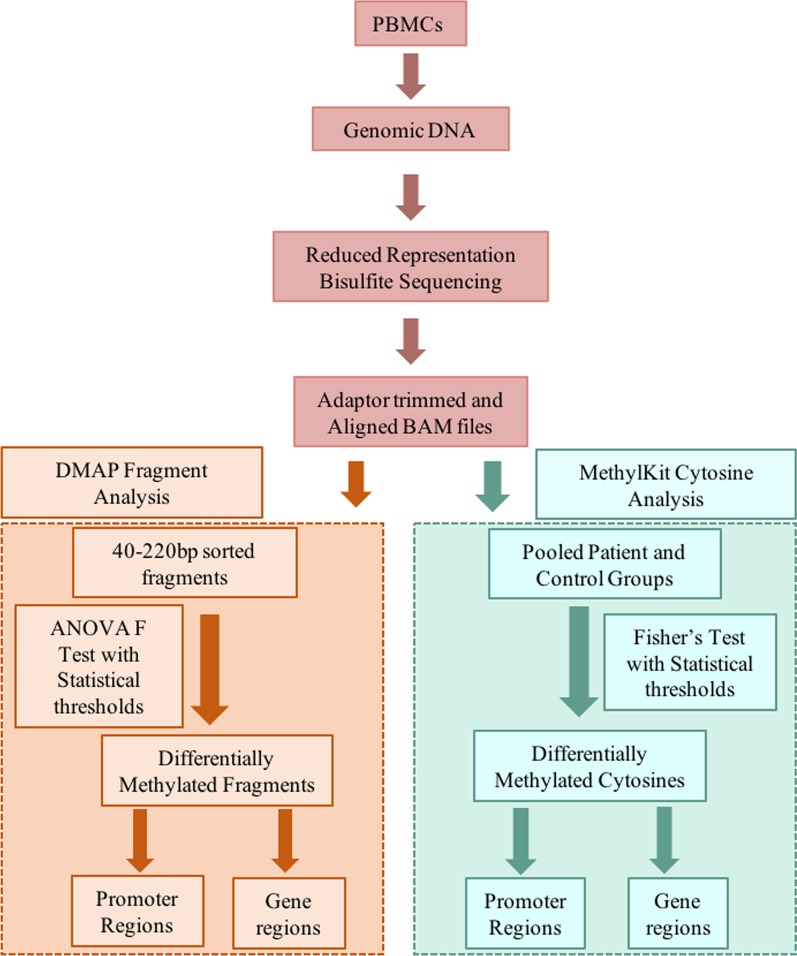
Fig. 1.
The study and analysis workflow for ME/CFS patients and age-matched healthy control methylome: DNA was isolated from the Peripheral blood mononuclear cells (PBMCs) of 10 ME/CFS patients and 10 age/sex matched healthy controls. DNA was processed to produce reduced representation bisulphite sequencing libraries. This DNA was then sequenced and the data aligned and trimmed using Bismark. The aligned RRBS sequence data were then analyzed using two parallel analyses pipelines: DMAP and MethylKit. DMAP utilised an ANOVA F test comparing patients and controls (requiring at least seven from each group to be present in each comparison between fragments). Associated promoter and gene regions were then identified. Methylkit analysis pooled the patient and control groups before comparison using a Fisher’s test to identify differentially methylated cytosines (requiring at least seven from each group to be present in the comparison between cytosines). Promoter and gene regions were then isolated and investigated