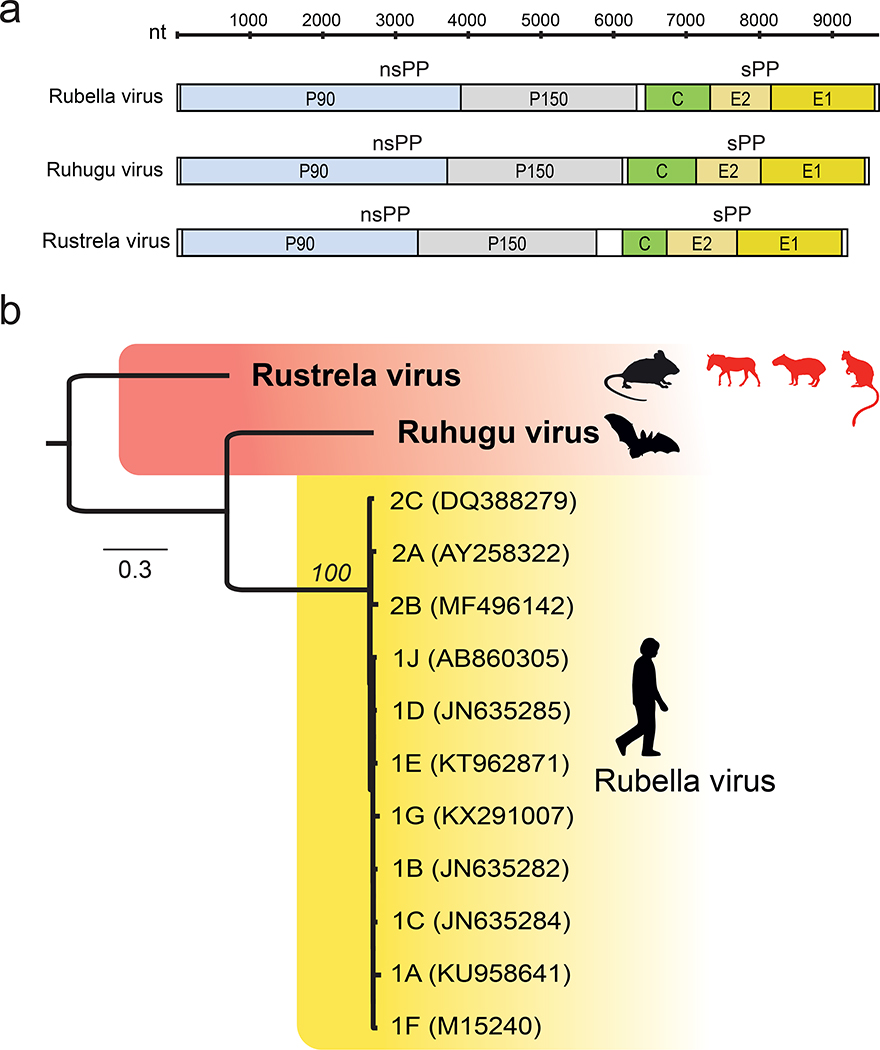
Fig. 3 |.
Evolutionary relationships among viruses. a) Comparative genome architecture of RuV, RuhV, and RusV, showing five open reading frames (colored), two untranslated regions at the 5′ and 3′ termini (white), and an intergenic region (white) between the ORFs encoding the non-structural (nsPP) and structural (sPP) polyproteins. b) Maximum likelihood phylogenetic tree of rustrela virus, ruhugu virus, and rubella virus genotypes 1A–1J and 2A–2C. Black silhouettes represent natural hosts of each virus, and red silhouettes represent spillover hosts in the case of RusV. Numbers beside nodes indicate bootstrap values (%; only values for major branches are shown); the scale bar indicates amino acid substitutions per site.