Figure 5.
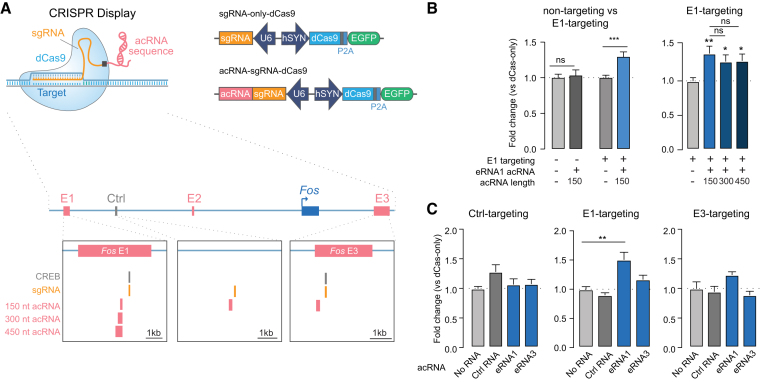
Fos eRNA1 is sufficient for Fos mRNA expression. (A) Illustration of Display plasmids (top), and CREB binding motifs, sgRNA target sites and eRNA regions chosen for CRISPR-Display constructs (acRNA1–3) (bottom). (B) RT-qPCR analysis reveals that while the lacZ-targeting 150 nt eRNA1 acRNA does not affect Fos mRNA (n = 9 per group, two-tailed Mann–Whitney test, U = 36, P = 0.7304), targeting eRNA1 to Fos enhancer-1 results in increased Fos mRNA expression (n = 21 per group, two-tailed Mann–Whitney test, U = 77, P = 0.0002, graph contains data from 12 replicates (from four experiments) shown in the right graph and none additional replicates (from 2 experiments)). Constructs with increasing acRNA lengths (150, 300 and 450 nt) did not result in stronger Fos mRNA induction (one-way ANOVA F(3,44) = 3.791, P = 0.0167, Tukey's post hoc test for multiple comparison). (C) Tethering 150 nt acRNAs based on eRNA1, eRNA3, and a control RNA revealed that only eRNA1 tethering to its own enhancer induced Fos mRNA expression (one-way ANOVA for control-targeting F(3,19) = 3.191, P = 0.0472, and Kurskal–Wallis test for E1-targeting F(3,44) = 23.27, P < 0.0001, E3 targeting F(3,40) = 5.183, P = 0.1589). Data expressed as mean ± s.e.m. Multiple comparisons, *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.