Fig. 5.
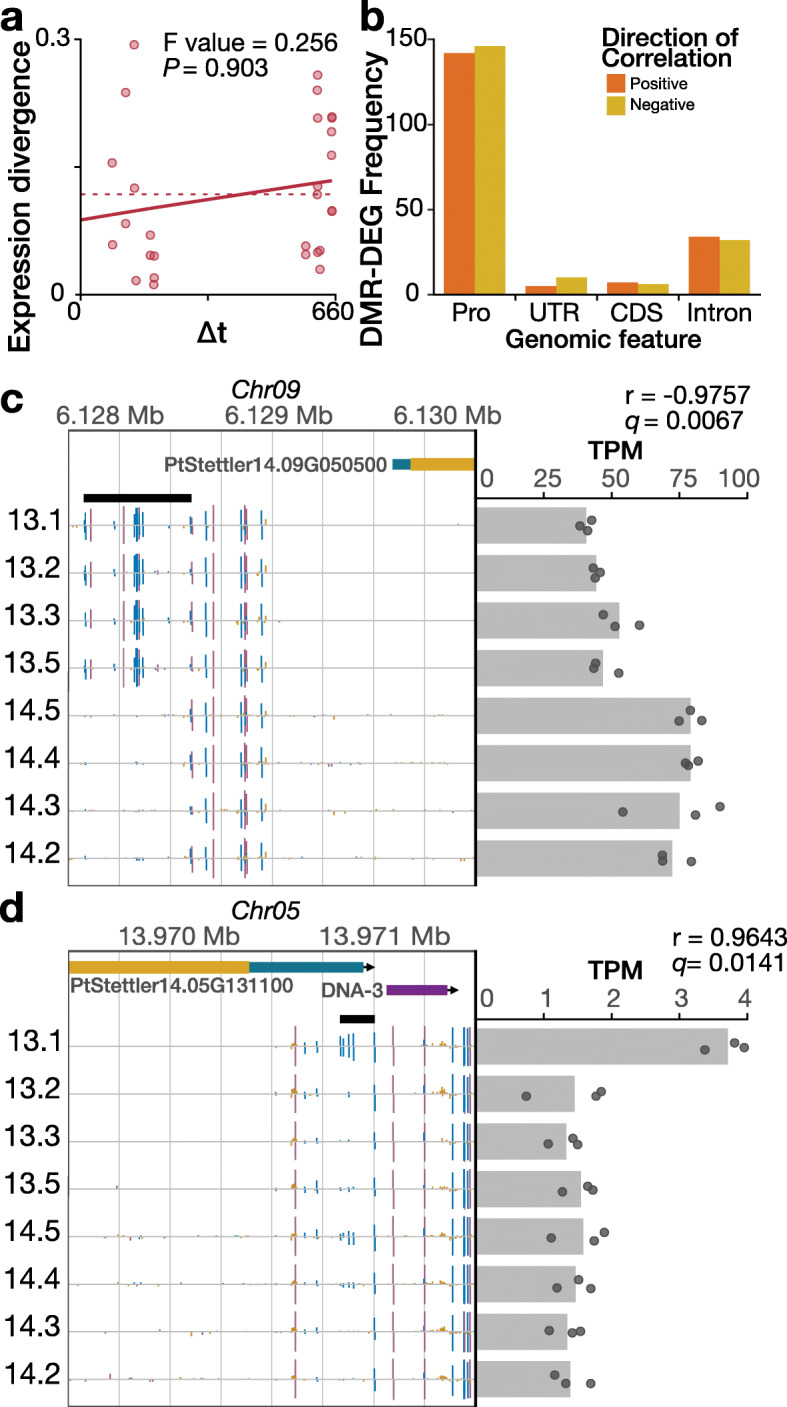
Gene expression is largely independent from divergence age and nearby cytosine methylation except in a few examples. a Gene expression divergence is not significantly associated with divergence age. The dots show the individual observed divergences, whereas the line represents the fit of the data to the model. b Distribution of positive and negative correlations for differentially expressed genes and overlapping/nearby DMRs. Positive correlation occurs when the higher methylation level is associated with higher gene expression among the samples. c A significantly negatively correlated, tree-specific DMR and DEG where the DMR occurs in the promoter region of the gene (Pearson’s correlation test, two-sided, N = 8, adjusted P value = 0.0067). The higher methylation levels in the DMR for tree 13 branches are associated with lower gene expression. d A significantly positively correlated, single gain DMR and DEG where the DMR occurs in the 5′ untranslated region of the gene (Pearson’s correlation test, two-sided, N = 8, adjusted P = 0.0141). The higher methylation level in the DMR for branch 13.1 is associated with greater gene expression. For c and d, gene expression, as transcripts per million (TPM), is represented as points for the individual replicates and as bar for mean among replicates. In the genome browser view, gene models and transposable elements are shown at the top and methylome tracks are below. Vertical bars indicate methylation at the position, where height corresponds to level and color is context, red for CG, blue for CHG, and yellow for CHH. DMR is indicated by thick black horizontal line
