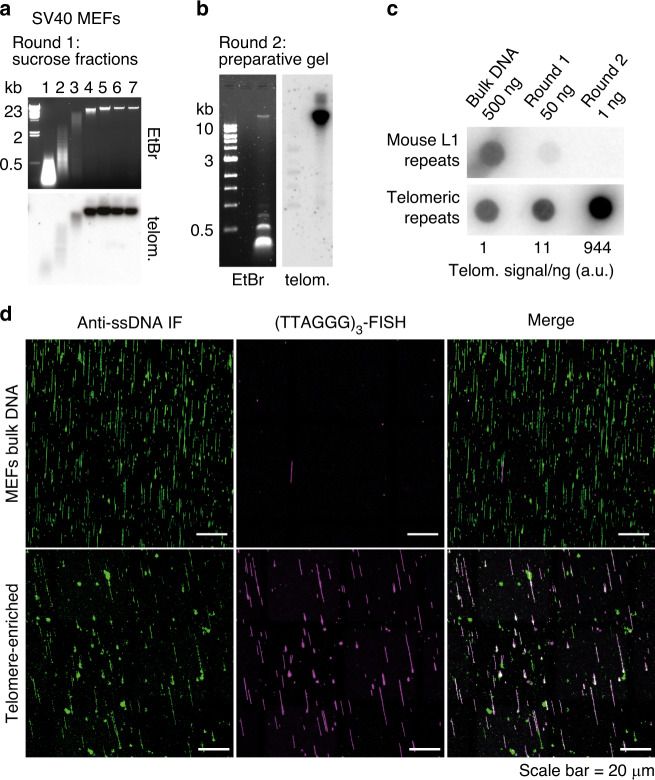
Fig. 1. A two-step procedure for the purification of mammalian telomeres.
a Top: agarose gel showing the separation of the large telomeric repeat fragments from the bulk genomic DNA in a sucrose gradient. Genomic DNA (∼2.5 mg) from SV40LT-immortalized MEFs was digested with HinfI and MspI. The digested DNA was separated by centrifugation on a sucrose gradient. Seven fractions were collected and an aliquot (∼1/500) of each fraction was loaded on an agarose gel. Bottom: the gel was blotted onto a membrane and hybridized with a TTAGGG repeats probe to verify that telomeric repeats remained in the high molecular weight (HMW) fractions. Source data are provided as a Source Data File. b Left: agarose gel showing the separation of the large telomeric repeat fragments from the remaining non-telomeric DNA, in the second purification round. The HMW DNA, contained in the last four fractions of the sucrose gradient described in (a), was recovered and digested with RsaI, AluI, MboI, HinfI, MspI, HphI, and MnlI. The digested DNA was separated on a preparative agarose gel and the DNA migrating in the area above 5 kb was extracted from the gel. The image shows an aliquot (∼1/100) of the digested DNA, separated on an agarose gel. Right: the gel was blotted onto a membrane and hybridized with a TTAGGG repeats probe to verify that telomeric repeats remained in the HMW area. Source data are provided as a Source Data File. c Dot blot analysis showing the enrichment of telomeric repeats. The indicated amounts of DNA from each enrichment step were spotted on a membrane and hybridized either with a probe recognizing the long interspersed L1 repeats or TTAGGG repeats. The amount of TTAGGG repeat signal/ng was quantified and reported relative to the signal/ng value in the initial, non-enriched DNA. Source data are provided as a Source Data File. d Single-molecule analysis showing the enrichment of the telomeric repeats. The DNA was combed onto silanized coverslips, denatured in situ and labeled sequentially with an antibody against single-stranded DNA and a Cy3-labeled (TTAGGG)3 PNA probe.