Figure 4.
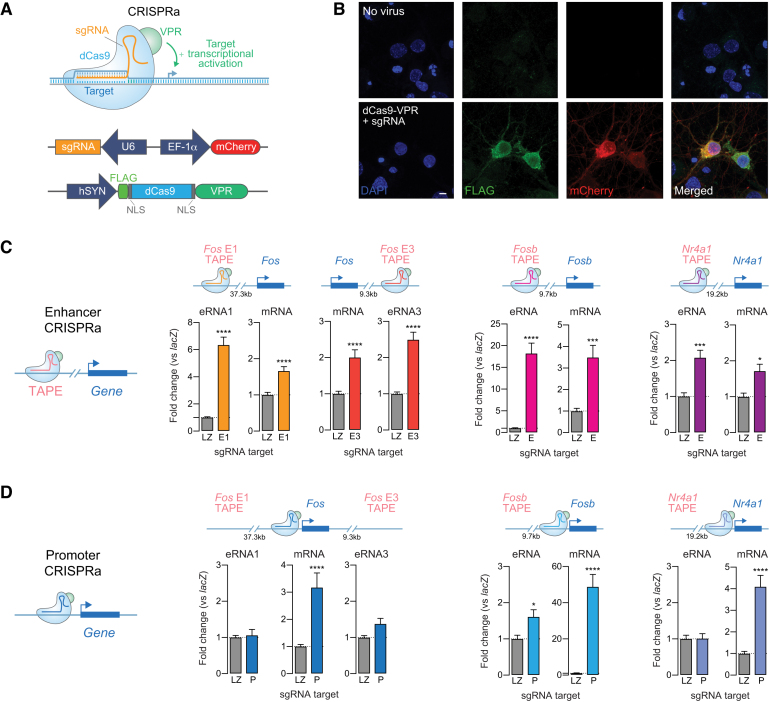
Transcriptional activation at enhancers is sufficient to induce linked genes. (A) Illustration of CRISPR activation (CRISPRa) strategy for site-specific targeting of the transcriptional activator VPR. (B) Immunocytochemistry on DIV 11 cortical neurons. Top, no virus control. Bottom, neurons co-transduced with lentiviruses expressing dCas9-VPR (marked by FLAG) and a custom sgRNA (mCherry reporter). Scale bar = 5 mm. (C) CRISPRa targeting to distal enhancers near Fos, Fosb, and Nr4a1 loci activates eRNA and mRNA from linked genes. Gene expression differences were measured with RT-qPCR (Fos, n = 18; Fosb and Nr4a1 n = 9 per group; two-tailed Mann-Whitney test for all comparisons as compared to non-targeting lacZ sgRNA control). (D) CRISPRa at Fos, Fosb, and Nr4a1 promoters increases mRNA but does not consistently increase eRNA (Fos, n = 18; Fosb and Nr4a1 n = 9 per group; two-tailed Mann–Whitney test for all comparisons as compared to non-targeting lacZ sgRNA control). Data expressed as mean ± s.e.m. Multiple comparisons, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.