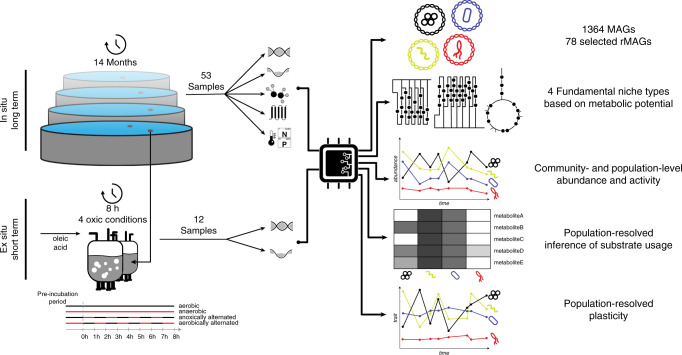
Fig. 1. Overview of the study design.
Samples are derived from in situ sampling of an anoxic tank of a municipal biological wastewater treatment plant. Metagenomic (MG), metatranscriptomic (MT), metaproteomic (MP), and meta-metabolomic (MM) data is generated. Physicochemical data is also collected. Additionally, MG and MT data is generated for ex situ experiments using biomass from the same system and fed with oleic acid under different oxic conditions to evaluate short-term responses to pulse disturbance. The time-series meta-omics data is integrated to define metagenome-assembled genomes (MAGs) over all time points. Representative MAGs (rMAGs) across time are selected for further analysis. The rMAGs’ functional potential is used to infer the fundamental niches. Abundance and activity data are derived from the functional omics and substrate usage is inferred per population. The variability of gene expression is used to assess the phenotypic plasticity of the individual populations.