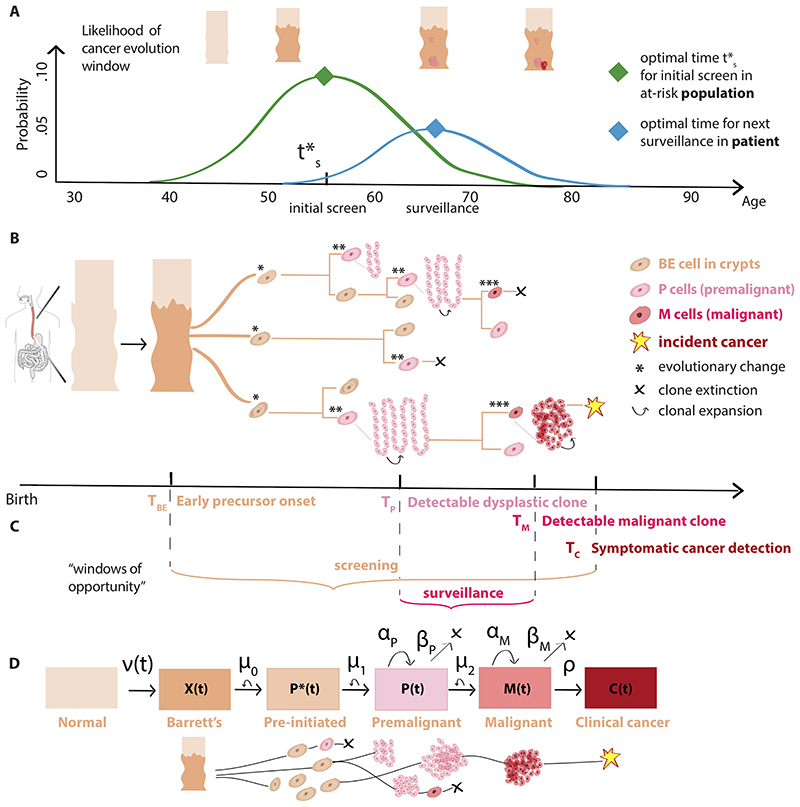
Figure 1. Optimizing screening and surveillance using branching process models of cancer evolution.
(A) After an initial screen to detect premalignant changes at time t’s, adaptive recommendations for patient-specific survei llance can account for the heterogeneity in a screened population. (B) Somatic evolution of stem cell lineages since birth leads to stochastic trajectories (example asymmetric divisions shown on tree nodes) that accumulate mutations and may be selected for advantageous phenotypes. Within an at-risk patient’s esophagus, normal squamous epithelium may transform to a columnar, crypt-filled Barrett’s esophagus (BE) segment. Stochastic initiation of pre-malignant P cells can occur (e.g., after ‘two-hit’ gene inactivation of TP53 in a cancer-promoting microenvironment), which can then undergo clonal expansion. Malignant cells M that are transformed, in turn, may undergo clonal expansions, be detected as a clinical EAC, or go extinct. (C) “Windows of opportunity” for optimal screening and surveillance can be formally defined using random variables T for each stage denoted by subscript (see Supplementary Methods 3.1 for details) to be used in objective functions in (A), example shown specific for BE-EAC pathway. (D) The multi-stage clonal expansion for EAC (MSCE-EAC) model. Model parameters: Transformation rate to X BE cells, ν(t); Pre-initiation Poisson rate, μ0 (P’ progeny); Initiation Poisson rate μ1 (P progeny); Premalignant birth-death-mutation process cell division rate, αp, cell death-or-differentiation rate, βp, malignant transformation rate, μ2; Malignant birth-death-detection process cell division rate, αM, cell death-or-differentiation rate, βM, size-based clinical detection rate, ρ (see Supplementary Methods 3.3 for model fits to cancer incidence).