Figure 1.
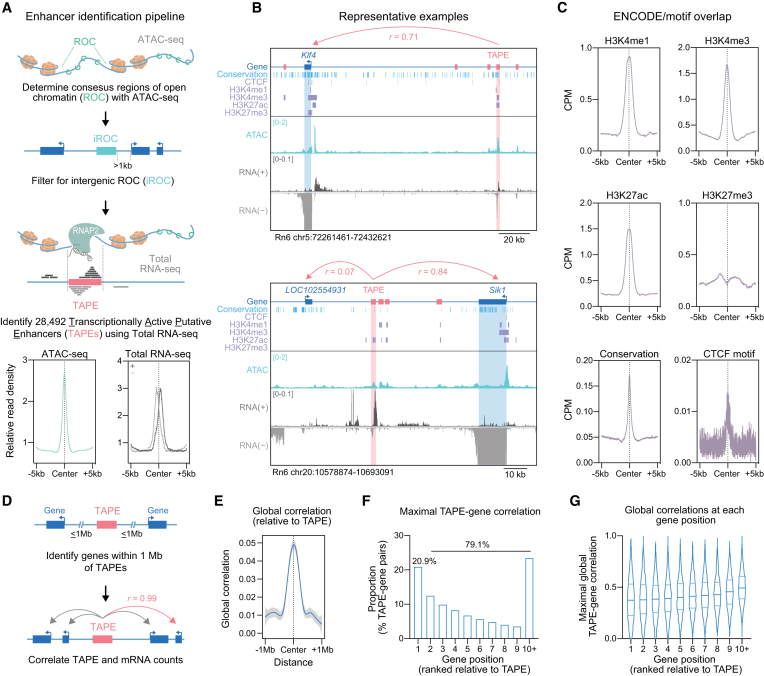
Genome-wide characterization of enhancers and eRNAs. (A) Analysis pipeline for localization and quantification of transcriptionally active putative enhancers (TAPEs). ATAC-seq datasets were generated using cultured cortical, hippocampal, and striatal rat neurons and used to identify regions of open chromatin (ROCs). ROCs were filtered to capture intergenic regions at least 1kb from annotated genes (iROCs). Total RNA-seq data from the same culture systems was used to identify 28 492 bidirectionally transcribed intergenic ROCs and termed TAPEs. TAPEs are characterized by an enrichment of ATAC-seq and bidirectional RNA-seq reads at TAPE centers. (B) Genome browser tracks showing ATAC-seq signal and total RNA expression at two example regions (Klf4 and Sik1), relative to tracks marking conserved DNA elements (phastConsElements20way), CTCF motifs, and enhancer-linked histone modifications. (C) TAPEs exhibit higher densities of H3K4me1, H3K4me3, H3K27ac, sequence conservation and CTCF motifs, and decreased H3K27me3 compared to surrounding regions. Chromatin immunoprecipitation datasets were obtained from the mouse forebrain at postnatal day zero (ENCODE project) and lifted over to the rat Rn6 genome assembly. (D) TAPE-gene pairs were determined by correlations of eRNA and mRNA levels at genes within a 1 Mb distance cutoff. (E) On average, TAPEs and closer genes show higher correlation values. (F) Only 20.9% of TAPEs have their maximal gene correlation at the closest gene, while the remaining 79.1% show higher correlations to genes at more distal positions. (G) For pairs with highest global TAPE–gene correlation, correlation strength does not decrease with gene distance from TAPE.