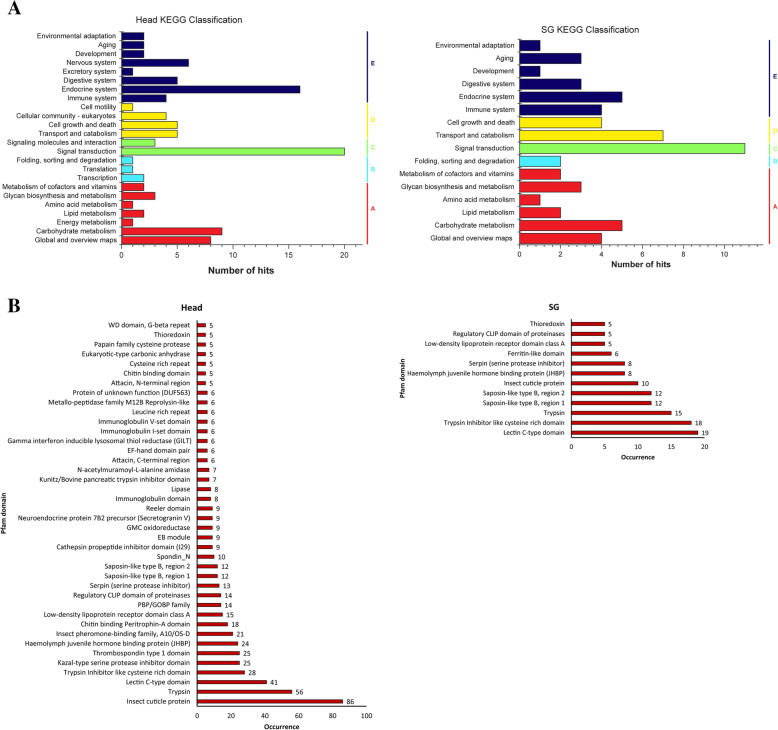
Fig. 7.
a The predicted KEGG pathways for all potential effector proteins. In KO database, a small number of proteins were annotated i.e. 142 out of 808 in the head and 80 out of 267 in the salivary gland were annotated. The proteins in the second hierarchy of the KEGG pathway were assigned to 5 categories A: Metabolism, B: Genetic Information Processing, C: Environmental Information Processing, D: Cellular Processes, E: Organismal Systems. b Predicted Pfam domains in the transcriptome of the head and Salivary gland of S. litura. In Pfam database, 558 out of 808 proteins in the head and 178 out of 267 proteins in the salivary gland were annotated. Pfam domains were mapped on potential effector proteins via Pfam standalone software using the default parameters. Numerical values at the top of each bar represent the number of times that domain has occurred in the annotated proteins. The domains which have occurred ≥5 times in the annotated proteins are shown here