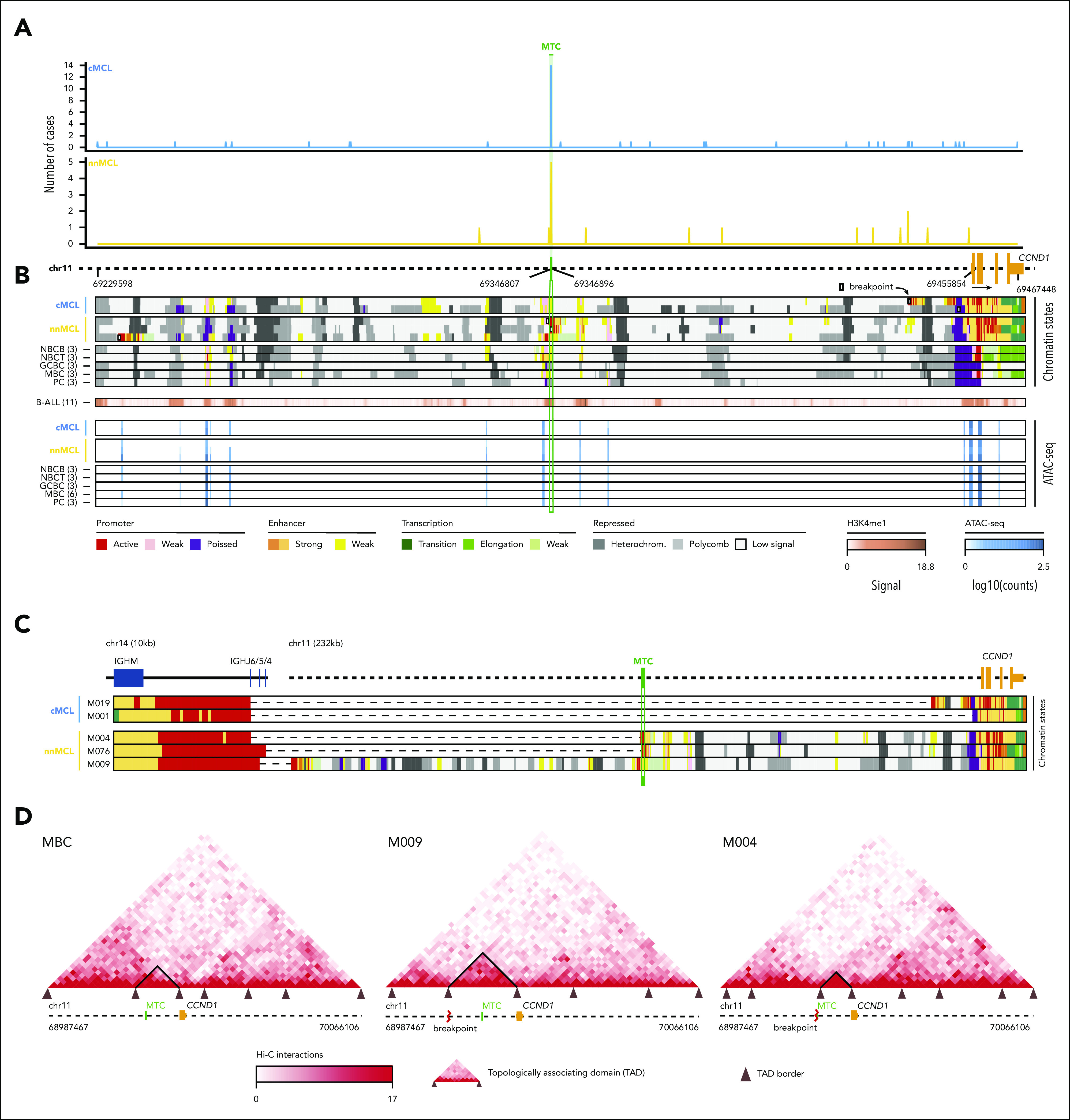
Integrative analysis of chr11 breaks of the IG/CCND1 translocation. (A) Distribution of breakpoints observed in chr11 in cMCL (top) and nnMCL (bottom). The number of cases with breakpoints in close proximity is summarized using a sliding window of 89 bp (MTC length) starting from the MTC region (shown in green). (B) Representation of the reference epigenomes of 5 MCL cases and 15 normal B cell samples spanning the B-cell maturation program. Numbers in brackets indicate the number of samples considered to build the consensus chromatin map of each cell type. (top) The breakpoints of each case are highlighted by a white square. (middle) Signal of H3K4me1 in B-cell acute lymphoblastic leukemia (B-ALL). (bottom) Chromatin accessibility (ATAC-seq peaks) showing the presence of an open chromatin region near the MTC region. B cells are grouped, and a consensus is depicted. (C) Reconstruction of the chromatin states of the IG/CCND1 translocated allele. Dashed lines indicate the junction of both chromosomes to maintain the breakpoints relative to the MTC and CCND1. (D) Hi-C contact matrices for memory B cells (MBC) and 2 nnMCL cases. The TAD associated with CCND1 and modulated upon the IG/CCND1 translocation is highlighted. GCBC, germinal center B cell; NBCB, naive B cell from peripheral blood; NBCT, naive B cell from tonsil; PC, plasma cell.
