Figure 2.
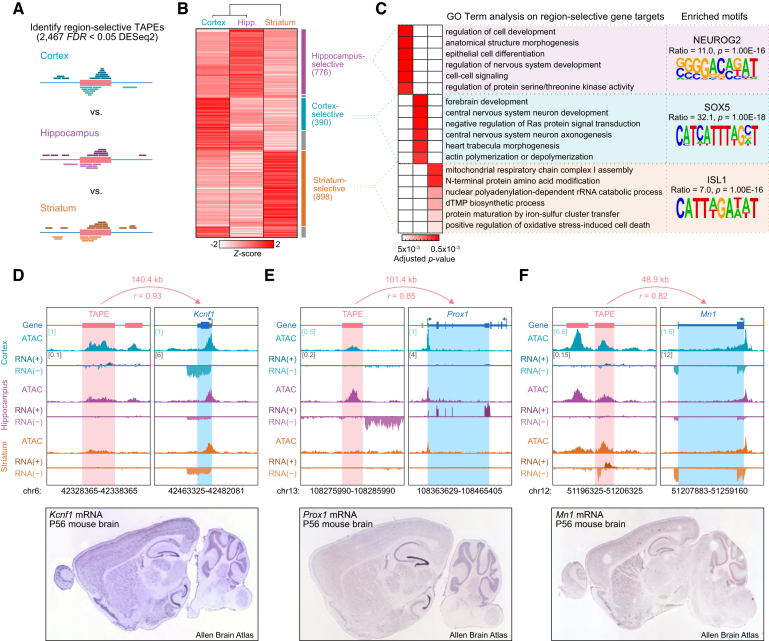
Identification of brain region-selective enhancers and eRNAs in primary neuronal culture systems. (A) Illustration of DESeq2-based identification of enhancers selective for cortex, hippocampus, and striatum. (B) Heatmap indicating transcription levels at region-selective TAPEs revealed 776, 390 and 898 TAPEs selective for hippocampus, cortex, and striatum, respectively. (C) Heatmap showing adjusted P-values of the six most enriched Gene Ontology Term groups for genes corresponding to region-selective TAPEs (left). HOMER analysis of transcription factor binding motifs shows an enrichment of characteristic transcription factors within region-selective TAPEs (representative examples on right, complete list in Supplementary Data Table S3). (D–F) Genome browser tracks showing ATAC-seq and total RNA-seq (reads mapped to + and – strand) signal at example loci separated by the three brain regions of interest (top, square brackets indicate y-axis max CPM value for all ATAC-seq and RNA-seq tracks at the respective loci). Region-selective TAPEs are represented by Kcnf1 for cortex, Prox1 for hippocampus, and Mn1 for striatum. Distance between TAPE and predicted target gene and global Pearson's correlation is indicated above each representative TAPE–gene pair, Rn6 chromosome coordinates are indicated below. Region-selective expression of the respective target genes is also evident in in situ hybridization images from the adult mouse brain (bottom, Image credit: Allen Brain Atlas, Allen Institute).