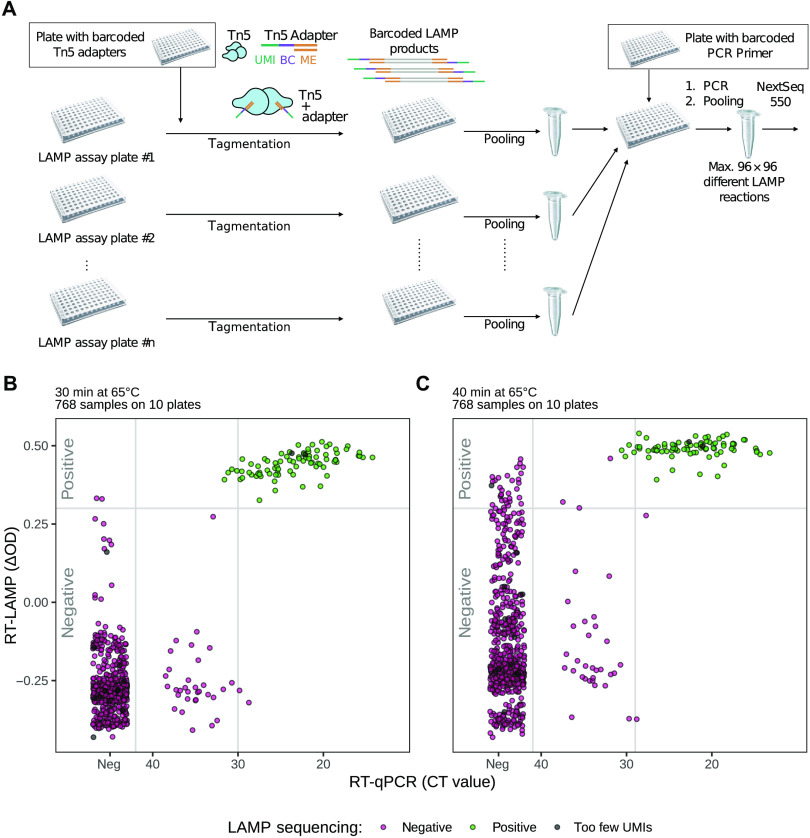
Fig. 4. Multiplexed sequencing of RT-LAMP reaction products (LAMP-sequencing).
(A) Workflow for LAMP-sequencing is shown. A plate of 96 barcoded (BC) adapters with unique molecular identifiers (UMIs) and mosaic ends (ME) was used as a seed plate for Tn5 tagmentation of all RT-LAMP reaction products. After tagmentation, each plate was pooled individually, followed by removal of excess adapters using size selection. Each pool of tagmentation products was then amplified using primers with plate-specific barcodes, and the PCR products were analyzed by Illumina sequencing. (B) Comparison of the outcome of the three assays: LAMP-sequencing (purple, negative; green, positive; gray, too few UMIs), RT-LAMP (after 30-min incubation, y axis), and RT-qPCR (x axis). Each dot represents one sample. If a substantial number of the sequencing reads contained SARS-CoV-2 RNA, the sample was called positive (green), if not, then it was called negative (purple). For some samples (gray), no LAMP-sequencing call could be made due to too few UMIs. (See also Table 2). (C) Although the RT-LAMP assay was scored after a 30-min incubation at 65°C (left), LAMP-sequencing was performed only after the samples had been incubated for another 10 min (15 min for one plate). This panel shows the RT-LAMP assay outcome (y axis) scored after the full incubation time, whereas the RT-qPCR CT values (x axis) and LAMP-sequencing results are the same as in (B).