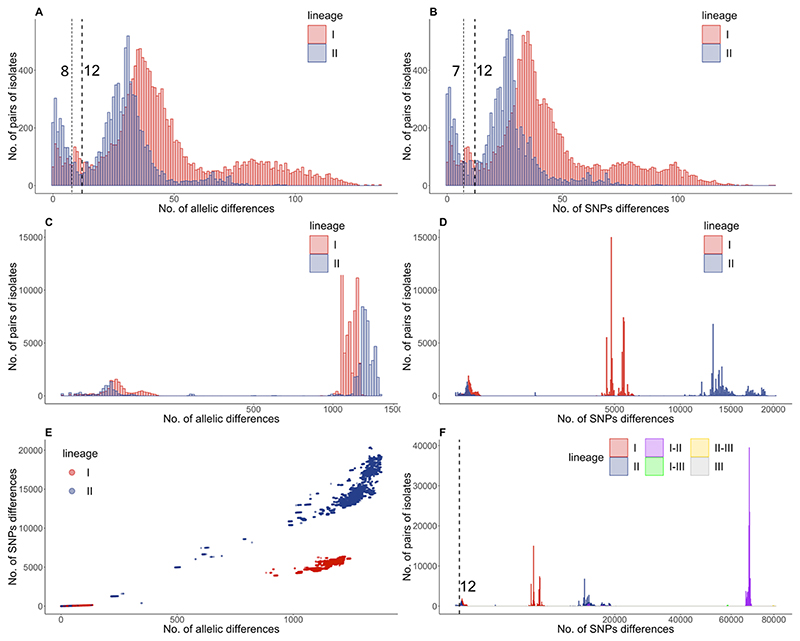
Figure 2. Lineage specific evolution of L. monocytogenes.
Pairwise cgMLST (A, B) and SNP (C, D, F) differences were analysed for 708 isolates form Switzerland, Germany and the Netherlands. Pairwise analyses are plotted separately for diversity within sub-lineage (A-B; up to 150 differences), between sub-lineage and within lineage (C-D; up to 1,500 cgMLST differences and 20,000 SNPs) and between lineages (F). The correlation between SNP and cgMLST differences are plotted in panel E. The pairwise SNP analysis considering the whole population (F) includes data for within lineage I differences (red), within lineage II differences (blue), within lineage III differences (grey) and the differences between lineage I and II (purple), between lineage I and III (green), and between lineage II and III (yellow).