Fig. 2.
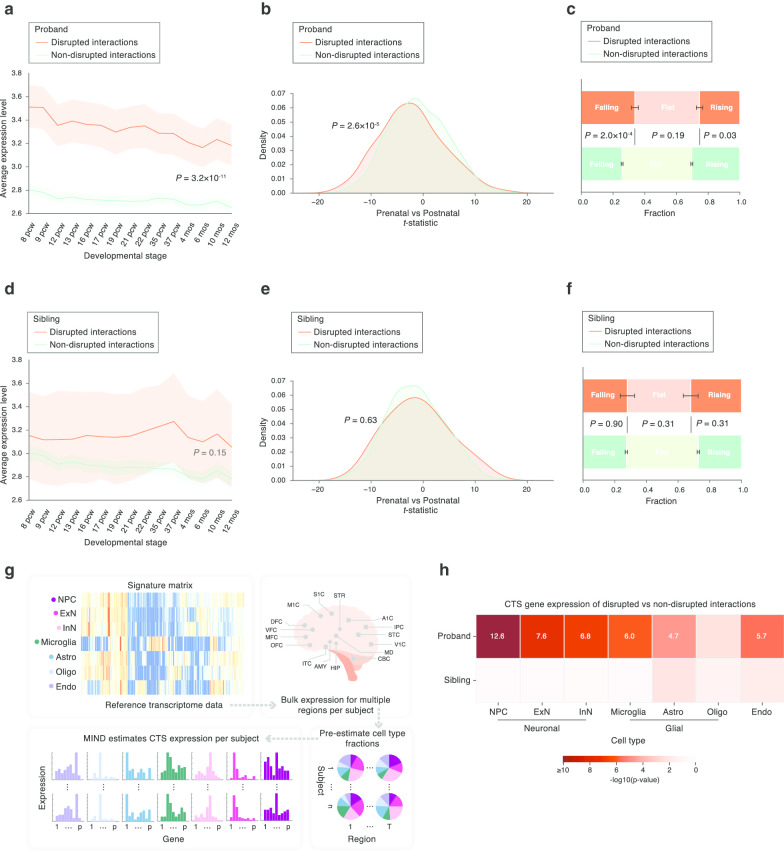
Transcriptome analyses of disrupted interactions implicate specific times and cell types in ASD. a–c Disrupted interaction genes identified in ASD probands a are highly expressed in brain, b exhibit prenatal expression bias, and c tend to have a falling expression trajectory. d–f, Disrupted interaction genes found in unaffected siblings do not present the above features. g MIND algorithm to estimate subject-level CTS expression from BrainSpan bulk RNA-seq data. h Disrupted interaction genes are most highly expressed in neuronal cell types. The heatmap shows the negative log10(P value) for comparing expression levels of disrupted versus non-disrupted interaction genes in seven cell types. Cell types that are significant after Bonferroni correction are noted with their negative log10(P value) written in white. pcw post-conception weeks, mos months, CTS cell-type-specific, NPC neuronal progenitor cell, ExN excitatory neuron, InN inhibitory neuron, Astro astrocyte, Oligo oligodendrocyte, Endo endothelial