Fig. 3.
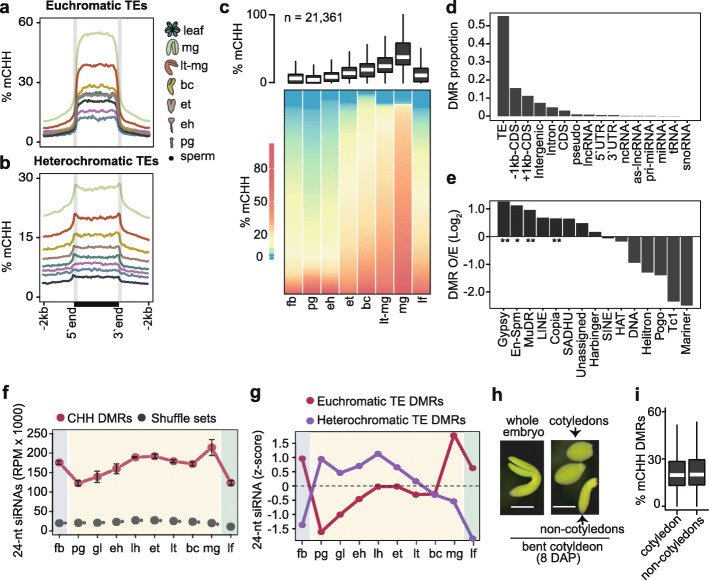
Embryonic methylome dynamics. a, b Metaplots of average weighted CHH methylation percentages across euchromatic (a) and heterochromatic (b) TEs in sperm, embryos, and leaves (key). pg, preglobular; eh, early heart; et, early torpedo; bc, bent cotyledon; lt-mg, late torpedo-to-early mature green; mg, mature green. c Boxplot and heat map illustrating the percentage of CHH methylation across DMRs in flowers, embryos, and leaves. The number of DMRs identified are indicated, and stages are labeled as in a and include floral buds (fb). Thick horizontal bars in the boxplot indicate medians, and the top and bottom edges of the box indicate the 75th and 25th percentiles, respectively. DMRs in heatmap were sorted by average methylation levels per column. d Proportion of genomic features overlapping DMRs. e Bar chart showing the enrichment of TE families observed overlapping DMRs relative to those expected based on their genomic proportions (O/E; log2). P values < 0.01 and < 0.05 based on Fisher’s exact test are represented by ** and *. f Line chart of 24-nt siRNA levels overlapping CHH DMRs (red) compared to randomly selected genomic regions with equal sizes and quantities (gray) in floral buds (fb), embryos, and leaves (lf). Embryo stages are labeled as in Fig. 1a. g Relative 24-nt siRNA levels (z-scores) on DMRs mapping to euchromatic (red) and heterochromatic (purple) TEs across development. Embryo stages are labeled as in Fig. 1a. h Representative image of a bent cotyledon-staged embryo 8 days after pollination (DAP) and dissected into cotyledon and non-cotyledon tissues for methylC-seq. Scale bar represents 0.25 mm. i Boxplot of CHH methylation percentages of DMRs defined in c for cotyledon and non-cotyledon tissues. Boxplots are as described in b. See also Additional file 2: Figure S3