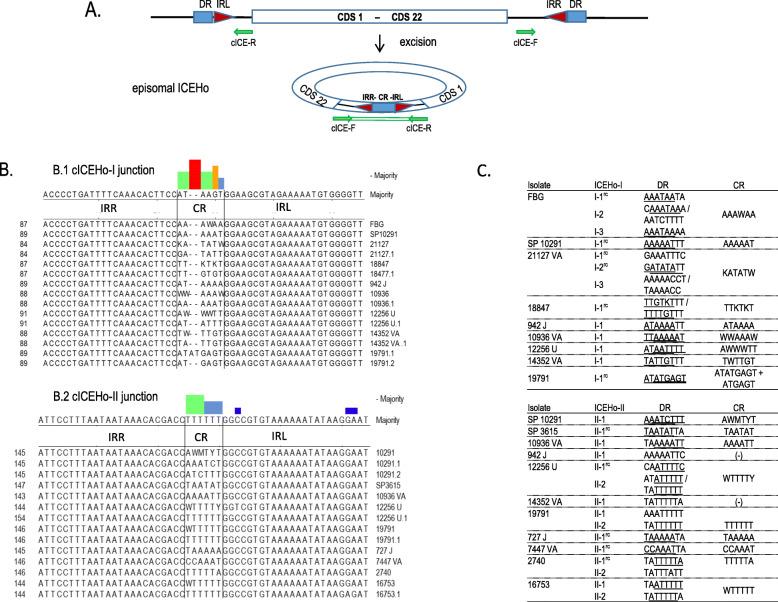
Fig. 4.
Chromosomal and episomal ICEHo-I and –II. a Schematic representation of the excision of a chromosomal integrated ICEHo element to build the circularized episomal cICE-I or -II. ICEHo-flanking left- and right-positioned inverted repeats (IRL and IRR) are represented by red triangles, and the direct repeats (DRs) are represented by blue squares. After excision of ICEHo, direct repeats are fused to the coupling region (CR) in the episomal cICEHo. For the detection of episomal cICEHo-I and -II, specific primer pairs (represented as green arrows) were used that do not lead to an amplification when targeting the chromosomal ICEHo-I and –II in outwards-facing position, and which result in 0.2 (cICE-I) or 0.3 kb (cICE-II) PCR products when targeting the circularized ICEHo (see Methods). b Multiple sequence alignments of IRR-IRL junction sequences with the central CR of cICE-I (B.1) and cICE-II (B.2), determined by Sanger sequencing of cICE PCR products. Ambiguity characters in the coupling region (M = A or C; K = G or T; W = A or T; Y = C or T) indicate the presence of minor sequence variants as detected by Mixed Sequence Reader [27]. The respective sequences are listed as strain.1 for the major sequence and strain.2 for the minor sequence. Coloured bar charts above the consensus sequence, created with the Lasergene software package (DNAStar, Madison, WI), indicate the presence of non-consensus characters (disagreements) in the corresponding column, ranging from red (high frequency of non-consensus characters or gaps) to blue (low frequency of non-consensus characters). c List of direct repeats (DR) of the chromosomal ICEHo-I and cICEHo-II copies (I-1 to I-3; II-1 to II-2) detected in the nanopore-sequenced M. hominis strains in comparison to the episomal coupling regions (CR). Identical (or comparable) sequences are underlined; if two different regions of one DR are underlined, both regions used for recombination explain the mismatch in CR. IUAPC ambiguity characters in the genomic DR sequences were introduced during the removal of circular contig overlaps (see Methods). I-xrc = the respective ICEHo-element is inserted in a reverse-complementary fashion with respect to the genome assembly, requiring evaluation of the reverse complementary DR