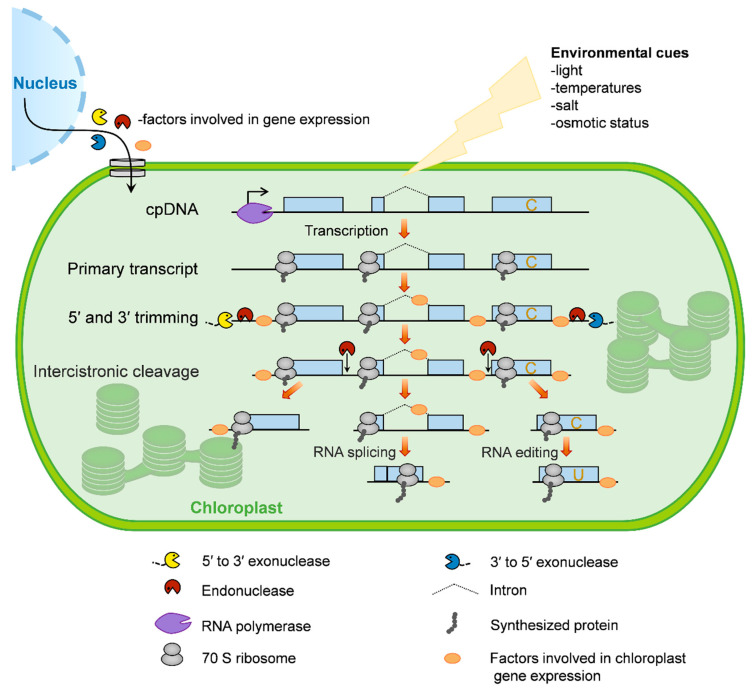
Figure 1.
Overview of chloroplast gene expression. In plants, most chloroplast genes are organized as operons and are controlled by single promoters (bent arrow). These genes are transcribed by two distinct types of RNA polymerase: Nucleus-encoded RNA polymerase (NEP) and plastid-encoded RNA polymerase (PEP). The resulting primary transcripts require several processing steps to form mature mRNA, including 5′ and 3′ trimming, intercistronic cleavage, RNA splicing, and RNA editing. In order for these events to take place, numerous nucleus-encoded proteins are translated in the cytosol and imported into the chloroplast, where they control and/or regulate chloroplast gene expression. Chloroplast gene translation is conducted by bacterial-type 70S ribosomes, which occurs cotranscriptionally. Since the mRNA turnover rate within chloroplasts is slow, most ribosomes function in posttranscriptional steps. Moreover, chloroplast gene expression is involved in responses to environmental cues.