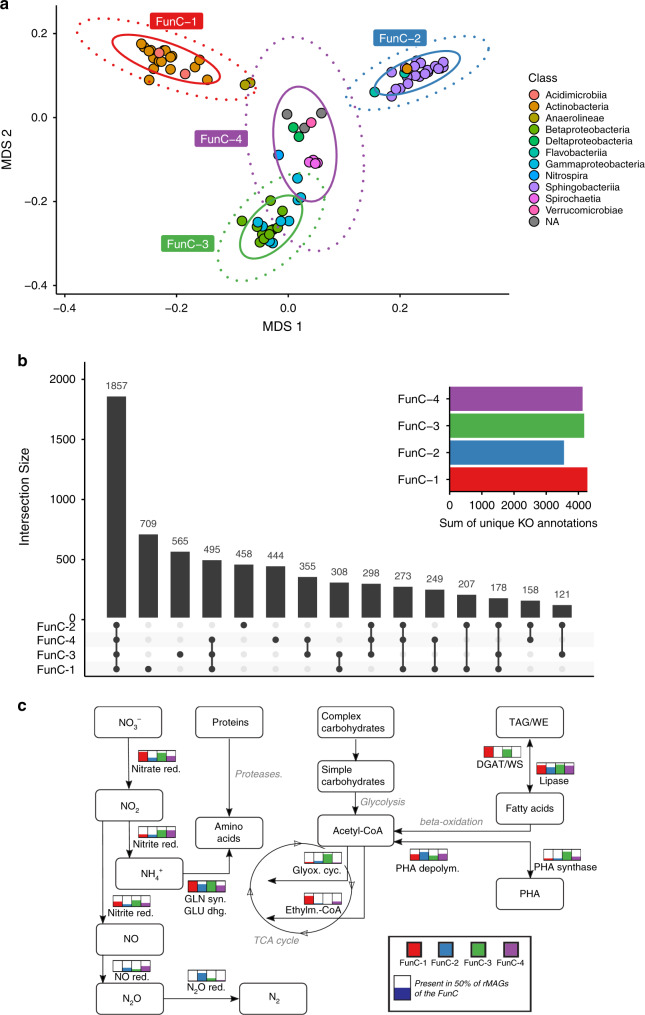
Fig. 2. Fundamental niche types.
a Multidimensional scaling (MDS) of Jaccard distances for the functional repertoire (presence of KEGG ortholog groups [KOs]) for each rMAG. Ellipses containing 95% (inner) or 99% (outer) of cluster-assigned data points are shown resulting in four distinct functional clusters (FunCs). Colors indicate the class-level taxonomy of the rMAGs. b Numbers of shared and unique KO assignments between the FunCs. Colored bars show the total number of nonredundant KO assignments within the individual FunCs. Overlaps between different sets of FunCs and their unique KOs are represented by the central black bars with the points below defining the members of the respective sets. c Presence of key functions within the four FunCs. Bars next to metabolic conversions show the proportion of rMAGs encoding characteristic enzymes for the respective reaction or pathway adjusted for mean rMAG completeness. Pathways ubiquitously present across rMAGs are shown in gray color. Source data are provided as a Source Data file. red. reductase, GLN syn. glutamine synthetase, GLU dhg. glutmatate dehydrogenase, glyox. cyc. glyoxylate cycle, ethylm.-CoA ethylmalonyl-CoA pathway, PHA depolym. PHA depolymerase.