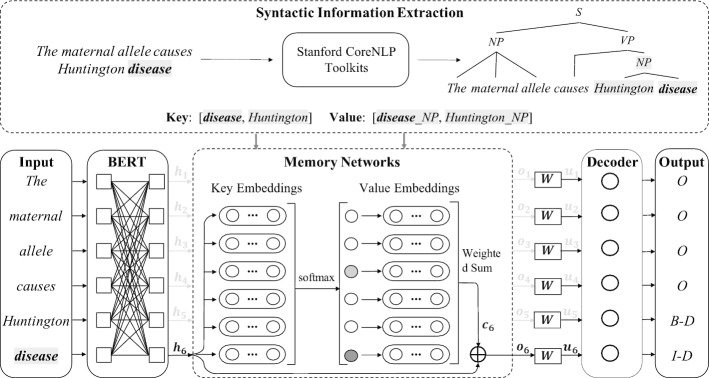
Fig. 3.
The overall architecture of BioKMNER. The top part of the figure shows the syntactic information extraction process: for the input word sequence, we firstly use off-the-shelf NLP toolkits to obtain its syntactic information (e.g., syntax tree), then map the context features and the syntactic information into keys and values, and finally convert them into embeddings. The bottom part is our sequence labeling based BioNER tagger, which uses BioBERT [19] as the encoder and a softmax layer as the decoder. Between the encoder and decoder are the key-value memory networks (KVMN) which weighs syntactic information (values) according to the importance of the context features (keys). The output of KVMN is fed into the decoder to predict output labels