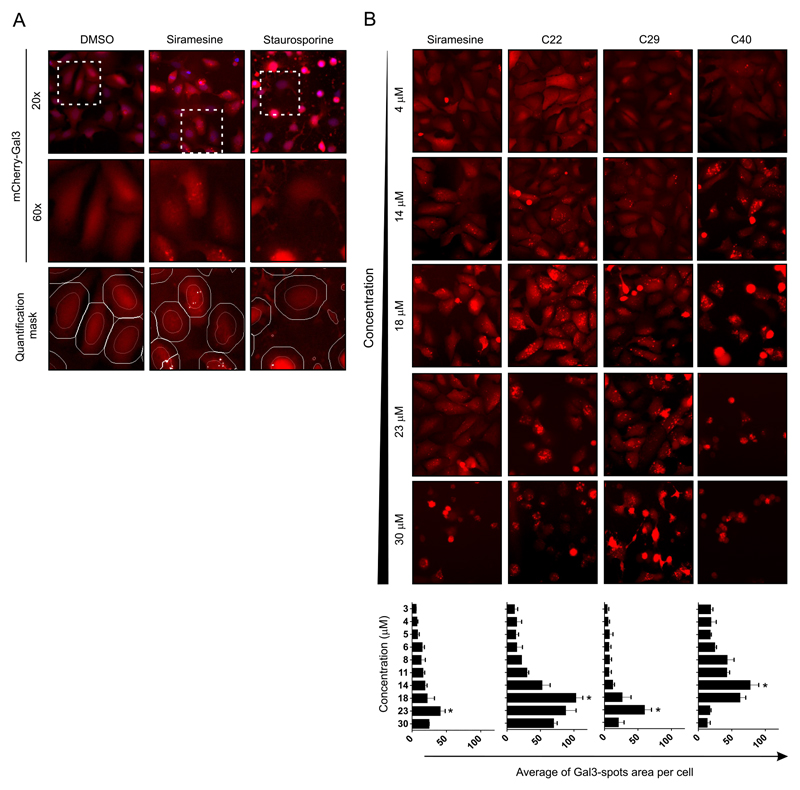
Figure 4. Quantification of Gal-3 re-localization to damages lysosomes by automatic microscopy.
A. LMP can be detected with U2OS-mCherry-Gal3 reporter cells using an automatic microscope. Cells were treated for 6 h, at concentration between 3 and 30 μM. Cells were stained with Hoechst (nuclear mask) and Calcein-AM (cell mask) and subjected to quantitative image analysis using the CellInsight™ CX5 High Content Screening (HCS) Platform. The quantification algorithm was optimized to detect the mCherry-Gal3 puncta (damaged lysosomes) within an artificially defined circular area centered on each nucleus. mCHerry-Gal3 puncta were quantified as the Mean_spot_area_per_cells in channel 2. Top row. CellInsight representative images for the vehicle (DMSO), positive (siramesine) and negative (staurosporine), overlays of the nuclei (Channel 1, blue) and mCherry-Gal3 (Channel 2, red). Middle row. Magnification of the mCherry-Gal3 channel for the indicated area. Bottom row. Magnification of the mCherry-Gal3 channel in overlay with the quantification marks for “nucleus” (orange) “cell mask” (white) and “detected spots” (white spots) as determined by the established automatic cell image quantification algorithm.
B. Automatic microscopy images of mCherry-Gal3 channel for four selected compounds at five selected concentrations. The assay was performed as indicated in panel A. There is an optimal concertation for the detection of Gal-3 puncta above which cell rounded-up. Bottom panel. Graphs showing the average “Gal-3 spot area per cells” across the 10 concentrations for the four selected compounds. Error bars are SDs for means of 3 wells. For each well, 10 images with a total of > 500 cells/sample.