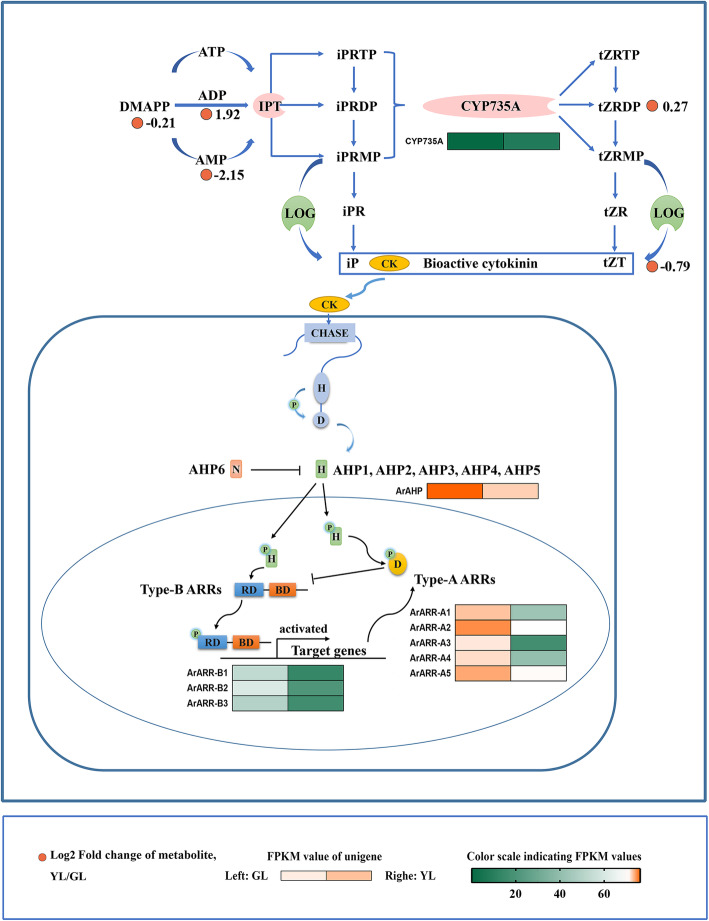
Fig. 4.
Biosynthetic and signal transduction pathways of cytokinins. DMAPP, dimethylallyl diphosphate; ATP, adenosine triphosphate; ADP, adenosine diphosphate; AMP, adenosine monophosphate; IPT, isopentenyl transferase; iPRTP, isopentenyl adenosine-5′-triphosphate; iPRDP, isopentenyl adenosine-5′-diphosphate; iPRMP, isopentenyl adenosine-5′-monophosphate; CYP735A, cytokinin trans-hydroxylase; tZRTP, trans-Zeatin riboside triphosphate; tZRDP, trans-Zeatin riboside diphosphate; tZRMP, trans-Zeatin riboside monophosphate; LOG, cytokinin-activating enzyme LONELY GUY; iPR, isopentenyl adenosine; tZR, trans-zeatin riboside; iP, isopentenyl adenine; tZT, trans-zeatin; AHP, histidine-containing phosphotransfer peotein; Type-A ARRs, type-A response regulators; Type-B ARRs, type-B response regulators. GL, non-senescing leaves, fully expanded green leaves; YL, senescing leaves, completely yellow; ~ 95% chlorophyll loss. The colored cell represents the expression profiles of differentially expressed genes (DEGs). The heatmaps were generated using the Rpackage and the color bar at the lower right. Orange dots show the fold change of the content of compounds in non-senescing and senescing leaves